Color Atlas of
Biochemistry
Second edition, revised and enlarged
Jan Koolman
Professor
Philipps University Marburg
Institute of Physiologic Chemistry
Marburg, Germany
Klaus-Heinrich Roehm
Professor
Philipps University Marburg
Institute of Physiologic Chemistry
Marburg, Germany
215 color plates by Juergen Wirth
Thieme
Stuttgart · New York
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
IV
Library of Congress Cataloging-in-
Important note: Medicine is an ever-changing
Publication Data
science undergoing continual development.
Research and clinical experience are continu-
ally expanding our knowledge, in particular
our knowledge of proper treatment and drug
This book is an authorized and updated trans-
therapy. Insofar as this book mentions any
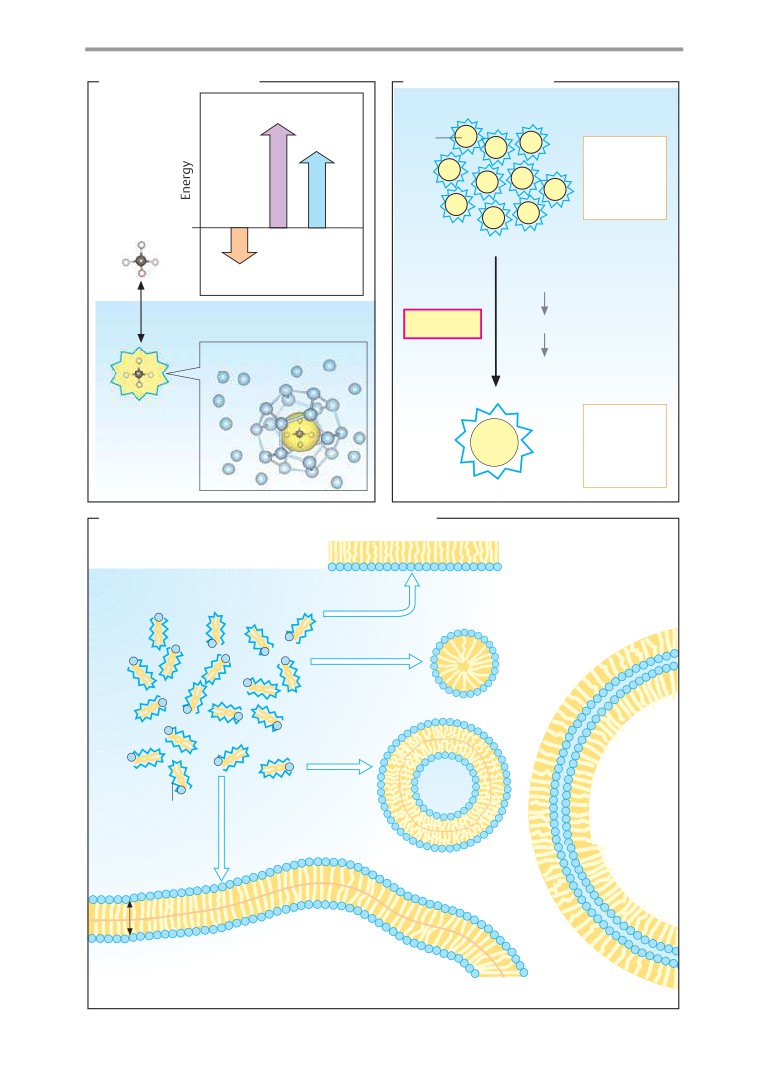
lation of the 3rd German edition published
dosage or application, readers may rest as-
and copyrighted 2003 by Georg Thieme Ver-
sured that the authors, editors, and publishers
lag, Stuttgart, Germany. Title of the German
have made every effort to ensure that such
edition: Taschenatlas der Biochemie
references are in accordance with the state of
knowledge at the time of production of the
book. Nevertheless, this does not involve, im-
Illustrator: Juergen Wirth, Professor of Visual
ply, or express any guarantee or responsibility
Communication, University of Applied Scien-
on the part of the publishers in respect to any
ces, Darmstadt, Germany
dosage instructions and forms of applications
stated in the book. Every user is requested to
examine carefully the manufacturers’ leaflets
Translator: Michael Robertson, BA DPhil,
accompanying each drug and to check, if nec-
Augsburg, Germany
essary in consultation with a physician or
specialist, whether the dosage schedules
mentioned therein or the contraindications
stated by the manufacturers differ from the
1st Dutch edition 2004
statements made in the present book. Such
1st English edition 1996
examination is particularly important with
1st French edition 1994
drugs that are either rarely used or have
2nd French edition 1999
been newly released on the market. Every
3rd French edition 2004
dosage schedule or every form of application
1st German edition 1994
used is entirely at the user’s own risk and
2nd German edition 1997
responsibility. The authors and publishers re-
1st Greek edition 1999
quest every user to report to the publishers
1st Indonesian edition 2002
any discrepancies or inaccuracies noticed. If
1st Italian edition 1997
errors in this work are found after publication,
1st Japanese edition 1996
1st Portuguese edition 2004
the product description page.
1st Russian edition 2000
1st Spanish edition 2004
Some of the product names, patents, and reg-
istered designs referred to in this book are in
© 2005 Georg Thieme Verlag
fact registered trademarks or proprietary
Rüdigerstrasse 14, 70469 Stuttgart,
names even though specific reference to this
Germany
fact is not always made in the text. Therefore,
the appearance of a name without designa-
Thieme New York, 333 Seventh Avenue,
tion as proprietary is not to be construed as a
New York, NY 10001 USA
representation by the publisher that it is in
the public domain.
This book, including all parts thereof, is legally
Cover design: Cyclus, Stuttgart
protected by copyright. Any use, exploitation,
Cover drawing: CAP cAMP bound to DNA
or commercialization outside the narrow lim-
Typesetting by primustype Hurler GmbH,
its set by copyright legislation, without the
Notzingen
publisher’s consent, is illegal and liable to
Printed in Germany by Appl, Wemding
prosecution. This applies in particular to pho-
tostat reproduction, copying, mimeograph-
ISBN 3-13-100372-3 (GTV)
ing, preparation of microfilms, and electronic
ISBN 1-58890-247-1 (TNY)
data processing and storage.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
V
About the Authors
Jan Koolman (left) was born in Lübeck, Ger-
took his doctorate under the supervision of
many, and grew up with the sea wind blowing
Friedhelm Schneider, and his postdoctoral de-
off the Baltic. The high school he attended in
gree in 1980 was in the Department of Chem-
the Hanseatic city of Lübeck was one that
istry. He has been an Honorary Professor since
focused on providing a classical education,
1986. His research group is concerned with
which left its mark on him. From 1963 to
the structure and function of enzymes in-
1969, he studied biochemistry at the Univer-
volved in amino acid metabolism. He is mar-
sity of Tübingen. He then took his doctorate
ried to a biologist and has two children.
(in the discipline of chemistry) at the Univer-
Jürgen Wirth (center) studied in Berlin and at
sity of Marburg, under the supervision of bio-
the College of Design in Offenbach, Germany.
chemist Peter Karlson. In Marburg, he began
His studies focused on free graphics and illus-
to study the biochemistry of insects and other
tration, and his diploma topic was “The devel-
invertebrates. He took his postdoctoral de-
opment and function of scientific illustration.”
gree in 1977 in the field of human medicine,
From 1963 to 1977, Jürgen Wirth was involved
and was appointed Honorary Professor in
in designing the exhibition space in the
1984. His field of study today is biochemical
Senckenberg Museum of Natural History in
endocrinology. His other interests include ed-
Frankfurt am Main, while at the same time
ucational methods in biochemistry. He is cur-
working as a freelance associate with several
rently Dean of Studies in the Department of
publishing companies, providing illustrations
Medicine in Marburg; he is married to an art
for schoolbooks, non-fiction titles, and scien-
teacher.
tific publications. He has received several
Klaus-Heinrich Röhm
(right) comes from
awards for book illustration and design. In
Stuttgart, Germany. After graduating from
1978, he was appointed to a professorship at
the School of Protestant Theology in Urach
the College of Design in Schwäbisch Gmünd,
—another institution specializing in classical
Germany, and in 1986 he became Professor of
studies—and following a period working in
Design at the Academy of Design in Darm-
the field of physics, he took a diploma in bio-
stadt, Germany. His specialist fields include
chemistry at the University of Tübingen,
scientific graphics/information graphics and
where the two authors first met. Since 1970,
illustration methods. He is married and has
he has also worked in the Department of
three children.
Medicine at the University of Marburg. He
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
VI
Preface
Biochemistry is a dynamic, rapidly growing
elements had to be found that make compli-
field, and the goal of this color atlas is to
cated phenomena appear tangible. The
illustrate this fact visually. The precise boun-
graphics were designed conservatively, the
daries between biochemistry and related
aim being to avoid illustrations that might
fields, such as cell biology, anatomy, physiol-
look too spectacular or exaggerated. Our
ogy, genetics, and pharmacology, are dif cult
goal was to achieve a visual and aesthetic
to define and, in many cases, arbitrary. This
way of representing scientific facts that would
overlap is not coincidental. The object being
be simple and at the same time effective for
studied is often the same—a nerve cell or a
teaching purposes. Use of graphics software
mitochondrion, for example—and only the
helped to maintain consistency in the use of
point of view differs.
shapes, colors, dimensions, and labels, in par-
For a considerable period of its history, bio-
ticular. Formulae and other repetitive ele-
chemistry was strongly influenced by chem-
ments and structures could be handled easily
istry and concentrated on investigating met-
and precisely with the assistance of the com-
abolic conversions and energy transfers. Ex-
puter.
plaining the composition, structure, and me-
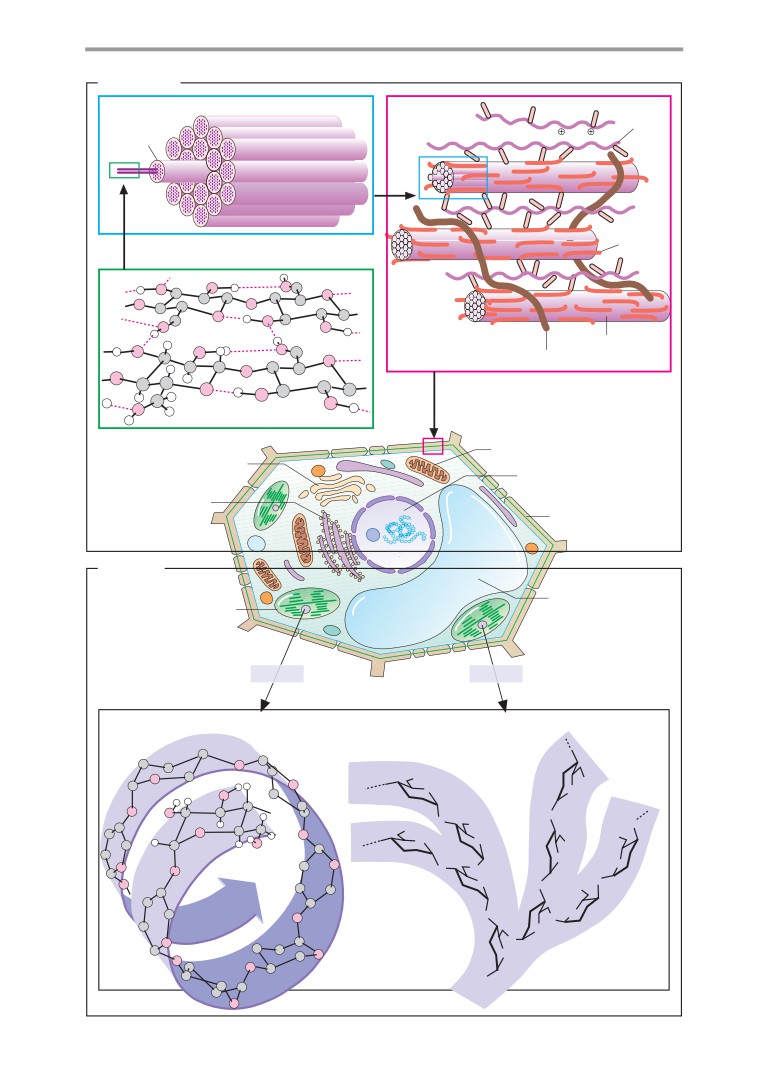
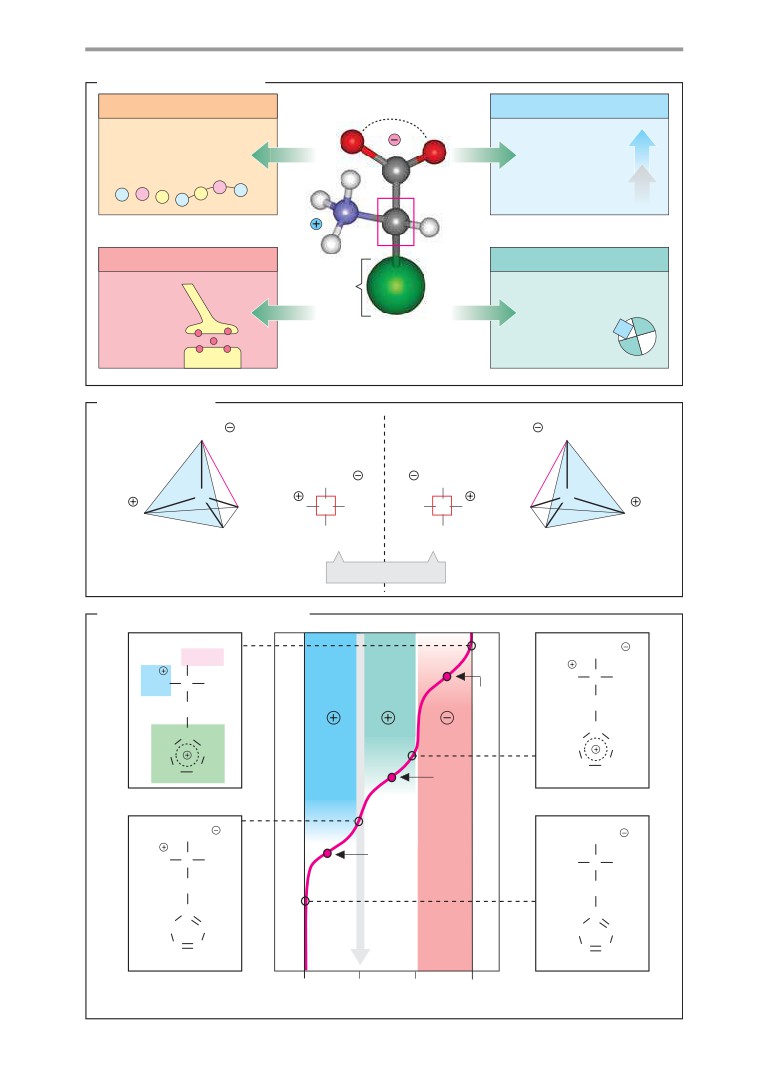
Color-coding has been used throughout to aid
tabolism of biologically important molecules
the reader, and the key to this is given in two
has always been in the foreground. However,
special color plates on the front and rear in-
new aspects inherited from biochemistry’s
side covers. For example, in molecular models
other parent, the biological sciences, are
each of the more important atoms has a par-
now increasingly being added: the relation-
ticular color: gray for carbon, white for hydro-
ship between chemical structure and biolog-
gen, blue for nitrogen, red for oxygen, and so
ical function, the pathways of information
on. The different classes of biomolecules are
transfer, observance of the ways in which
also distinguished by color: proteins are al-
biomolecules are spatially and temporally dis-
ways shown in brown tones, carbohydrates in
tributed in cells and organisms, and an aware-
violet, lipids in yellow, DNA in blue, and RNA
ness of evolution as a biochemical process.
in green. In addition, specific symbols are
These new aspects of biochemistry are bound
used for the important coenzymes, such as
to become more and more important.
ATP and NAD+. The compartments in which
Owing to space limitations, we have concen-
biochemical processes take place are color-
trated here on the biochemistry of humans
coded as well. For example, the cytoplasm is
and mammals, although the biochemistry of
shown in yellow, while the extracellular space
other animals, plants, and microorganisms is
is shaded in blue. Arrows indicating a chem-
no less interesting. In selecting the material
ical reaction are always black and those rep-
for this book, we have put the emphasis on
resenting a transport process are gray.
subjects relevant to students of human med-
In terms of the visual clarity of its presenta-
icine. The main purpose of the atlas is to serve
tion, biochemistry has still to catch up with
as an overview and to provide visual informa-
anatomy and physiology. In this book, we
tion quickly and ef ciently. Referring to text-
sometimes use simplified ball-and-stick mod-
books can easily fill any gaps. For readers
els instead of the classical chemical formulae.
encountering biochemistry for the first time,
In addition, a number of compounds are rep-
some of the plates may look rather complex. It
resented by space-filling models. In these
must be emphasized, therefore, that the atlas
cases, we have tried to be as realistic as pos-
is not intended as a substitute for a compre-
sible. The models of small molecules are
hensive textbook of biochemistry.
based on conformations calculated by com-
As the subject matter is often dif cult to vis-
puter-based molecular modeling. In illustrat-
ualize, symbols, models, and other graphic
ing macromolecules, we used structural infor-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Preface
VII
mation obtained by X-ray crystallography
We are grateful to many readers for their
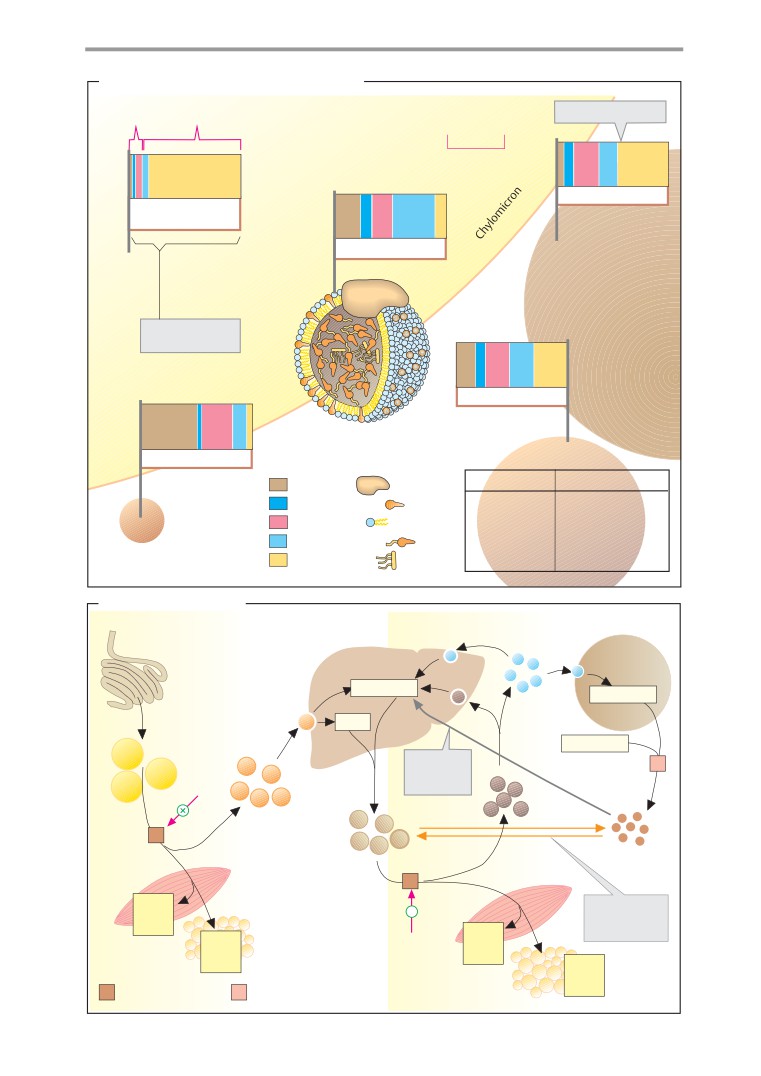
that is stored in the Protein Data Bank. In
comments and valuable criticisms during the
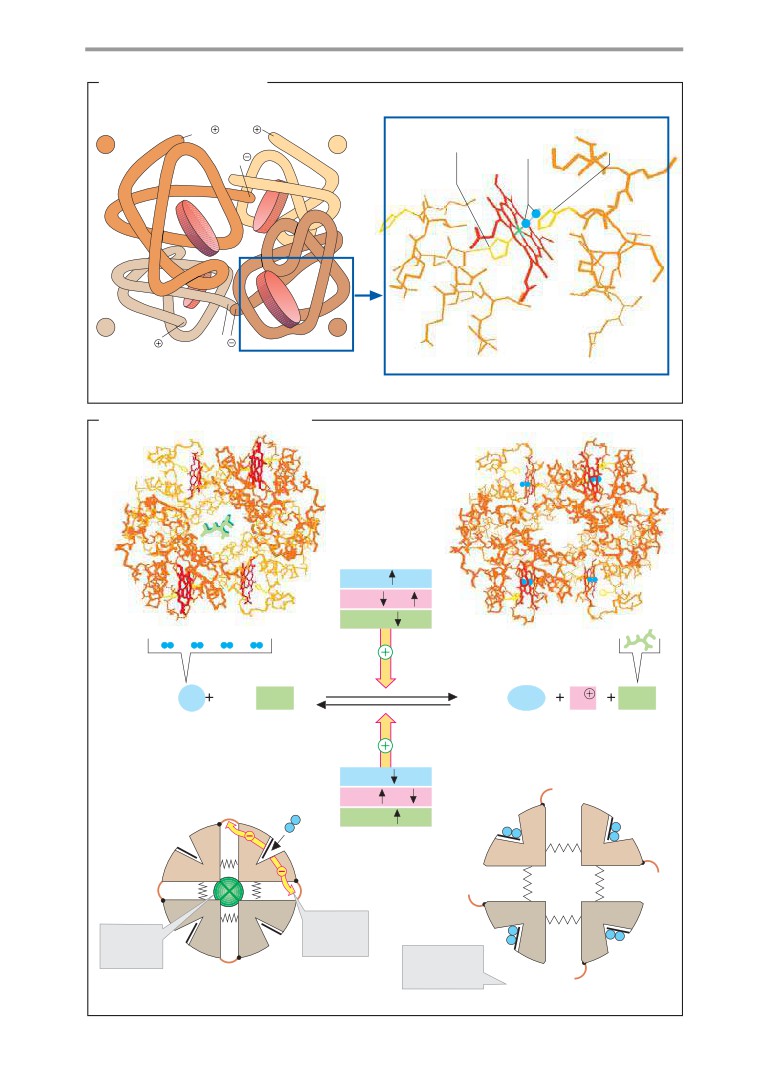
naming enzymes, we have followed the of -
preparation of this book. Of course, we would
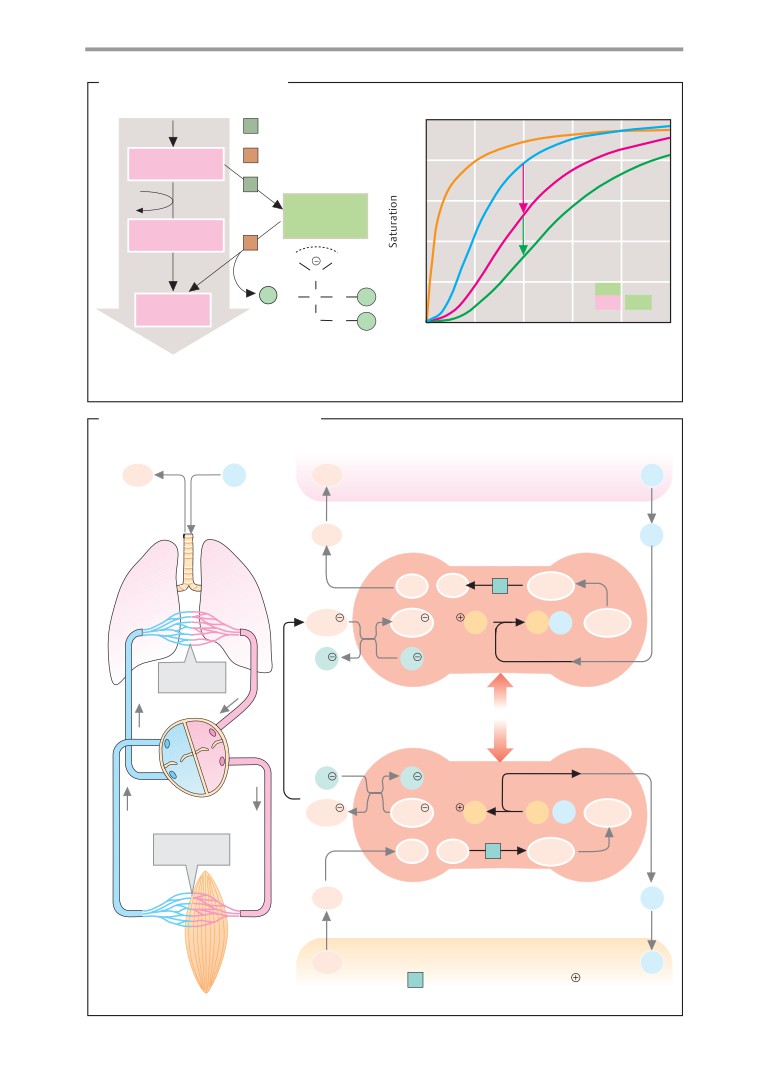
cial nomenclature recommended by the
also welcome further comments and sugges-
IUBMB. For quick identification, EC numbers
tions from our readers.
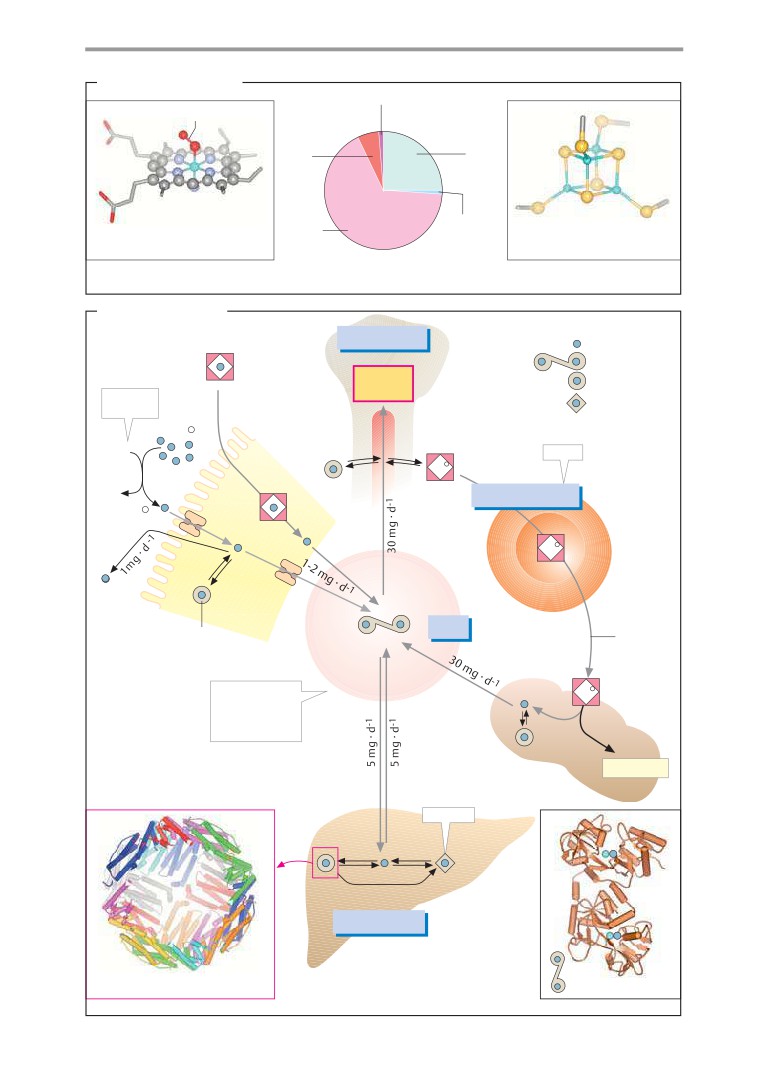
(in italics) are included with enzyme names.
To help students assess the relevance of the
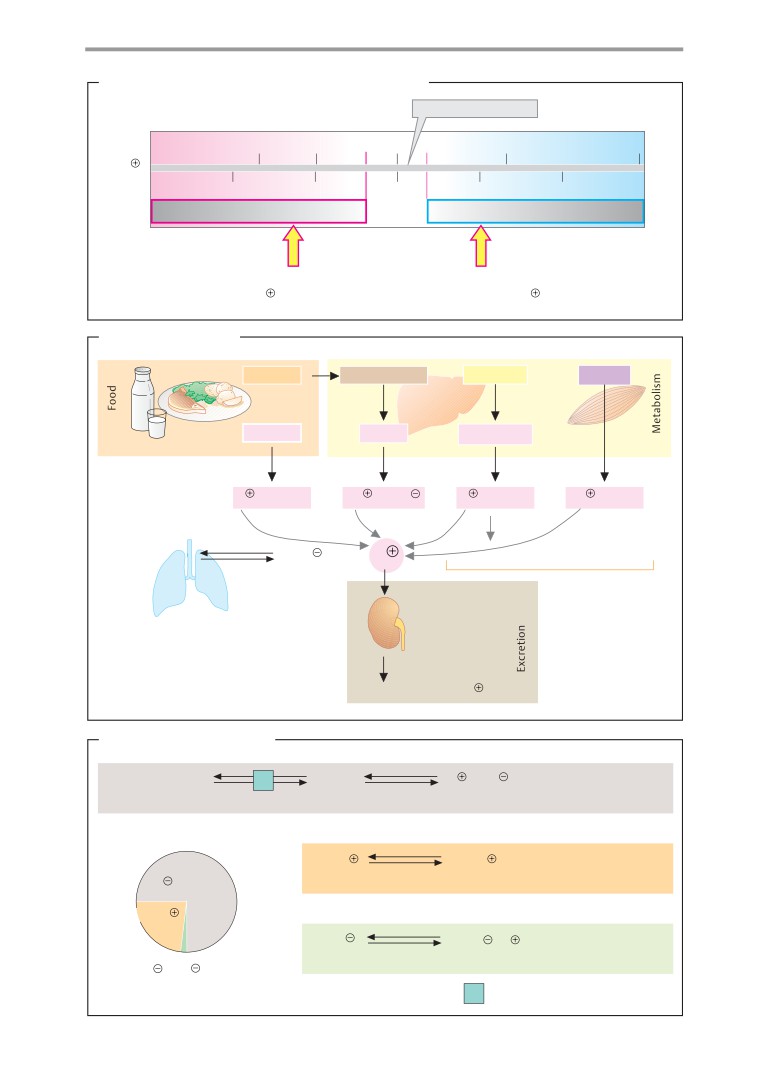
material (while preparing for an examination,
August 2004
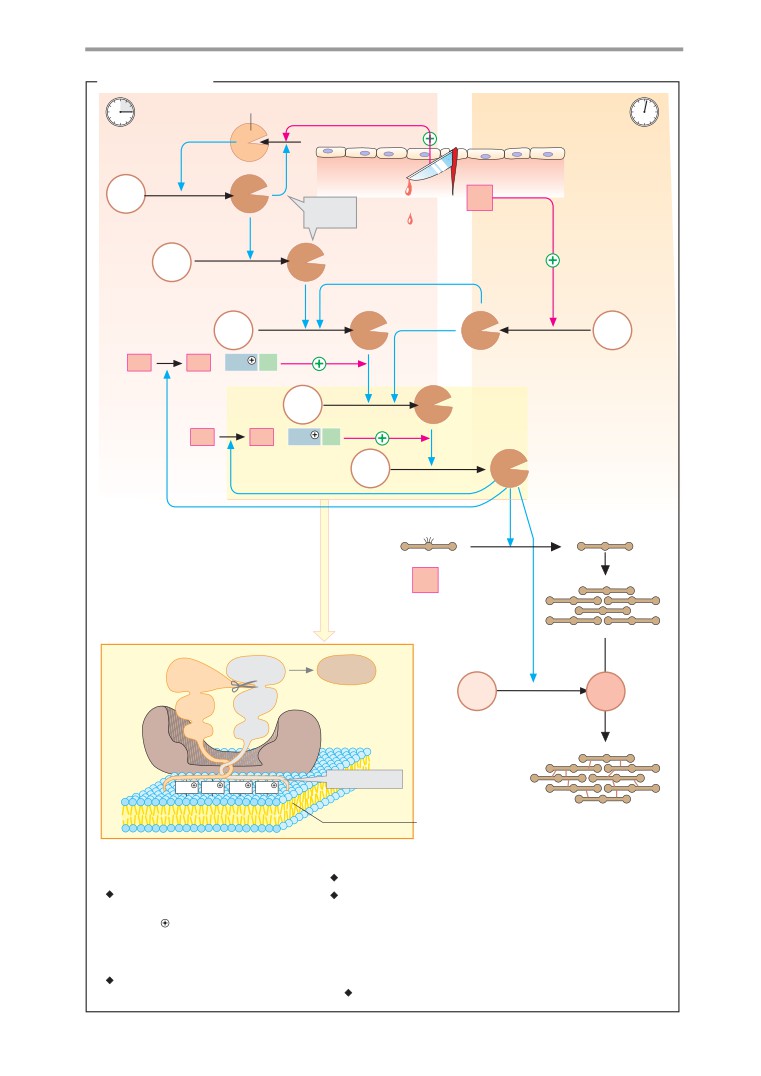
for example), we have included symbols on
the text pages next to the section headings to
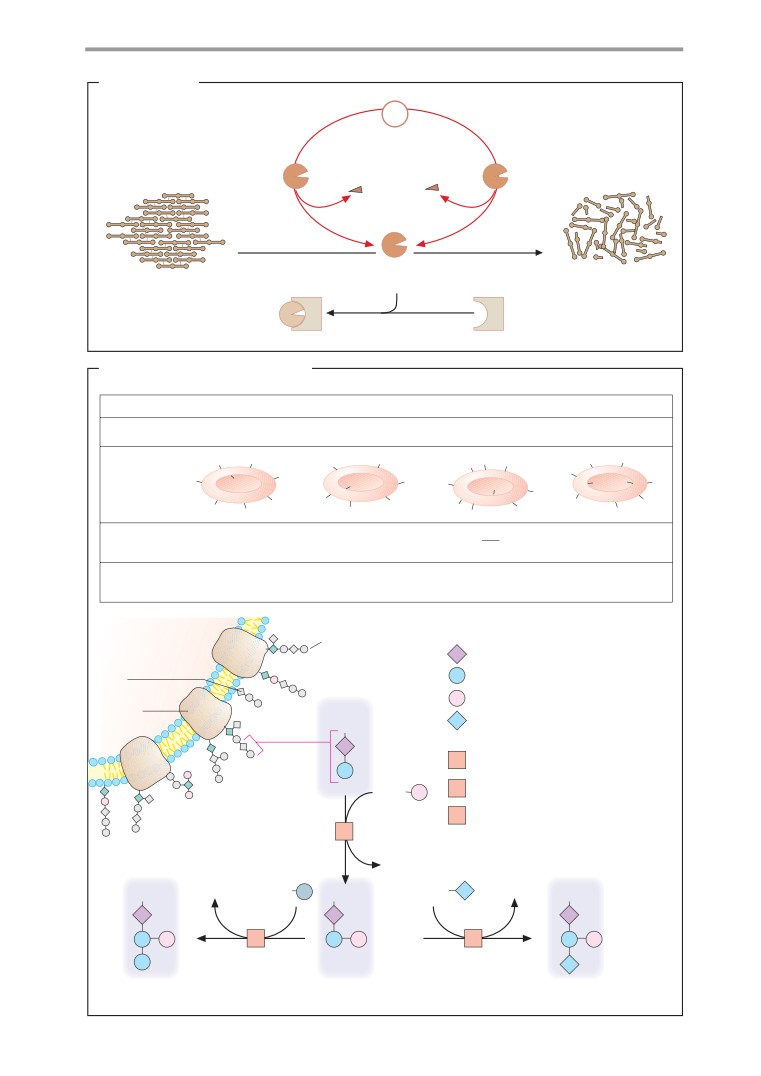
indicate how important each topic is. A filled
Jan Koolman,
circle stands for “basic knowledge,” a half-
Klaus-Heinrich Röhm
filled circle indicates “standard knowledge,”
Marburg
and an empty circle stands for
“in-depth
knowledge.” Of course, this classification
Jürgen Wirth
only reflects our subjective views.
Darmstadt
This second edition was carefully revised and
a significant number of new plates were
added to cover new developments.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
VIII
Contents
Introduction
1
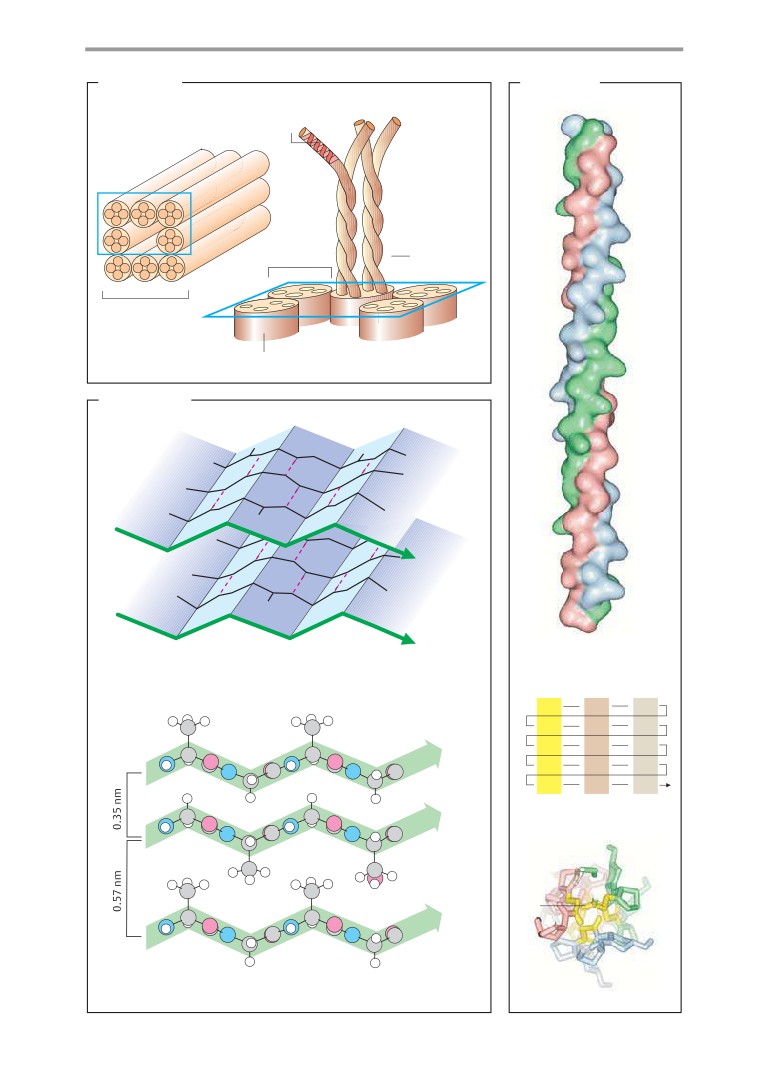
Structural proteins
70
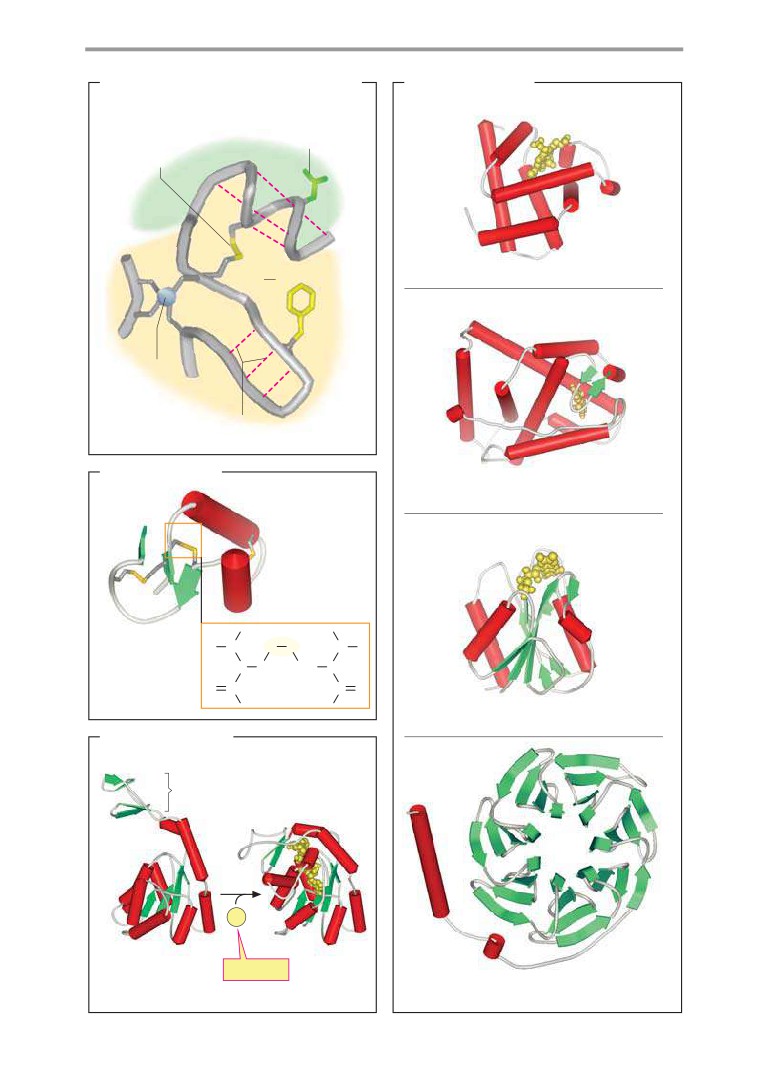
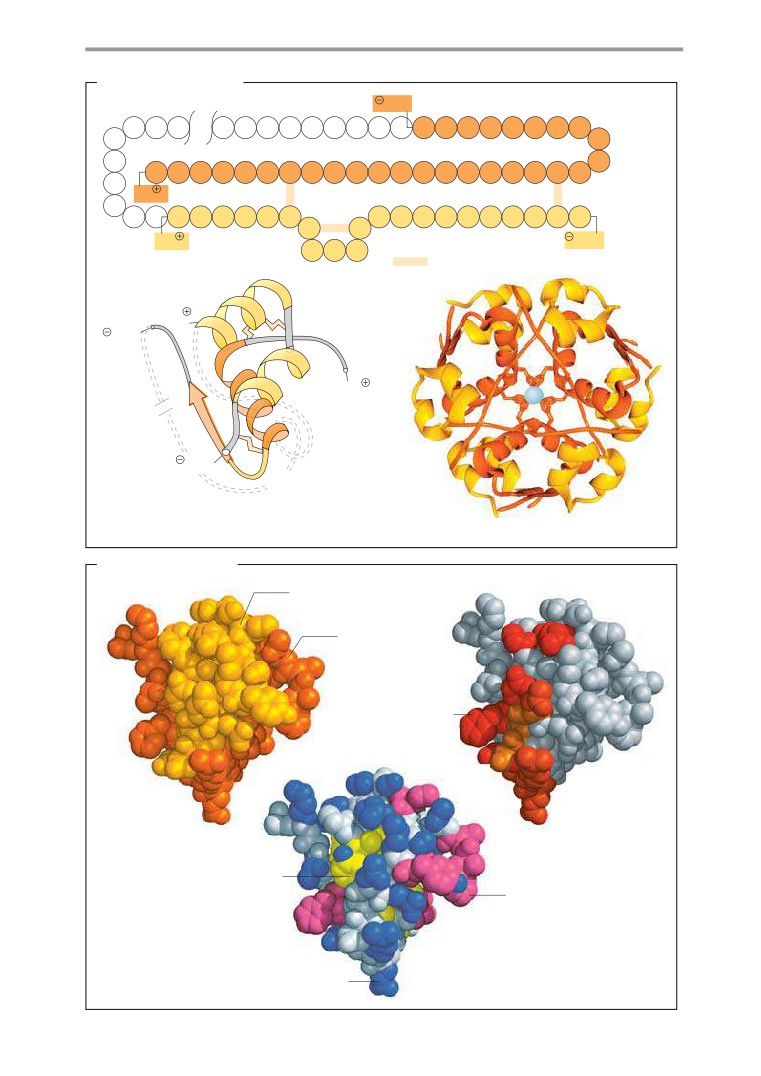
Globular proteins
72
Basics
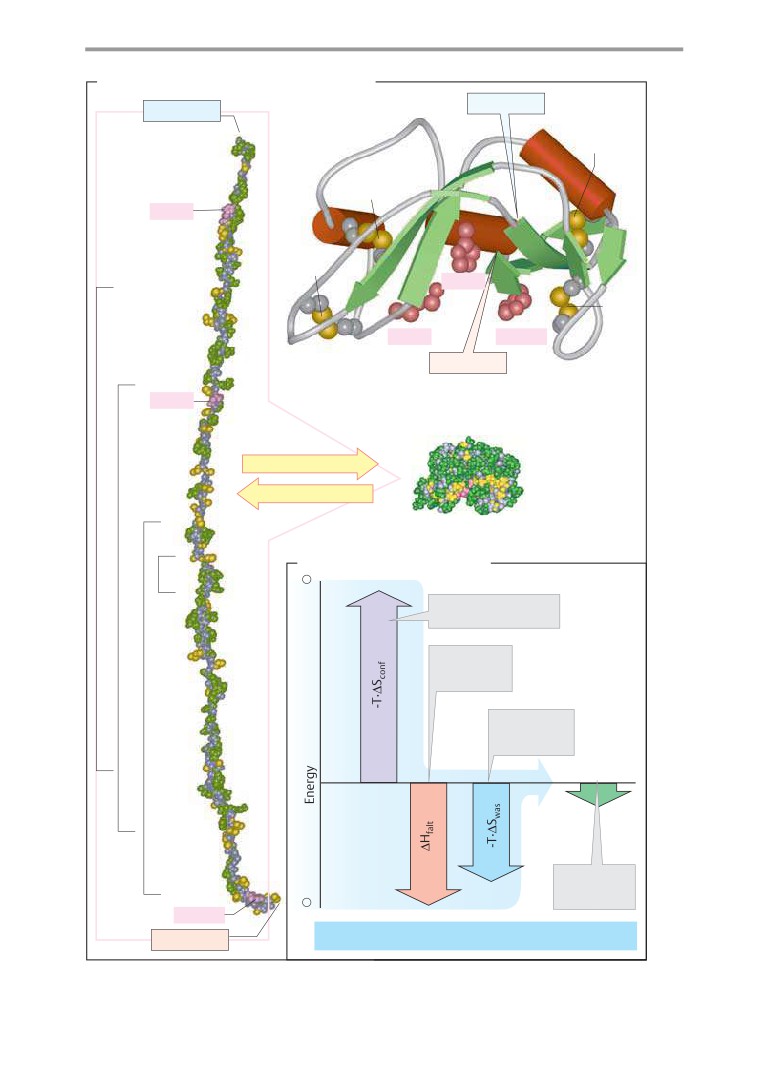
Protein folding
74
Molecular models: insulin
76
Chemistry
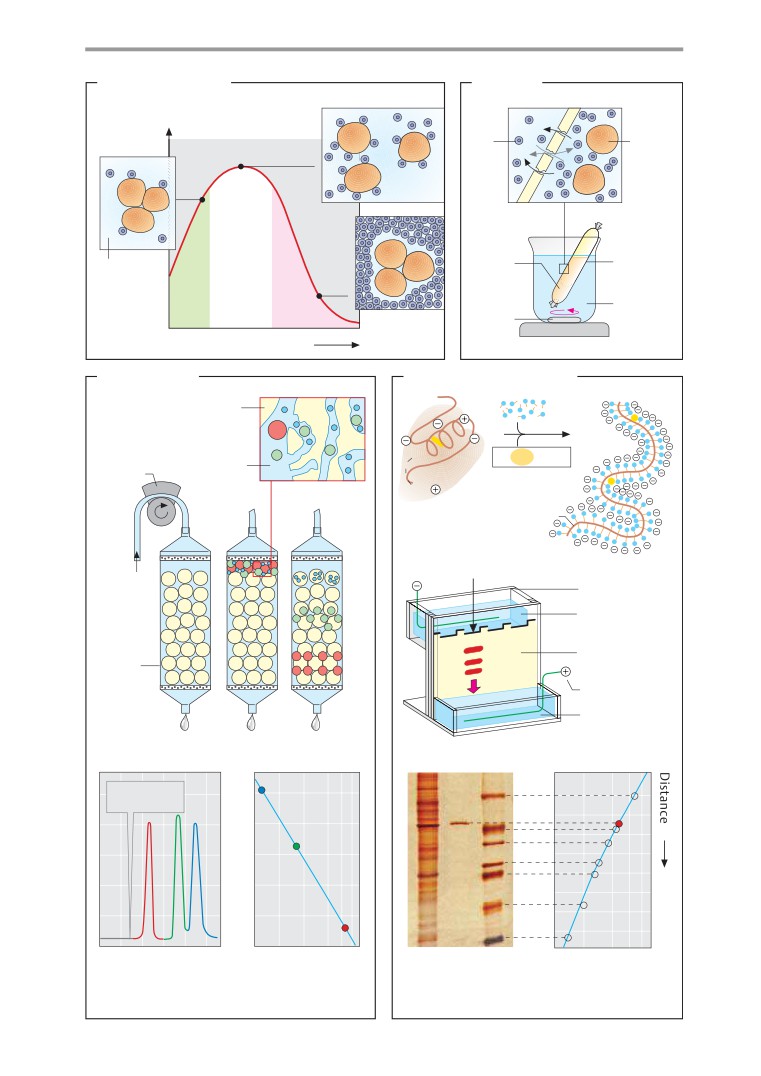
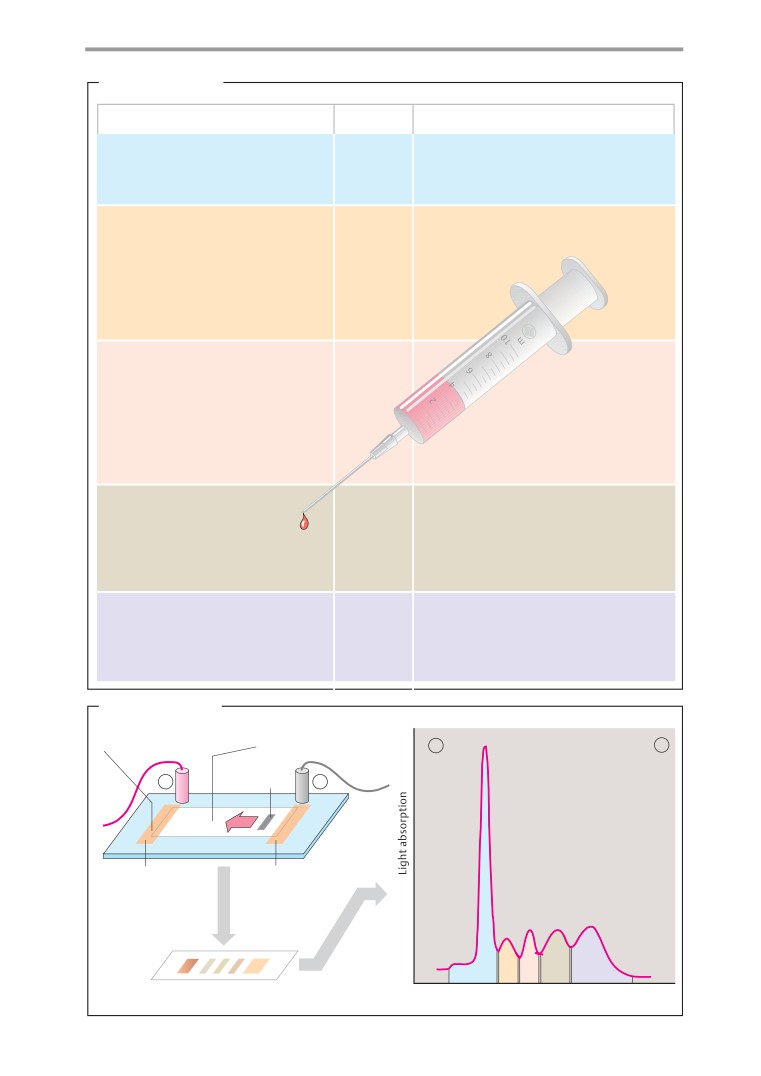
Isolation and analysis of proteins
78
Periodic table
2
Bonds
4
Nucleotides and Nucleic Acids
Molecular structure
6
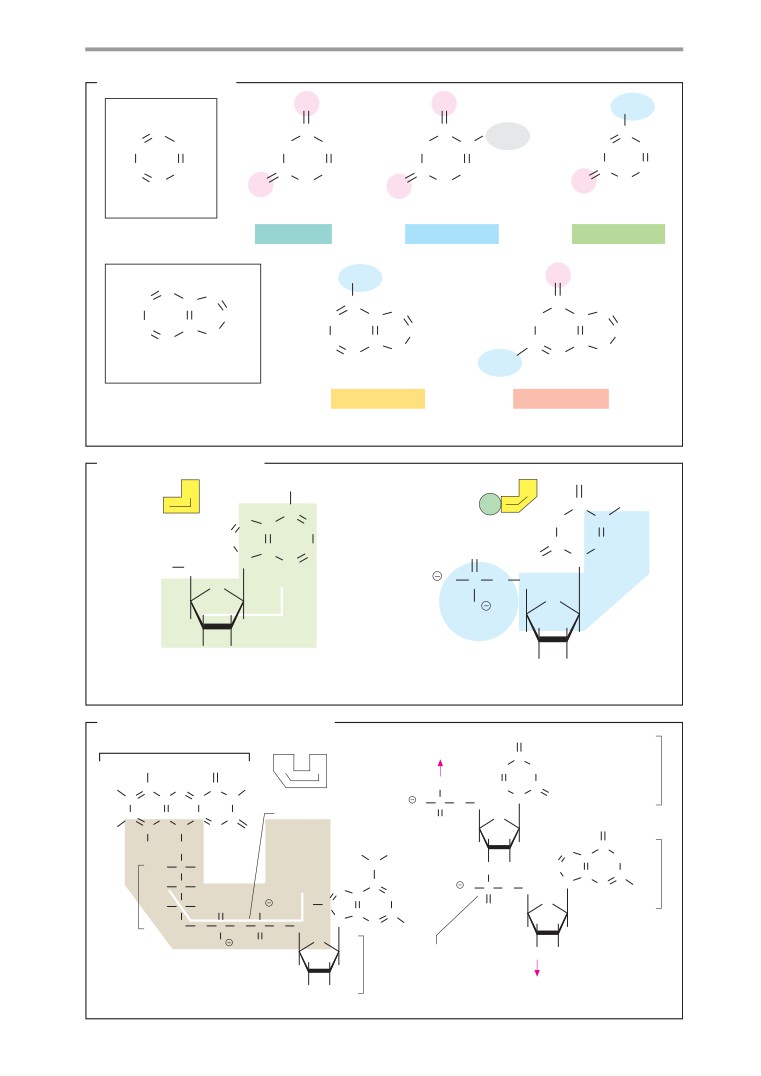
Bases and nucleotides
80
Isomerism
8
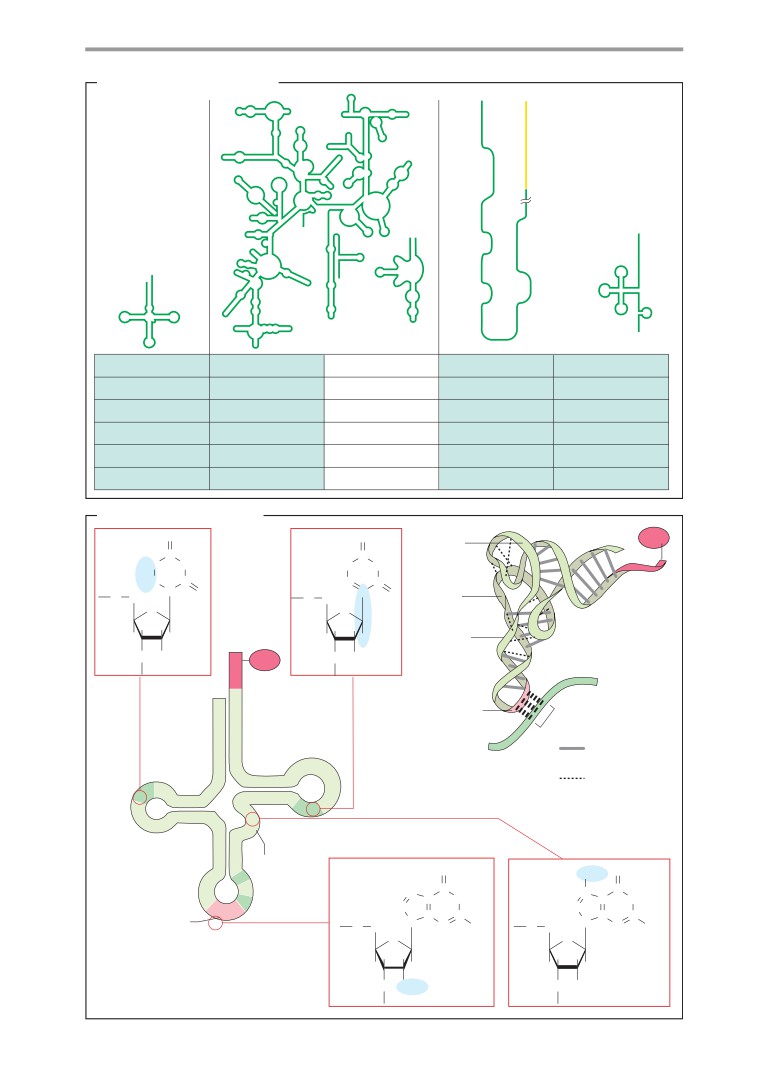
RNA
82
Biomolecules I
10
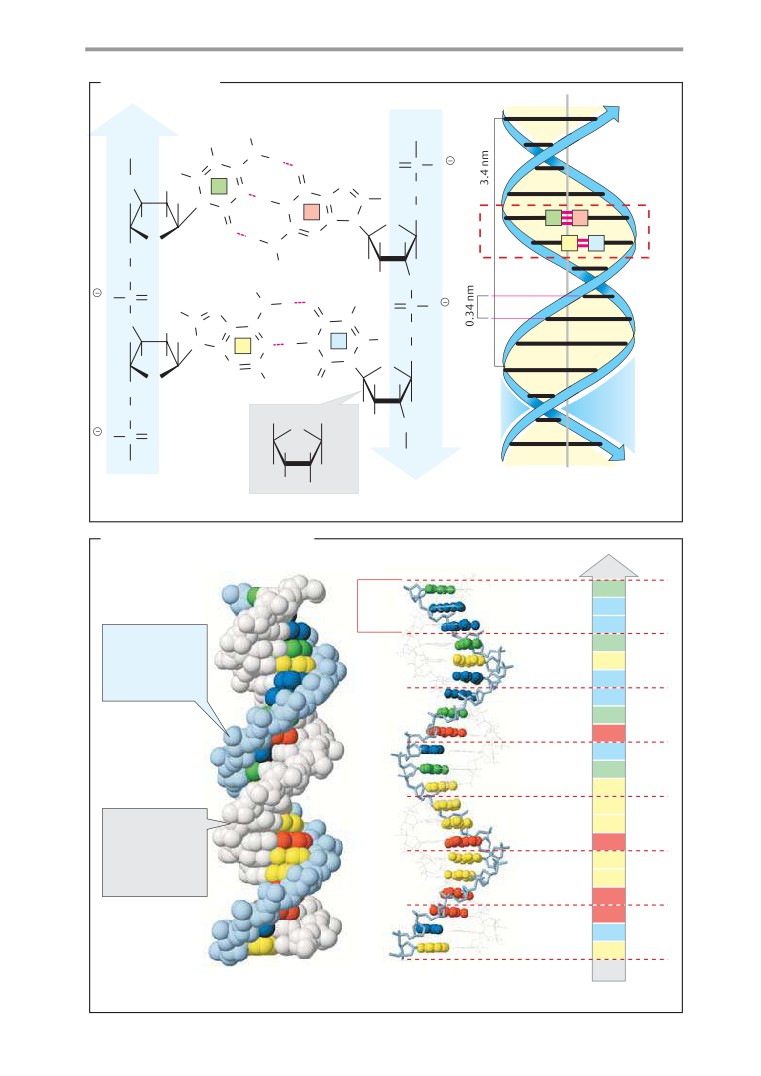
DNA
84
Biomolecules II
12
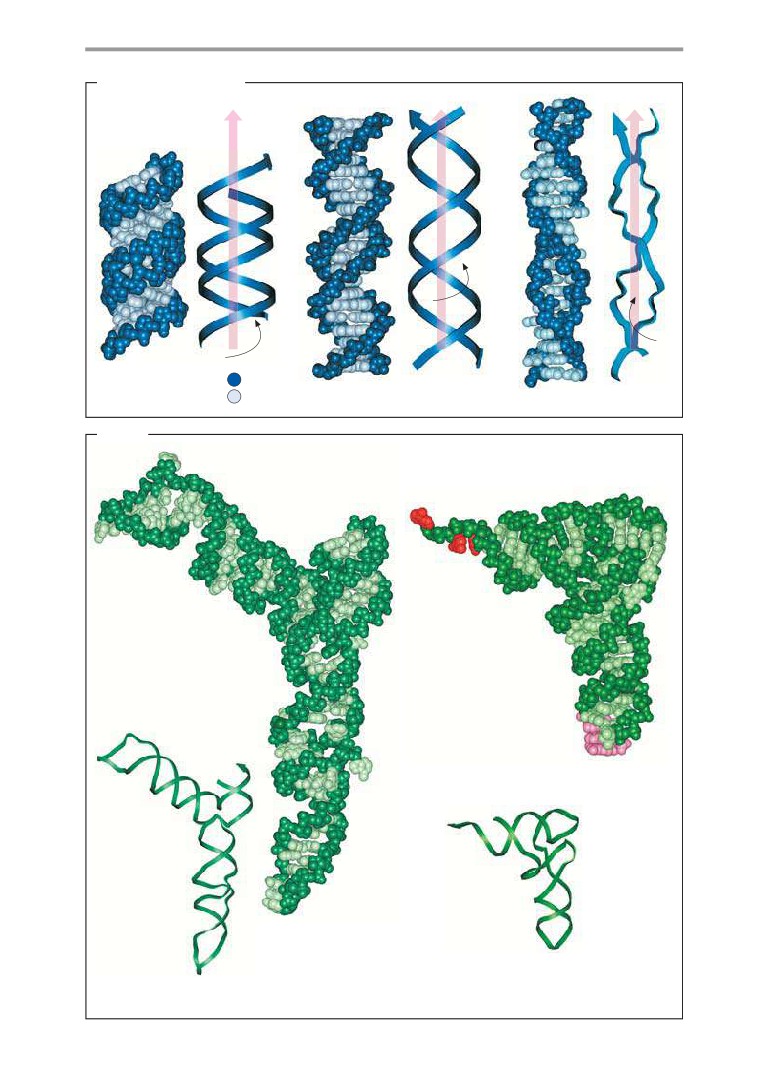
Molecular models: DNA and RNA
86
Chemical reactions
14
Metabolism
Physical Chemistry
Enzymes
Energetics
16
Basics
88
Equilibriums
18
Enzyme catalysis
90
Enthalpy and entropy
20
Enzyme kinetics I
92
Reaction kinetics
22
Enzyme kinetics II
94
Catalysis
24
Inhibitors
96
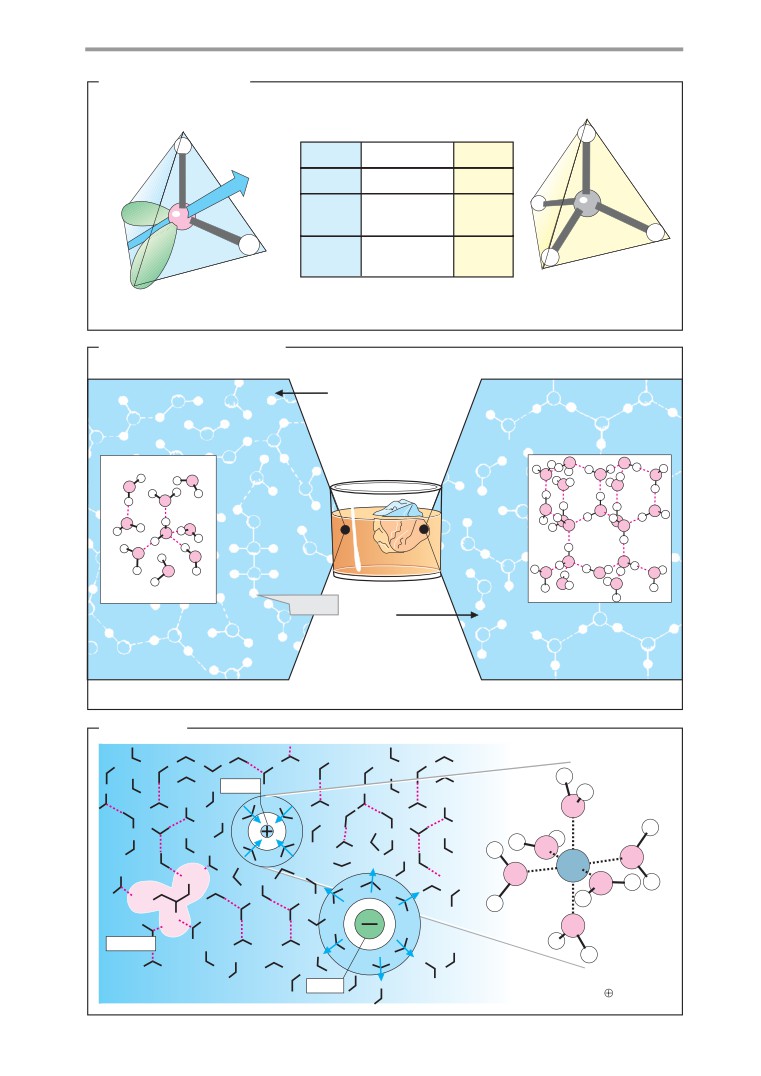
Water as a solvent
26
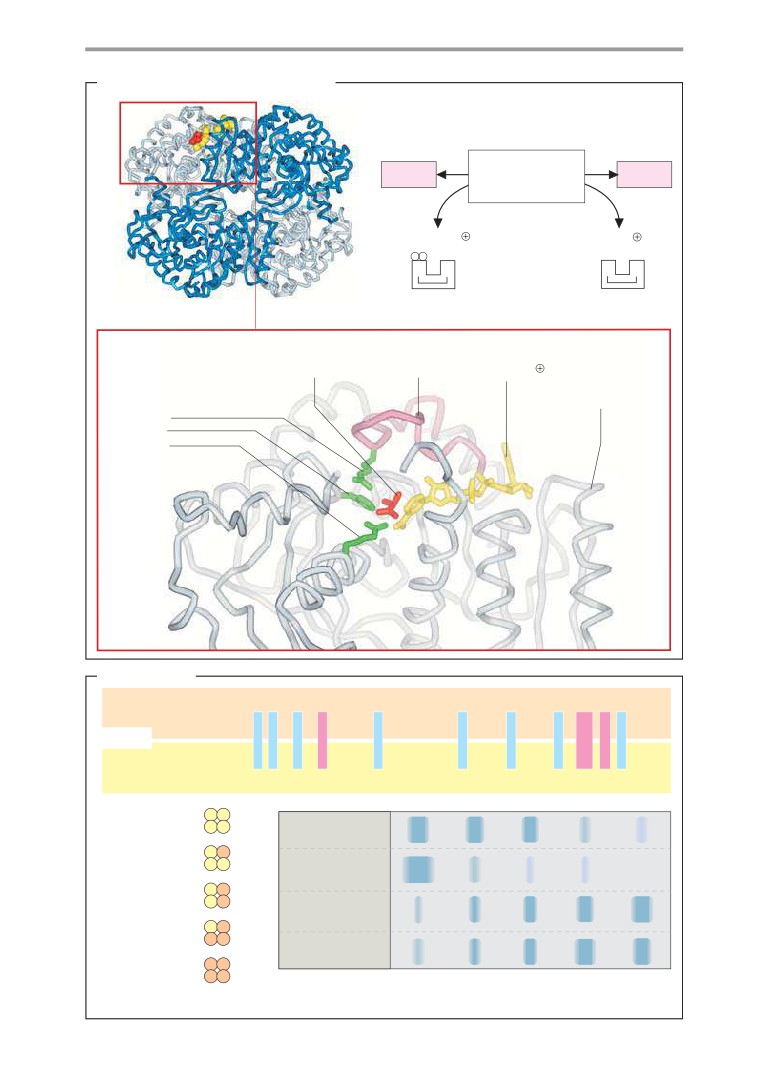
Lactate dehydrogenase: structure
98
Hydrophobic interactions
28
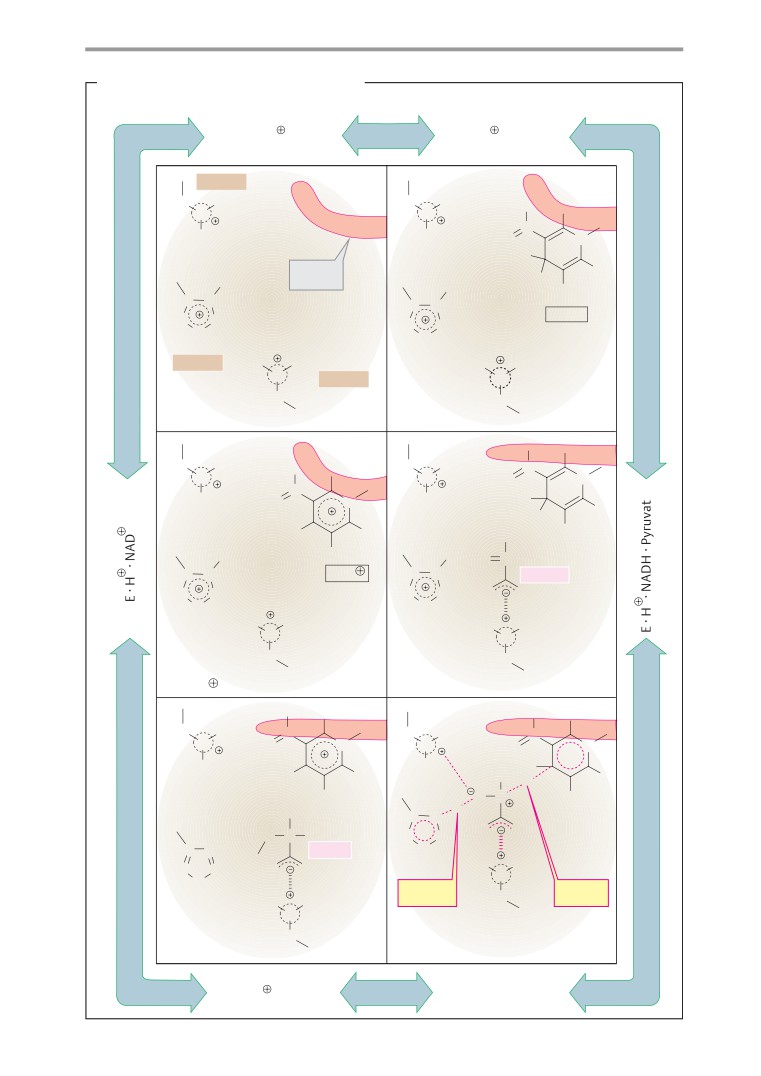
Lactate dehydrogenase: mechanism . . .
100
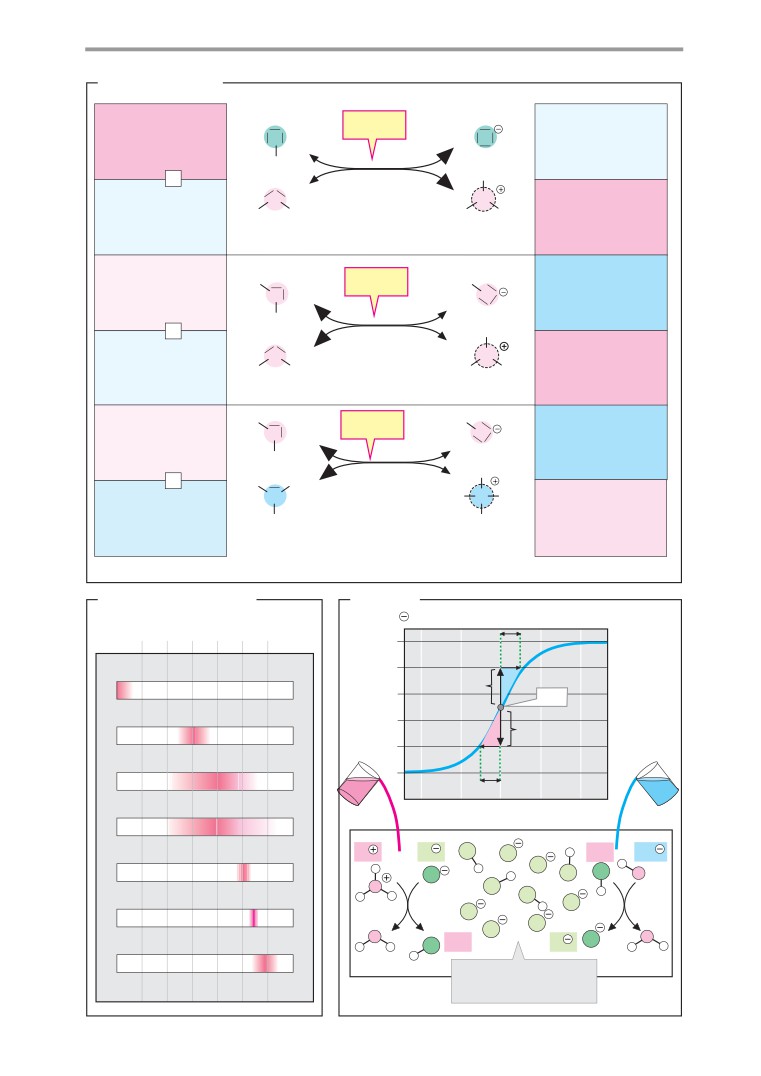
Acids and bases
30
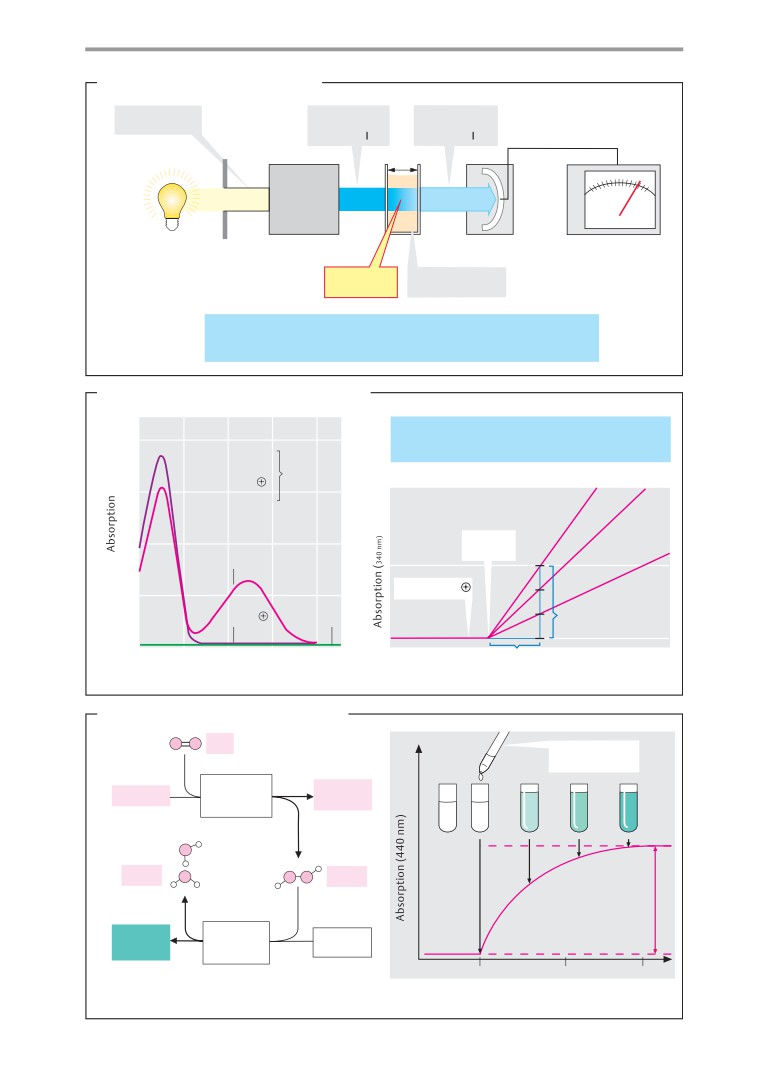
Enzymatic analysis
102
Redox processes
32
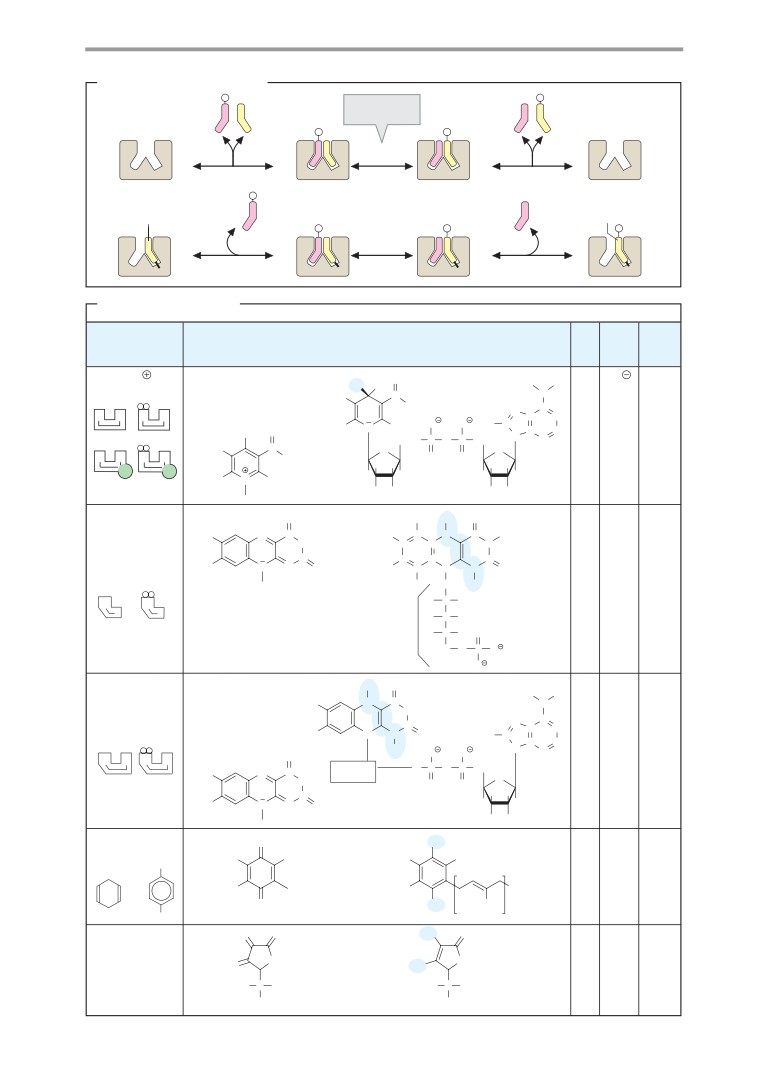
Coenzymes 1
104
Coenzymes 2
106
Biomolecules
Coenzymes 3
108
Carbohydrates
Activated metabolites
110
Overview
34
Metabolic Regulation
Chemistry of sugars
36
Intermediary metabolism
112
Monosaccharides and disaccharides . . .
38
Regulatory mechanisms
114
Polysaccharides: overview
40
Allosteric regulation
116
Plant polysaccharides
42
Transcription control
118
Glycosaminoglycans and glycoproteins .
44
Hormonal control
120
Lipids
Energy Metabolism
Overview
46
ATP
122
Fatty acids and fats
48
Energetic coupling
124
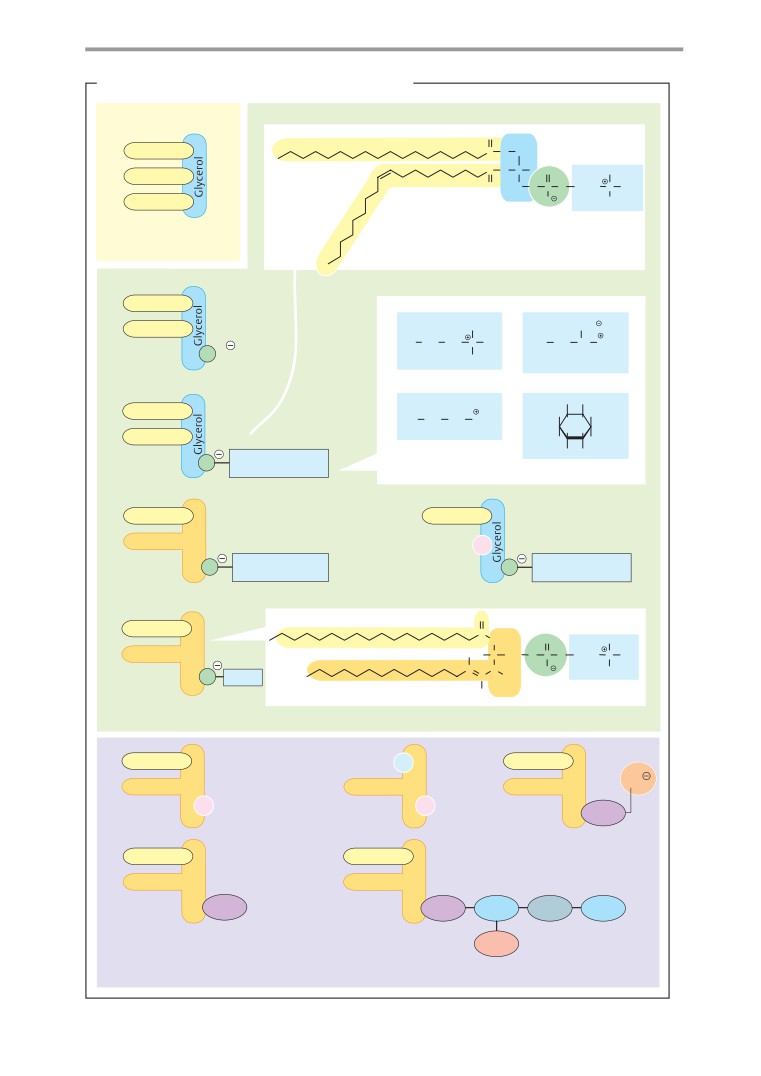
Phospholipids and glycolipids
50
Energy conservation at membranes
126
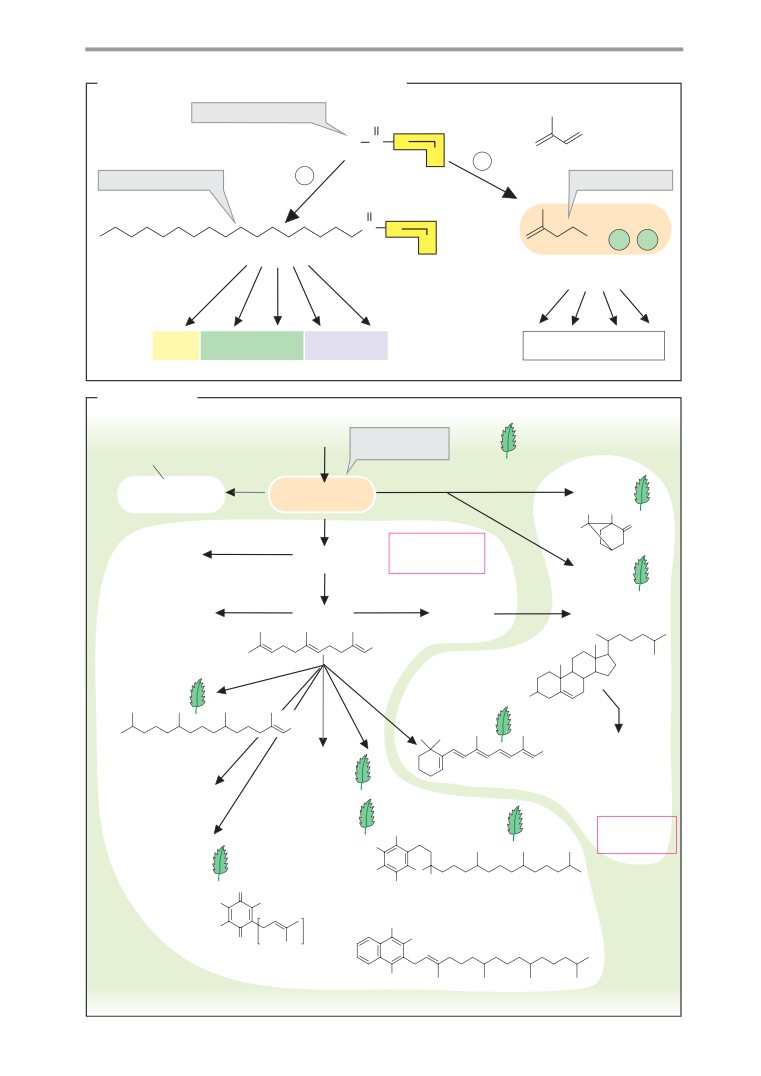
Isoprenoids
52
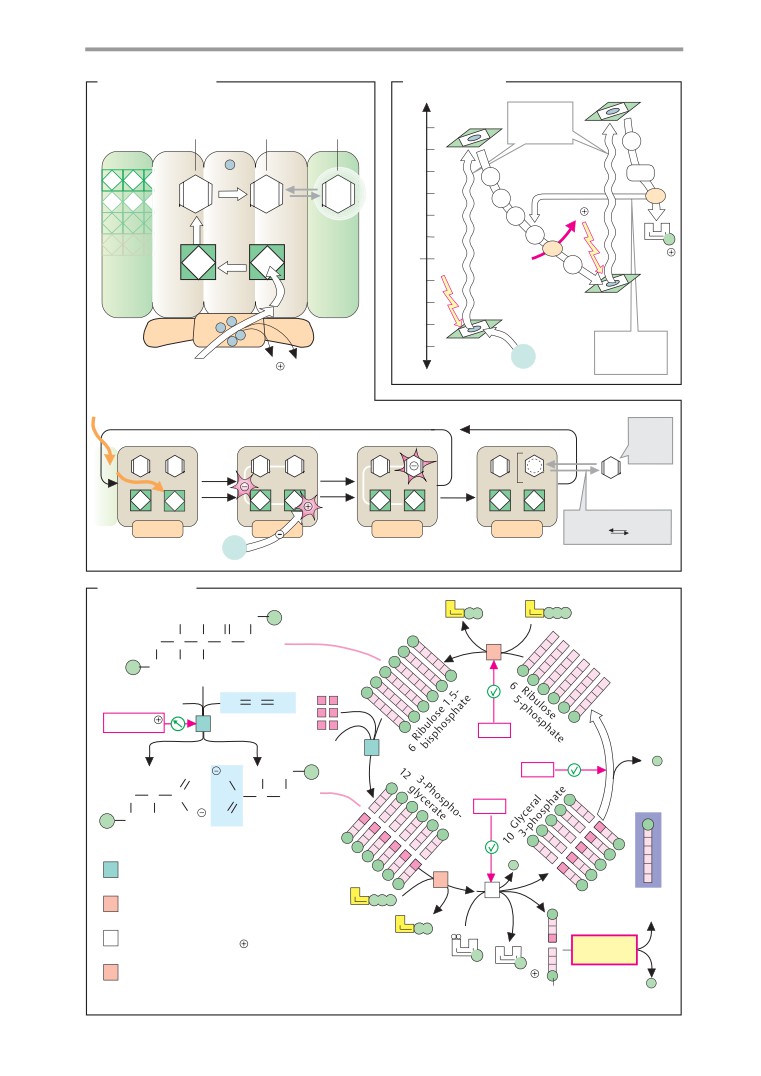
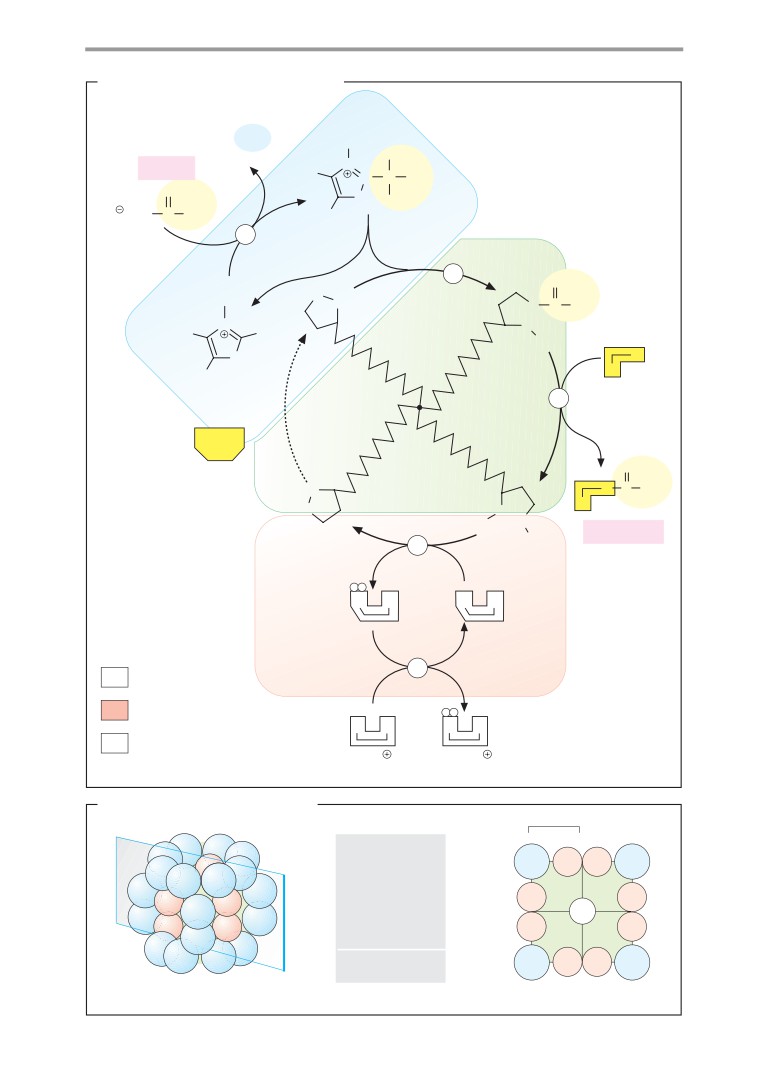
Photosynthesis: light reactions
128
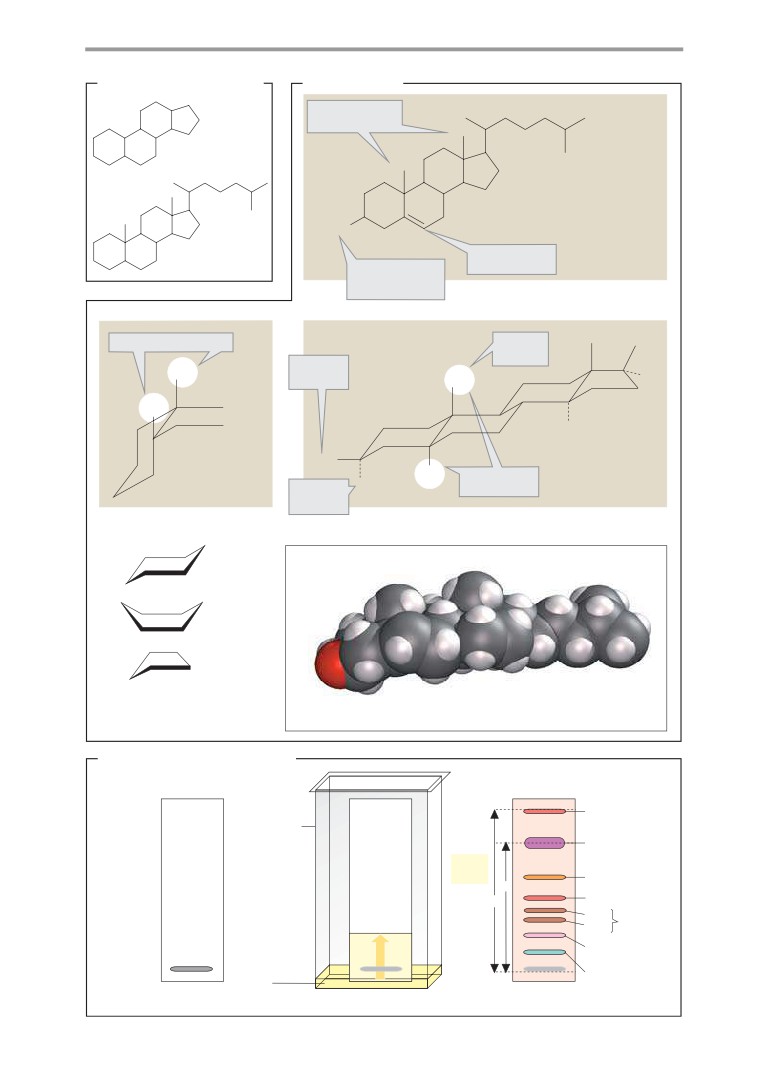
Steroid structure
54
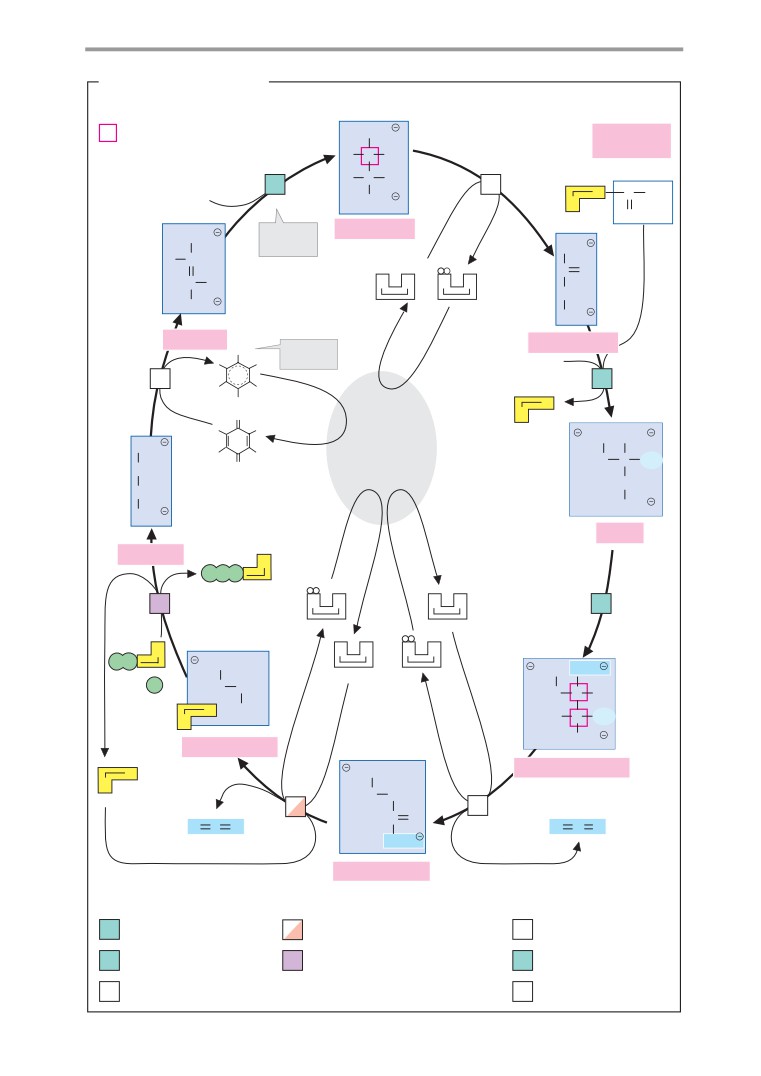
Photosynthesis: dark reactions
130
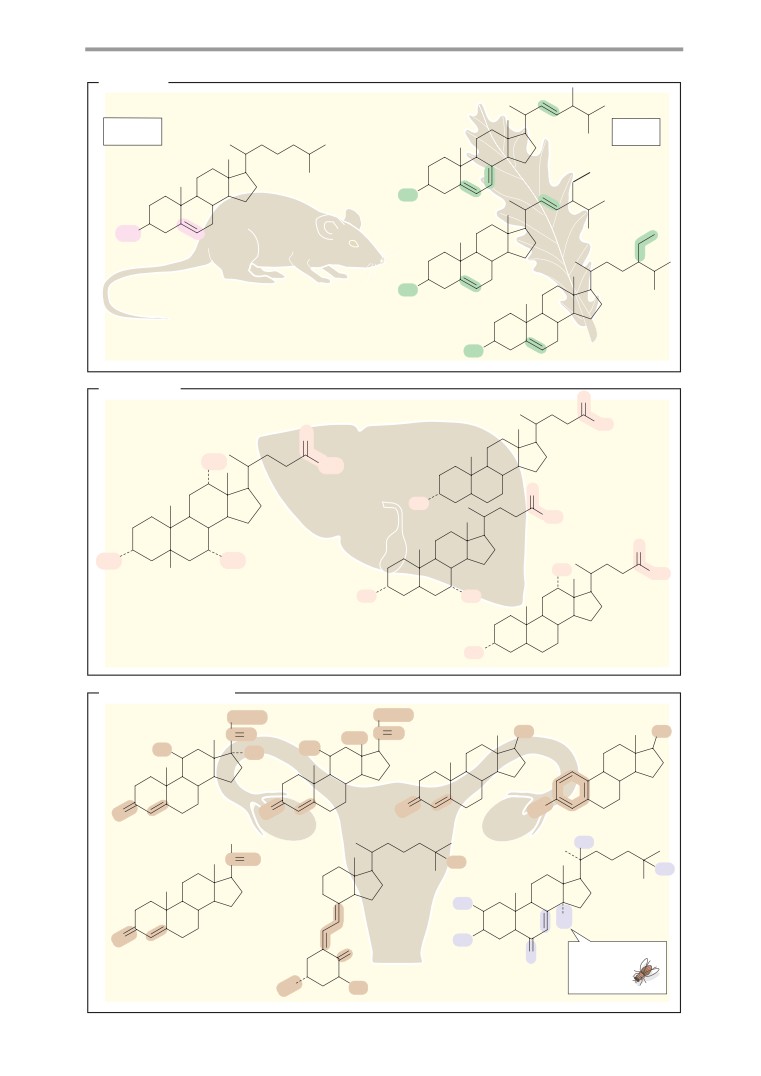
Steroids: overview
56
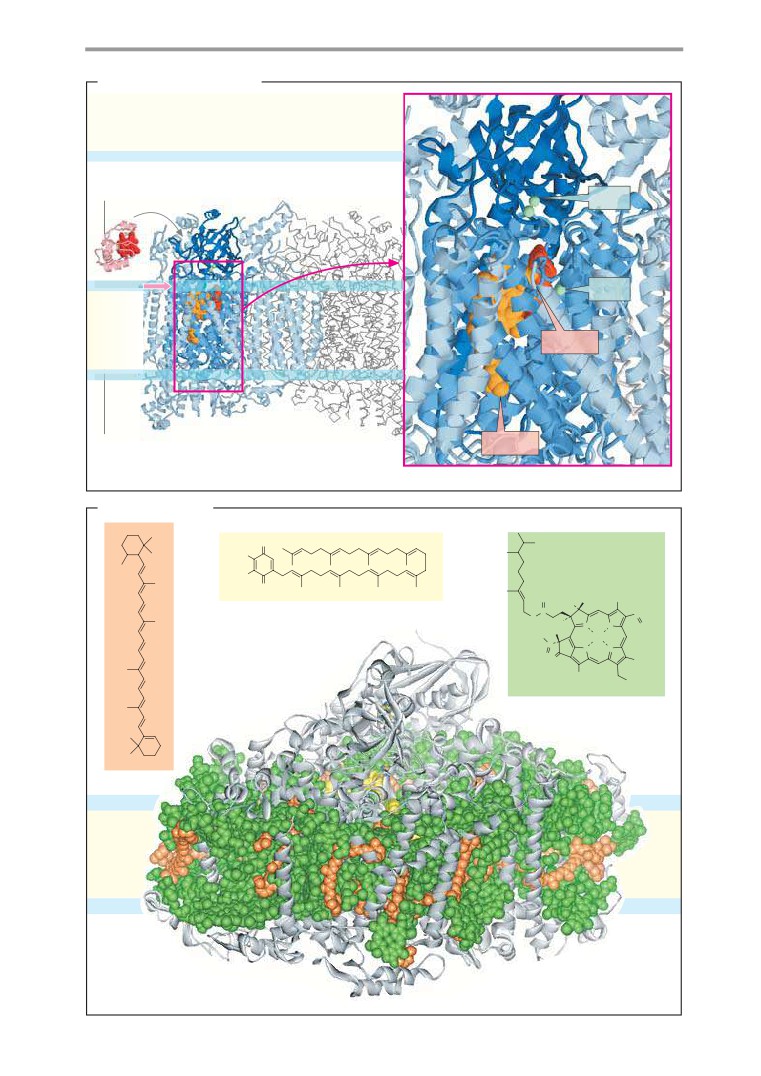
Molecular models: membrane proteins .
132
Amino Acids
Oxoacid dehydrogenases
134
Chemistry and properties
58
Tricarboxylic acid cycle: reactions
136
Proteinogenic amino acids
60
Tricarboxylic acid cycle: functions
138
Non-proteinogenic amino acids
62
Respiratory chain
140
Peptides and Proteins
ATP synthesis
142
Overview
64
Regulation
144
Peptide bonds
66
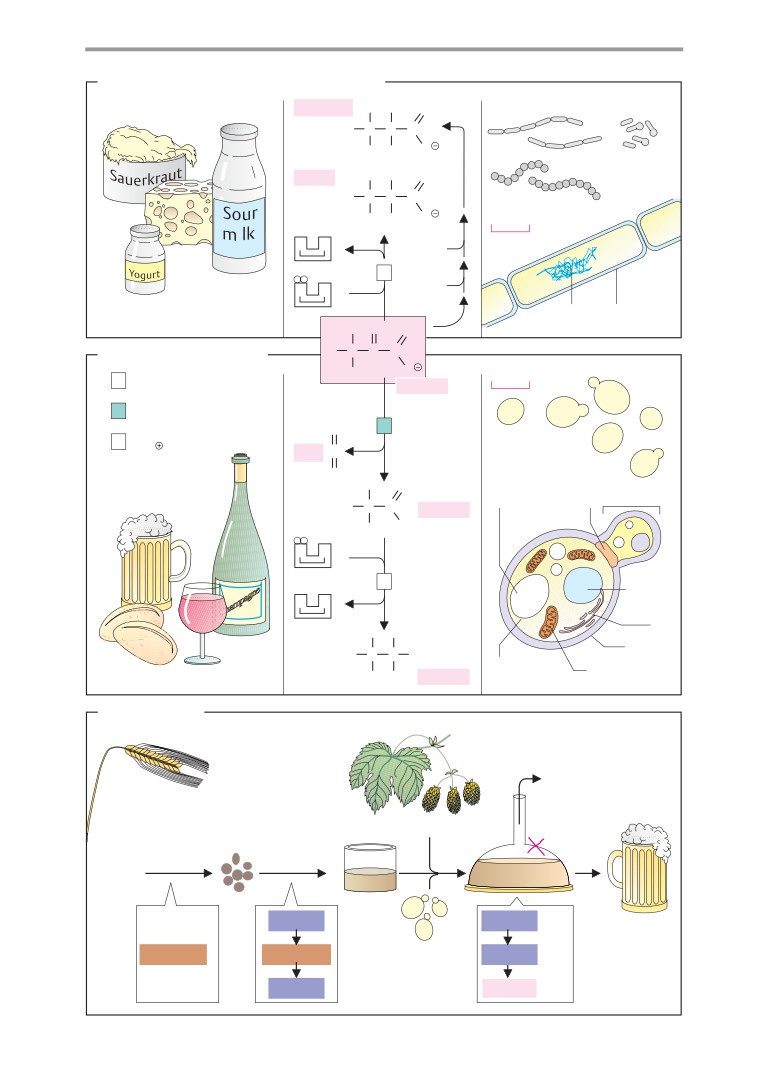
Respiration and fermentation
146
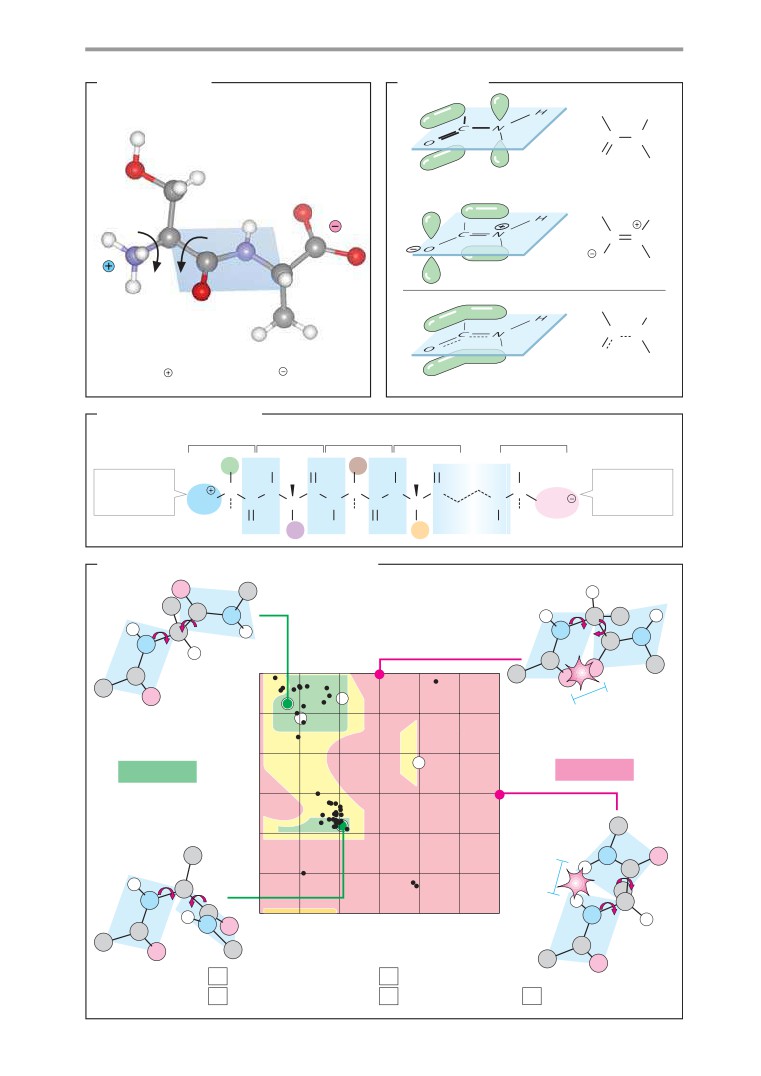
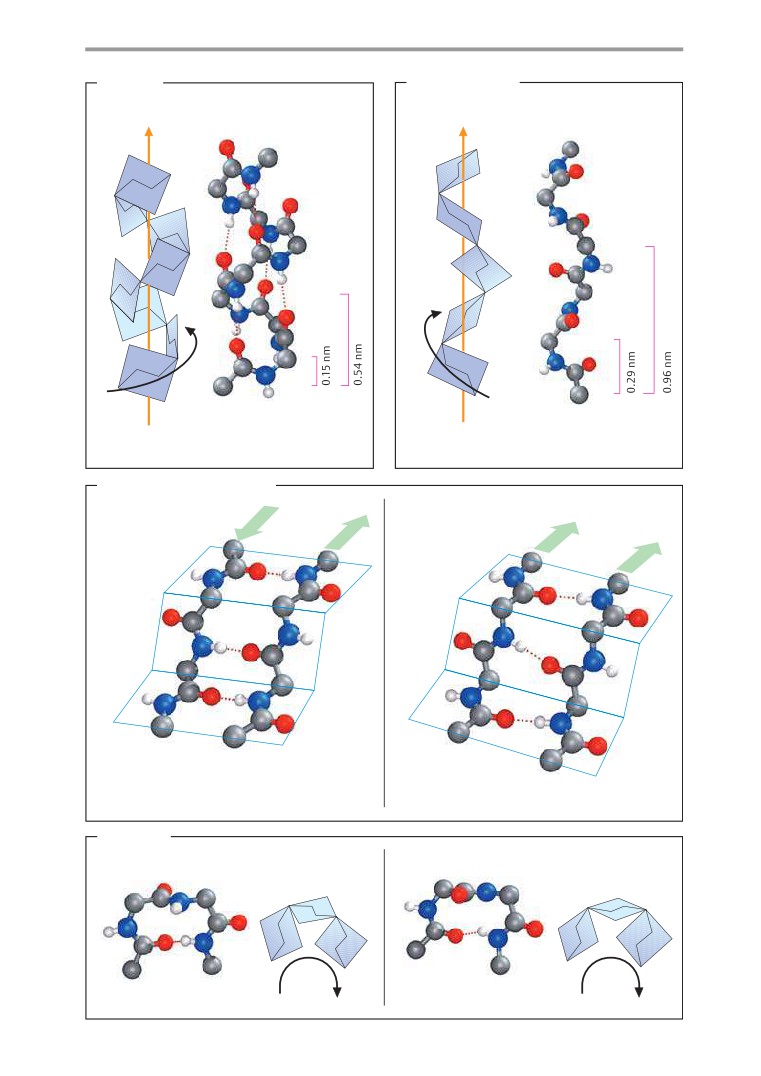
Secondary structures
68
Fermentations
148
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Contents
IX
Carbohydrate Metabolism
Endoplasmic Reticulum and Golgi Apparatus
Glycolysis
150
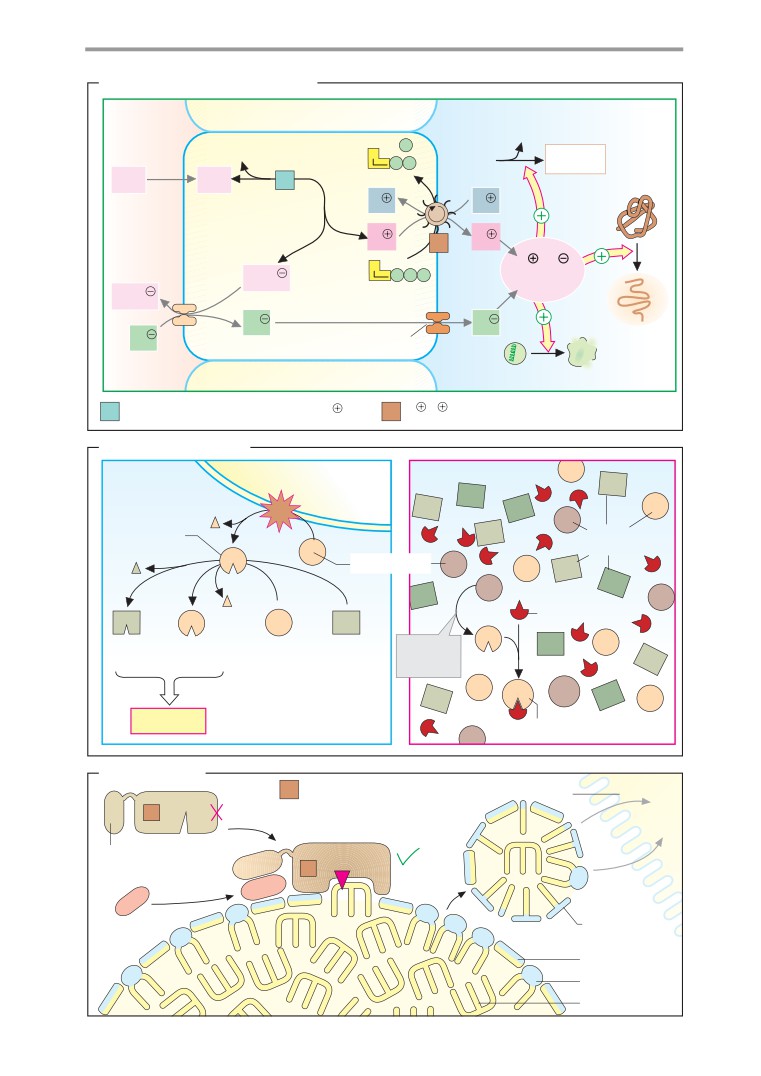
ER: structure and function
226
Pentose phosphate pathway
152
Protein sorting
228
Gluconeogenesis
154
Protein synthesis and maturation
230
Glycogen metabolism
156
Protein maturation
232
Regulation
158
Lysosomes
234
Diabetes mellitus
160
Lipid Metabolism
Molecular Genetics
Overview
162
Overview
236
Fatty acid degradation
164
Genome
238
Minor pathways of fatty acid
Replication
240
degradation
166
Transcription
242
Fatty acid synthesis
168
Transcriptional control
244
Biosynthesis of complex lipids
170
RNA maturation
246
Biosynthesis of cholesterol
172
Amino acid activation
248
Protein Metabolism
Translation I: initiation
250
Protein metabolism: overview
174
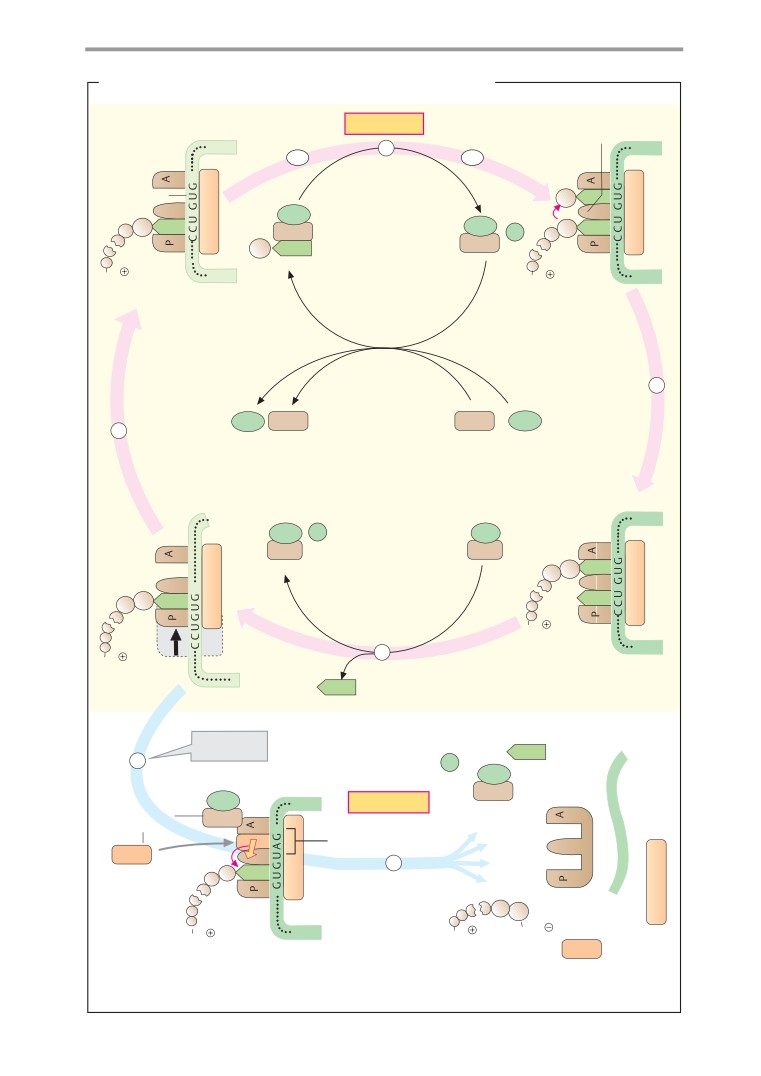
Translation II: elongation and
Proteolysis
176
termination
252
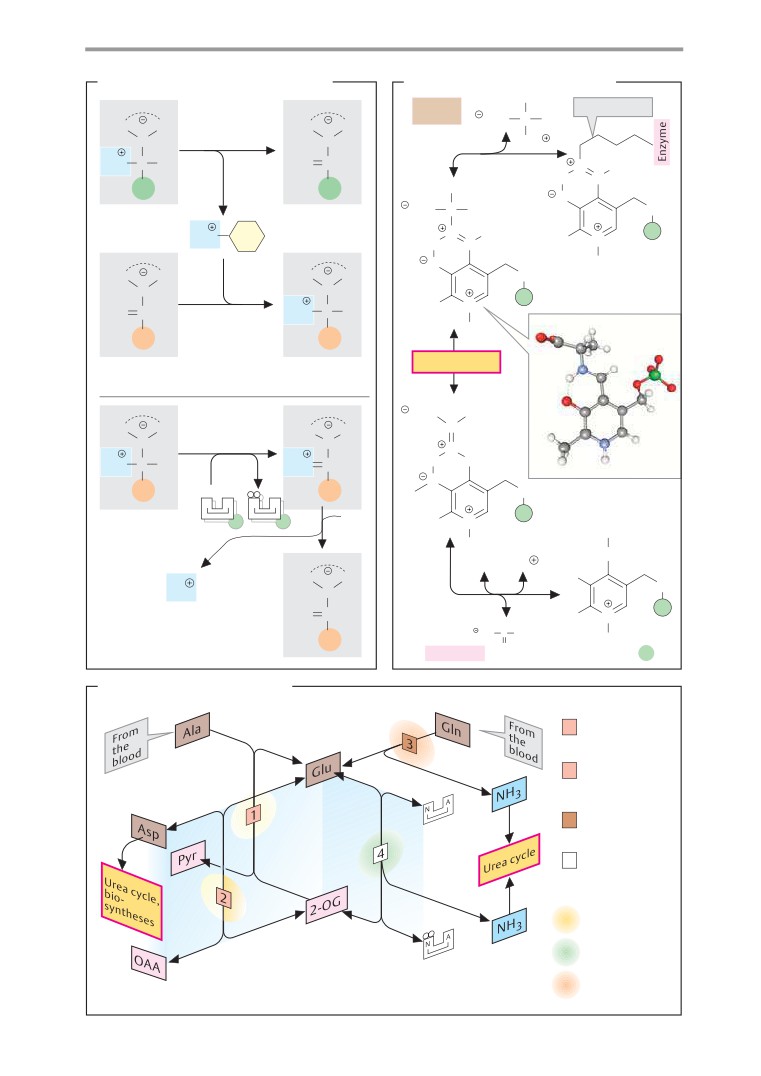
Transamination and deamination
178
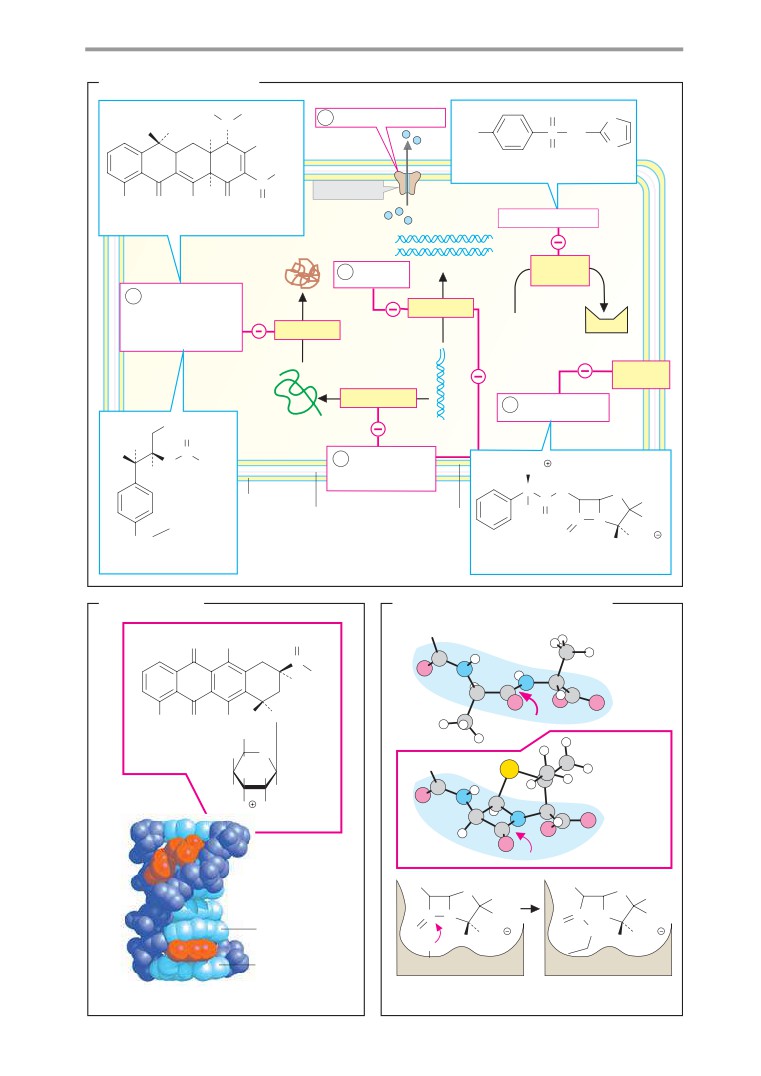
Antibiotics
254
Amino acid degradation
180
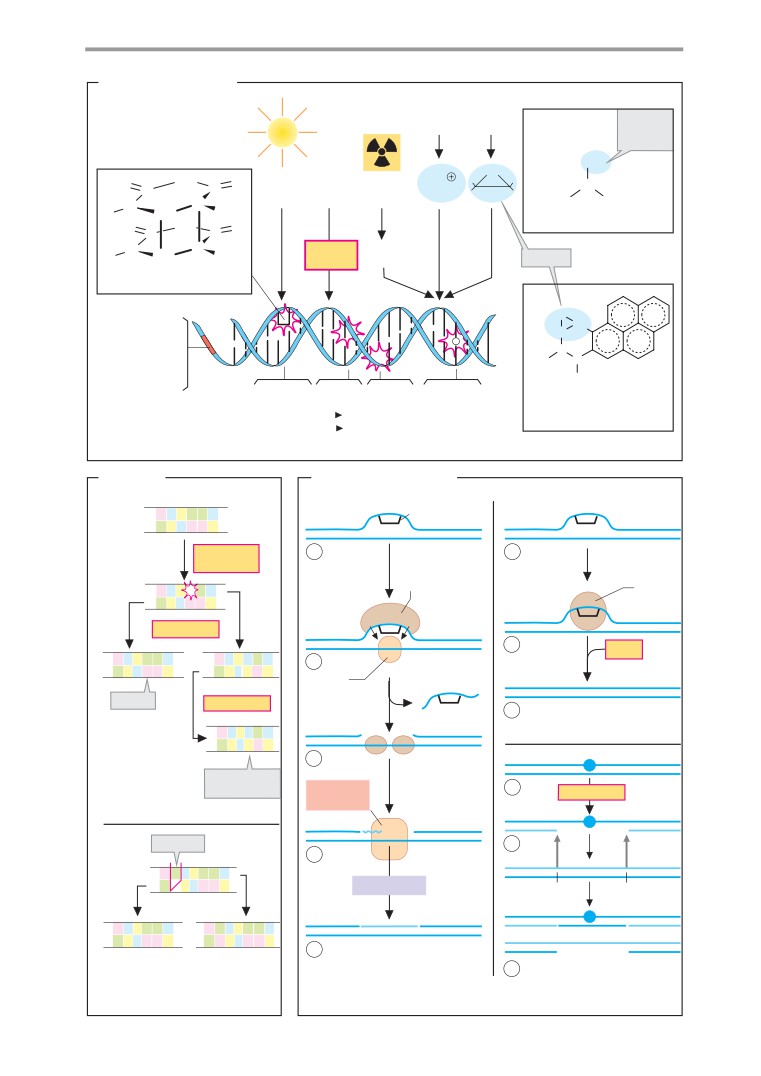
Mutation and repair
256
Urea cycle
182
Genetic engineering
Amino acid biosynthesis
184
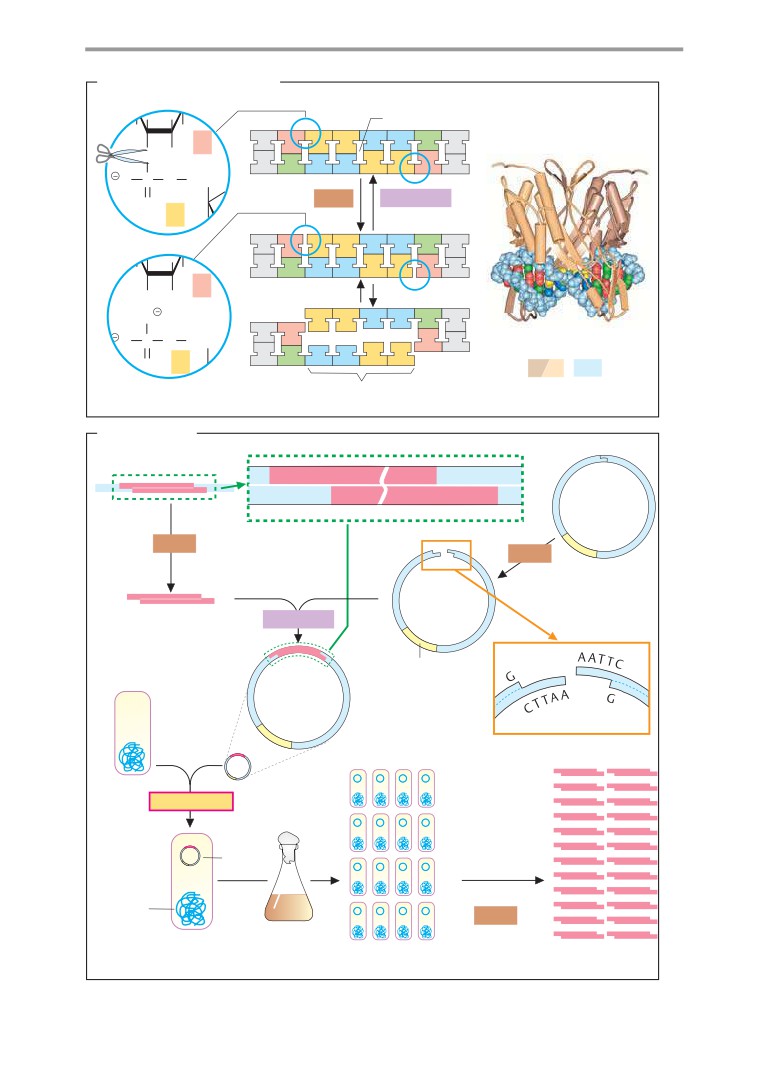
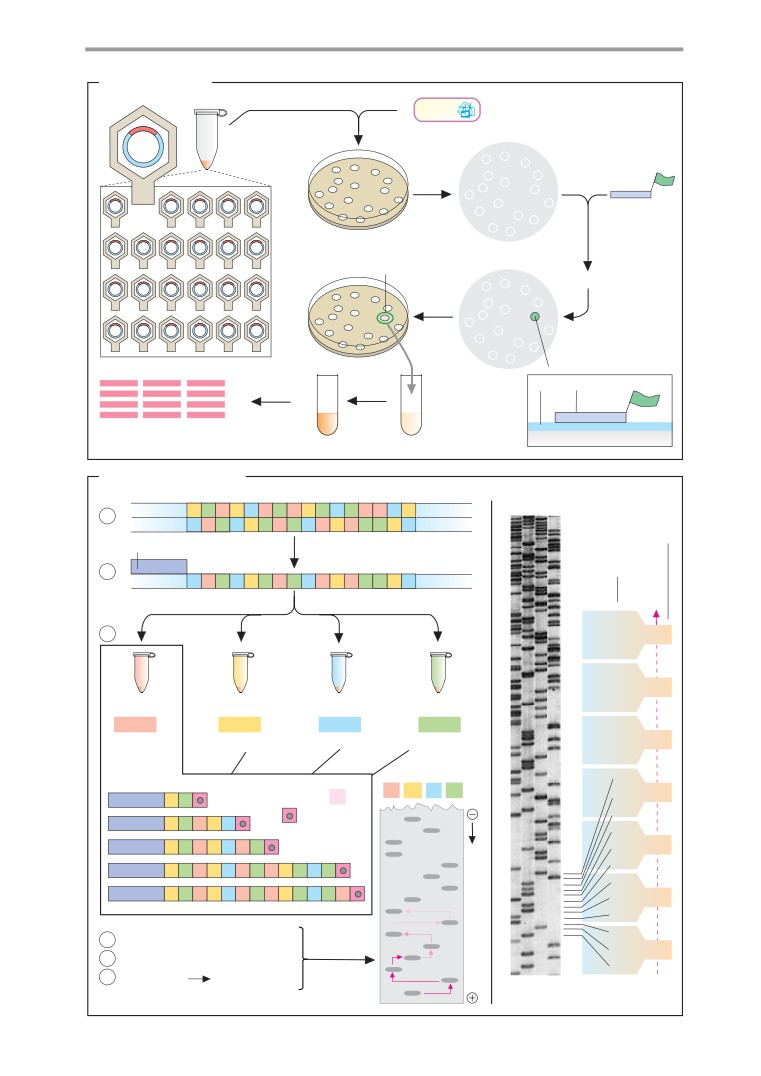
DNA cloning
258
Nucleotide Metabolism
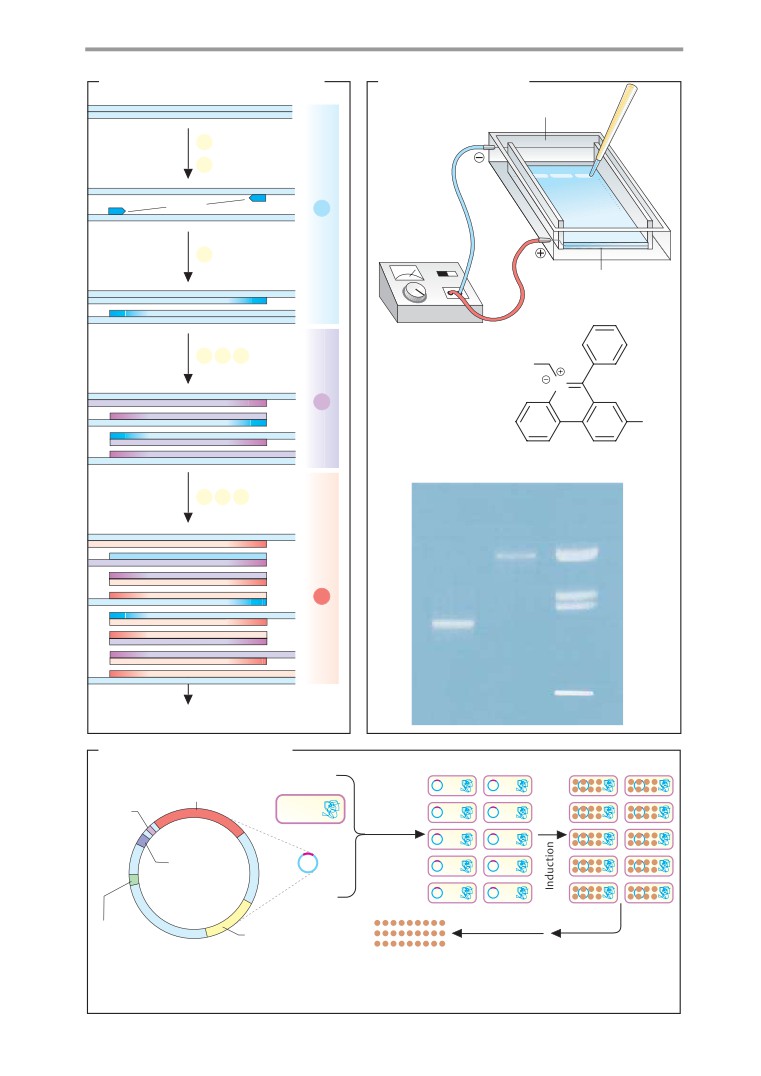
DNA sequencing
260
Nucleotide degradation
186
PCR and protein expression
262
Purine and pyrimidine biosynthesis
188
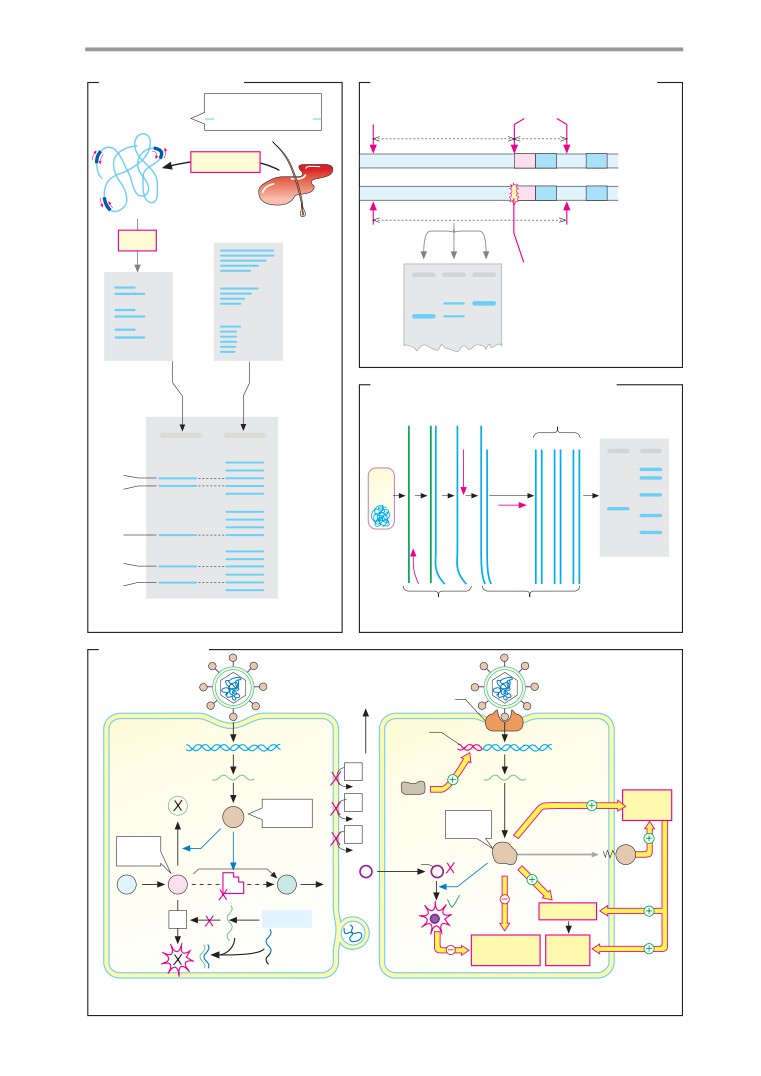
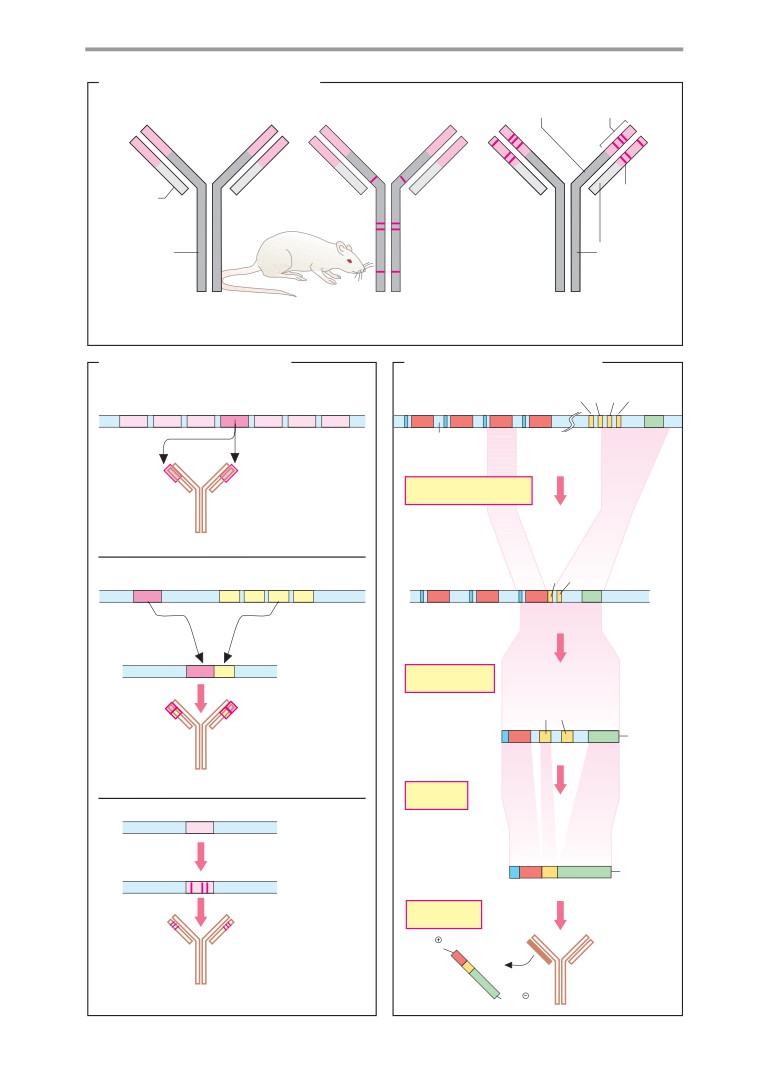
Genetic engineering in medicine
264
Nucleotide biosynthesis
190
Tissues and organs
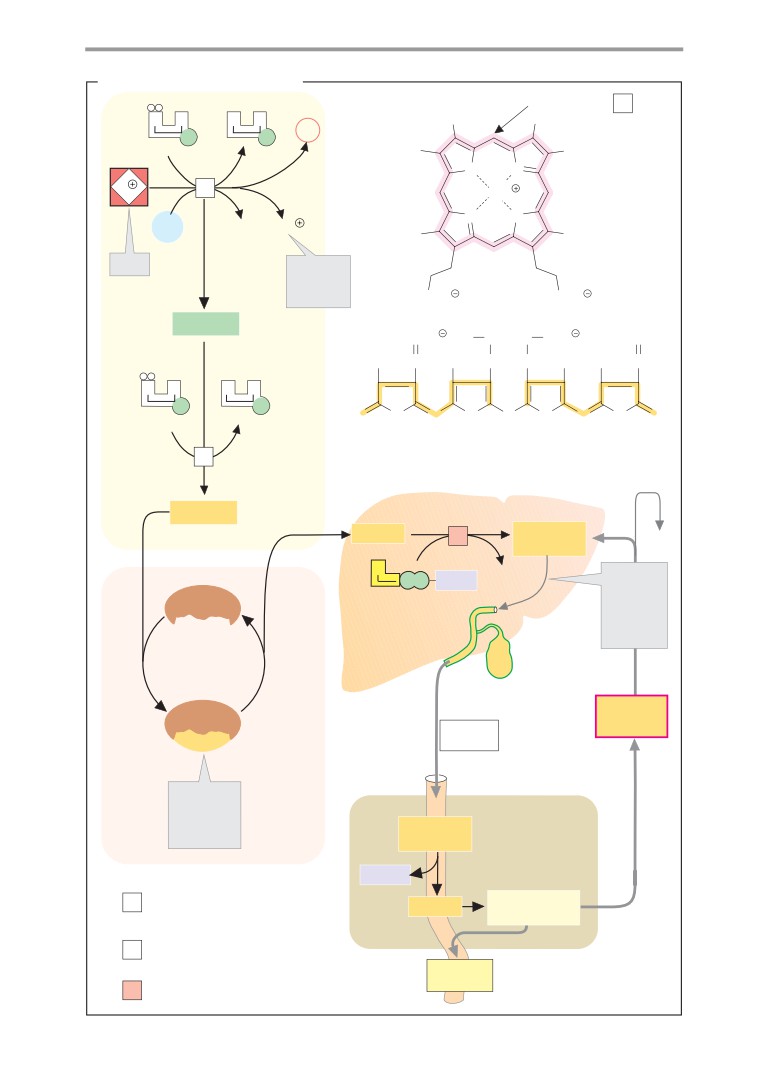
Porphyrin Metabolism
Heme biosynthesis
192
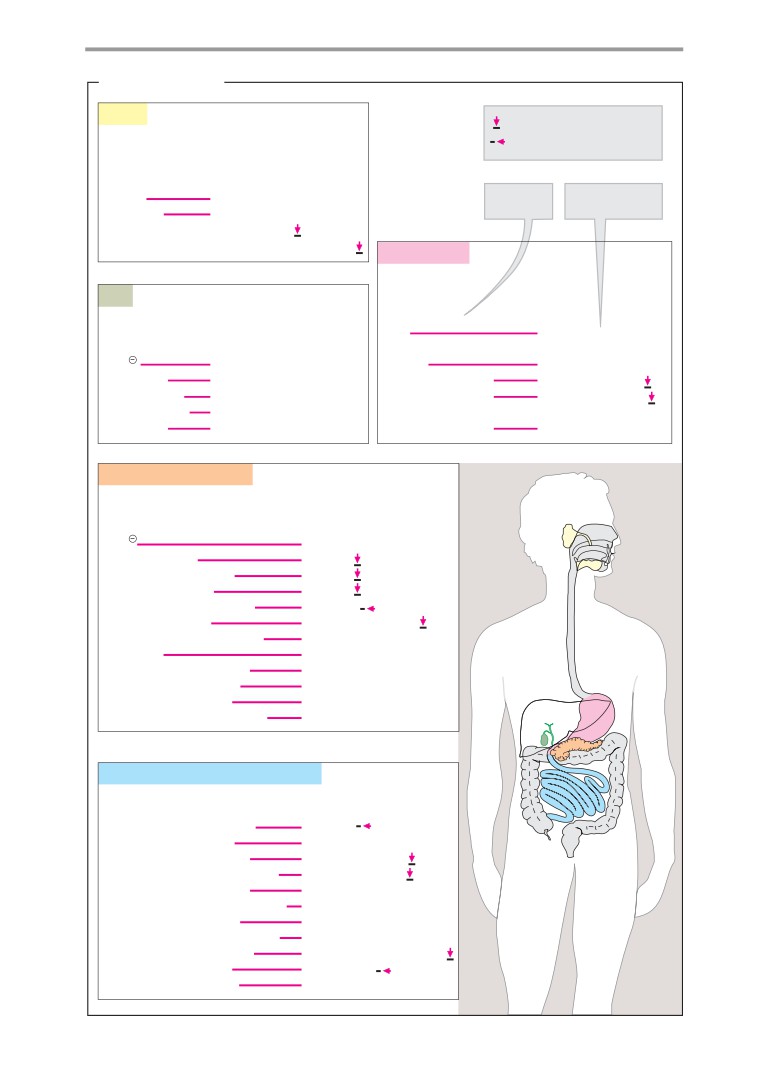
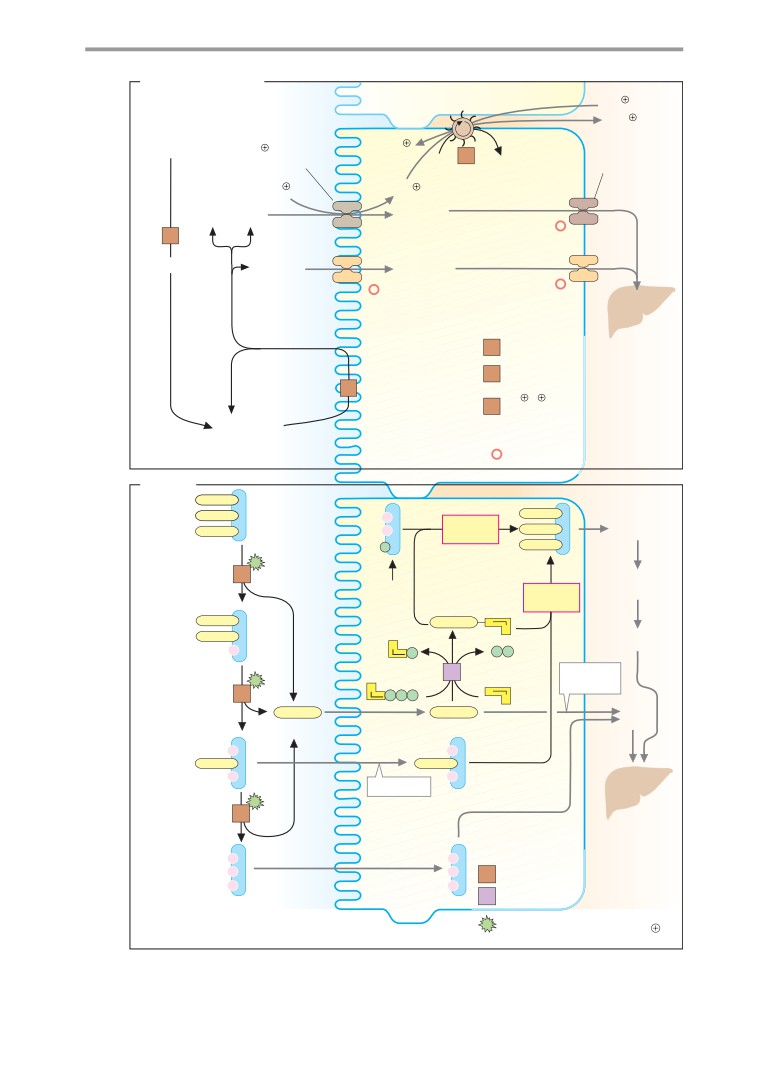
Digestion
Heme degradation
194
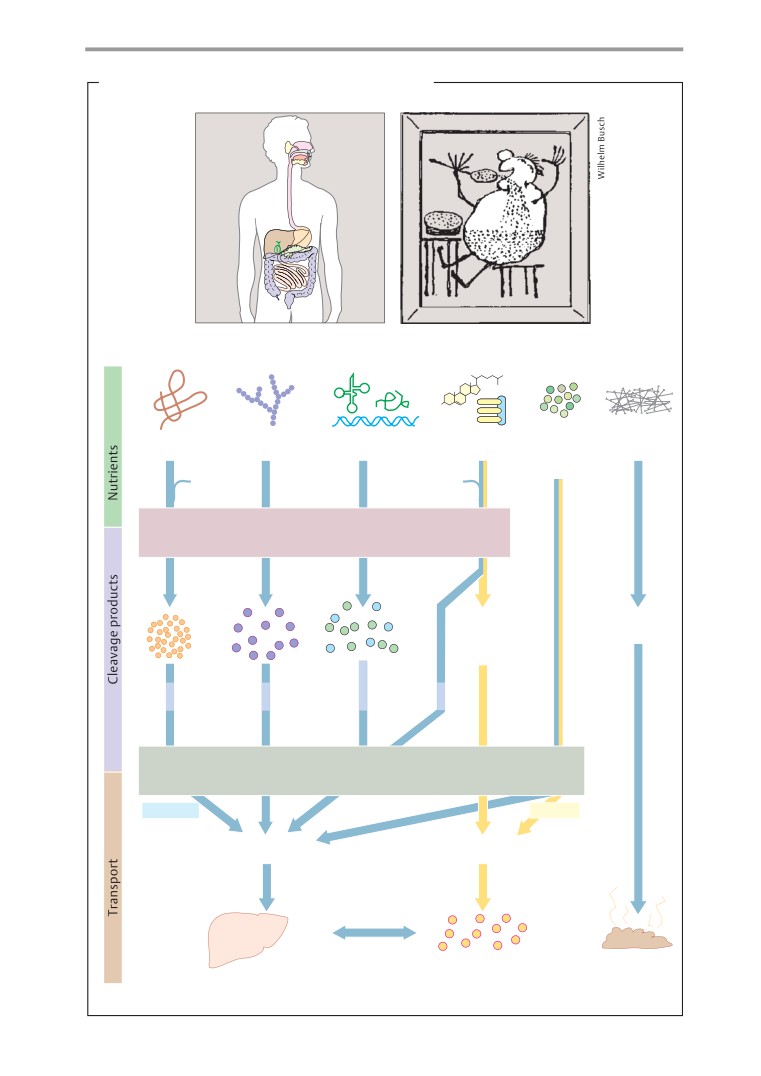
Overview
266
Digestive secretions
268
Organelles
Digestive processes
270
Resorption
272
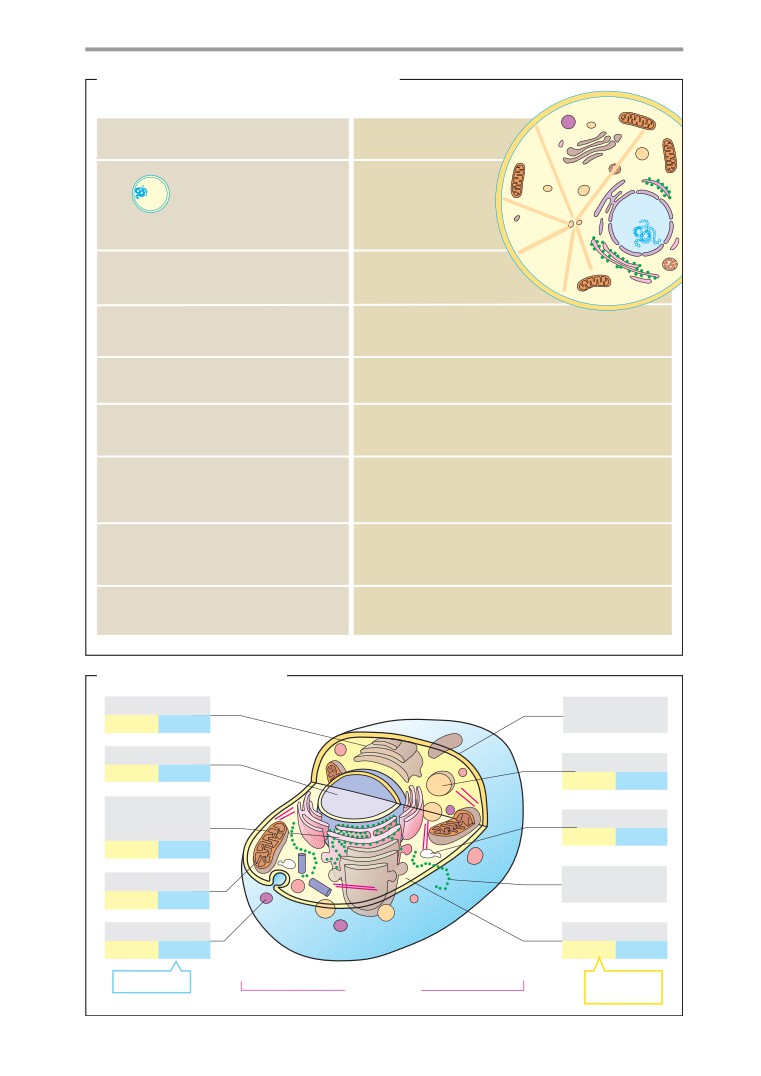
Basics
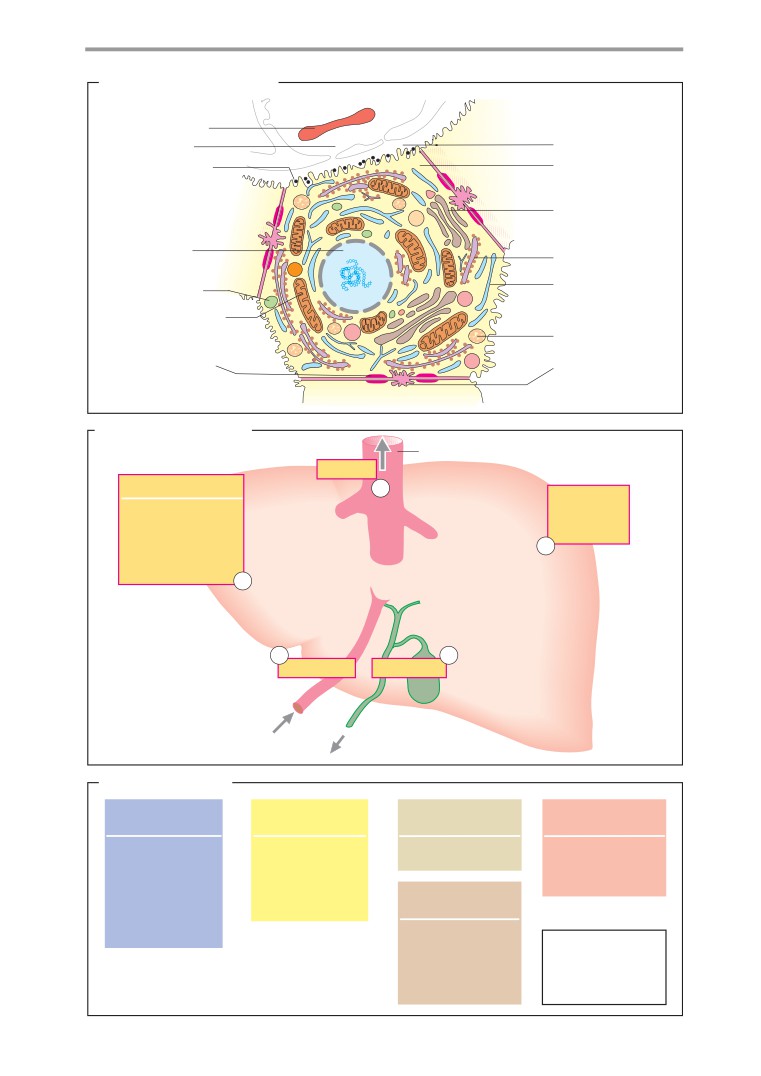
Structure of cells
196
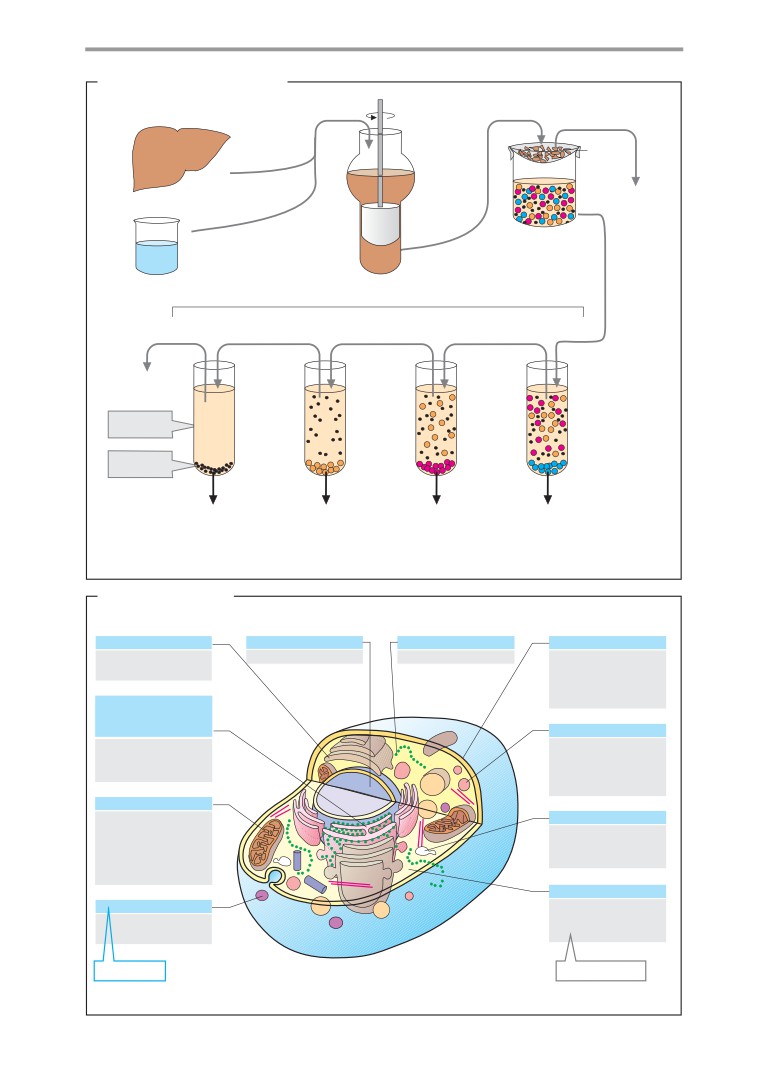
Blood
Cell fractionation
198
Composition and functions
274
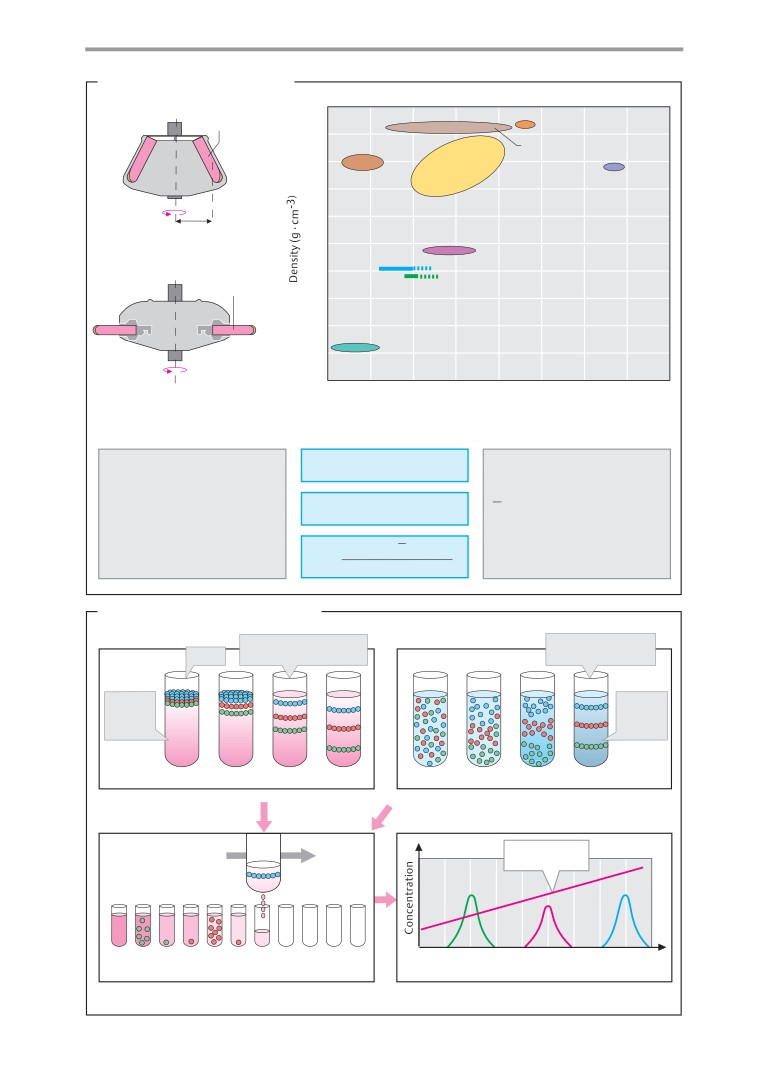
Centrifugation
200
Plasma proteins
276
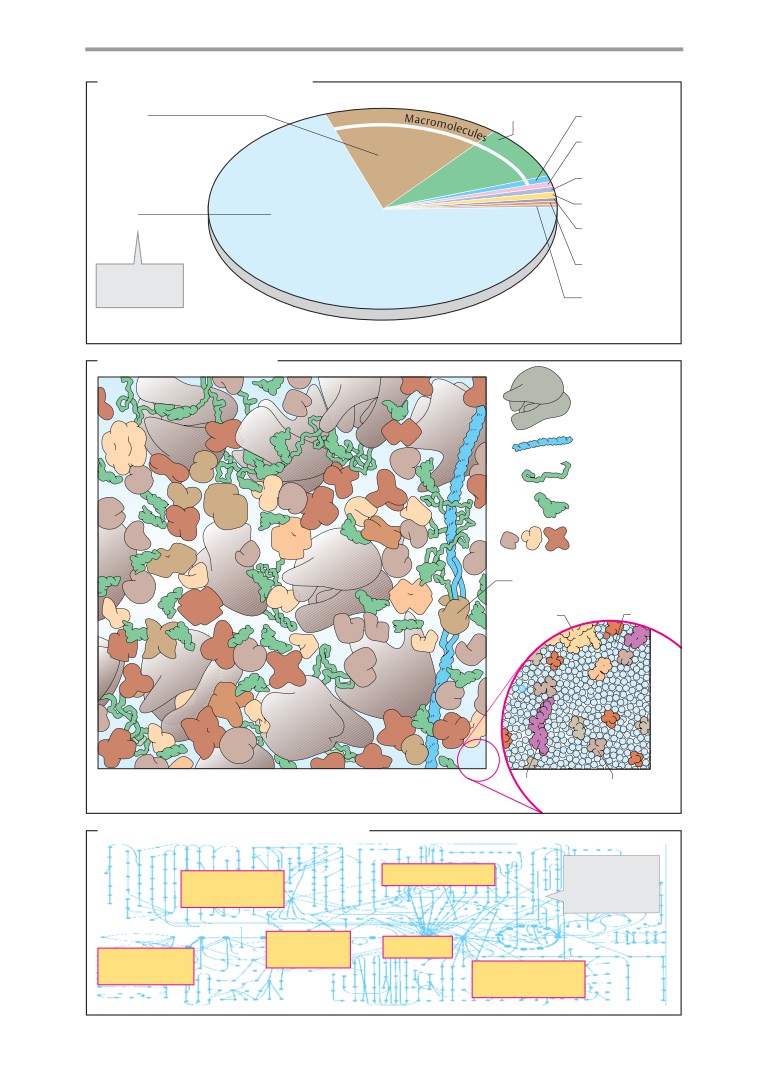
Cell components and cytoplasm
202
Lipoproteins
278
Hemoglobin
280
Cytoskeleton
Gas transport
282
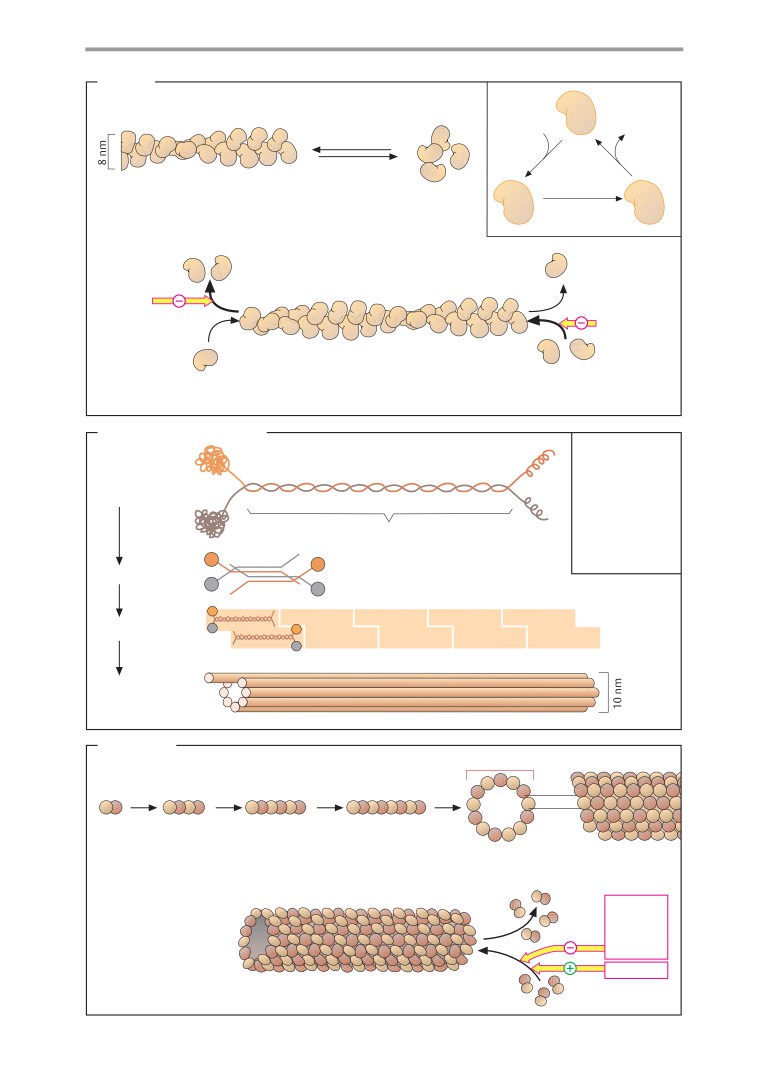
Components
204
Erythrocyte metabolism
284
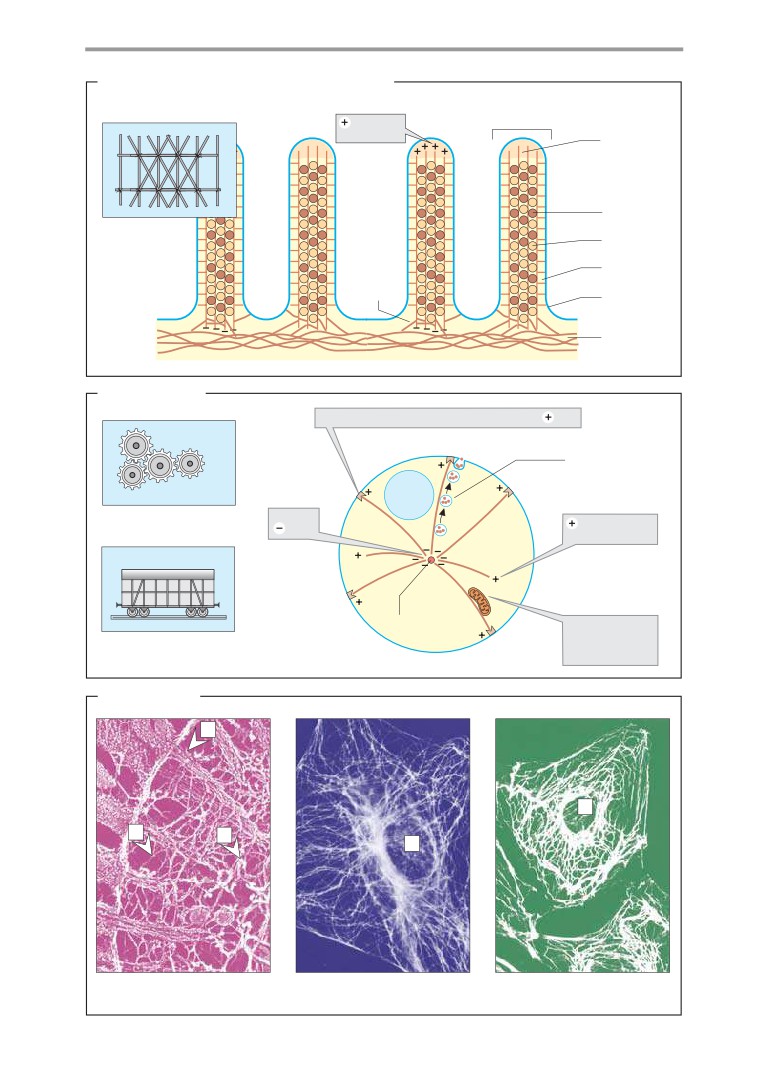
Structure and functions
206
Iron metabolism
286
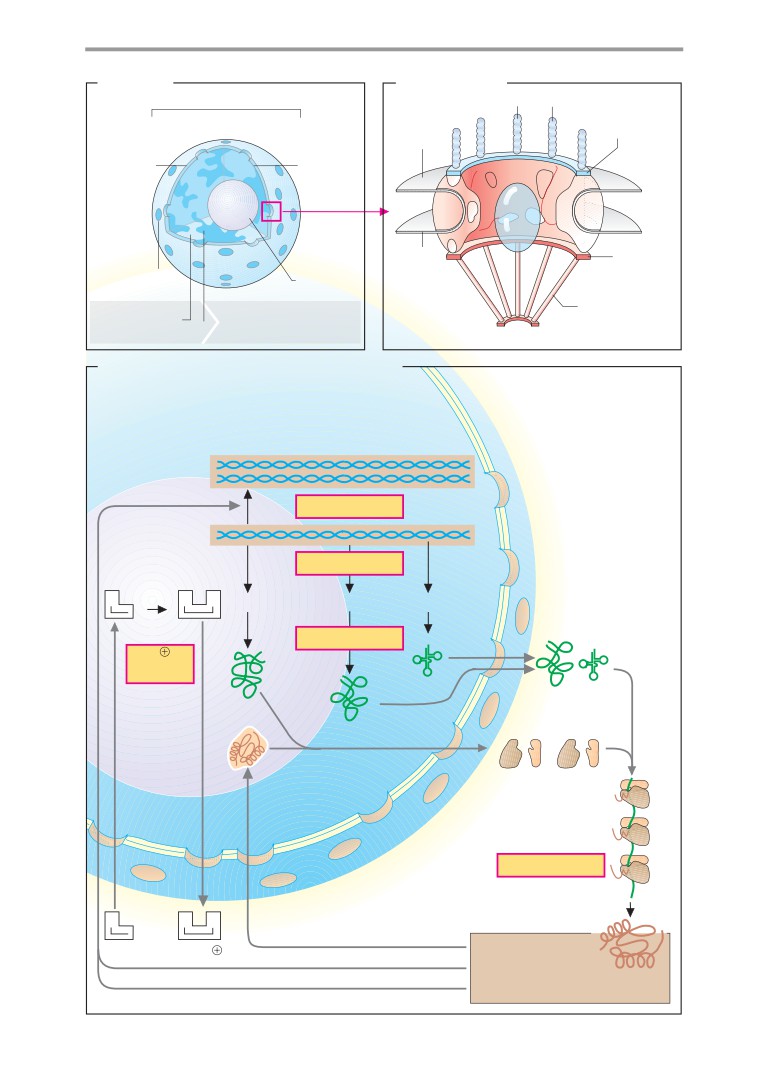
Nucleus
208
Acid-base balance
288
Mitochondria
Blood clotting
290
Structure and functions
210
Fibrinolysis, blood groups
292
Transport systems
212
Immune system
Biological Membranes
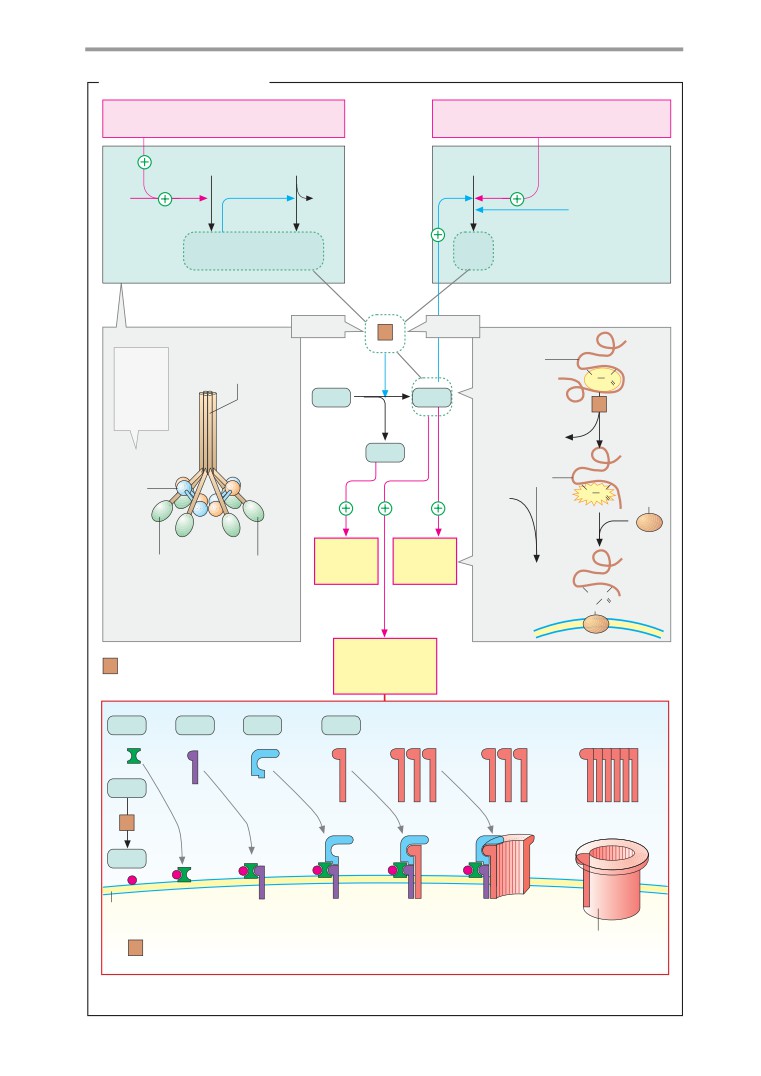
Immune response
294
Structure and components
214
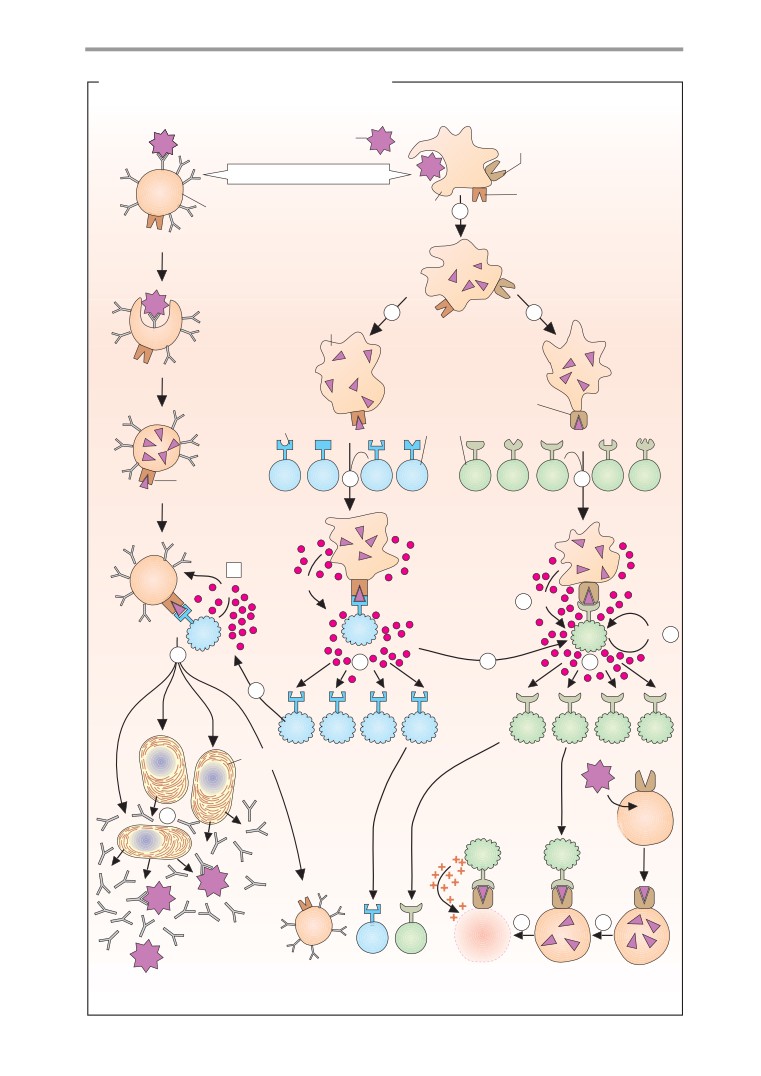
T-cell activation
296
Functions and composition
216
Complement system
298
Transport processes
218
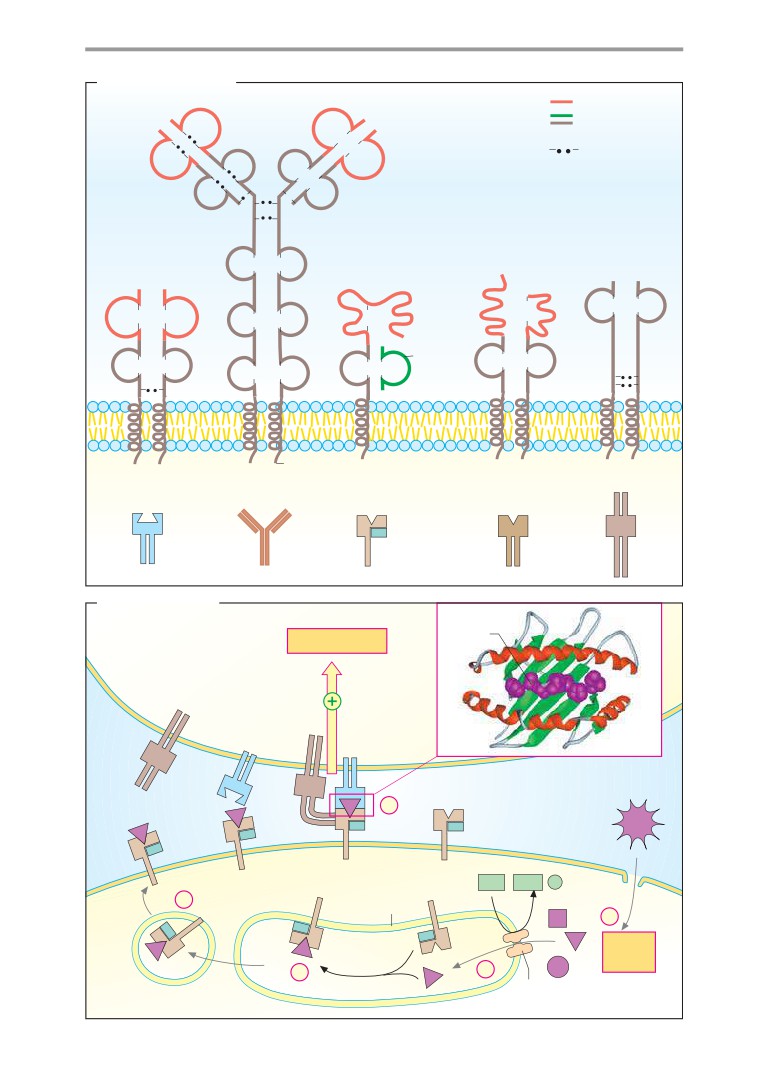
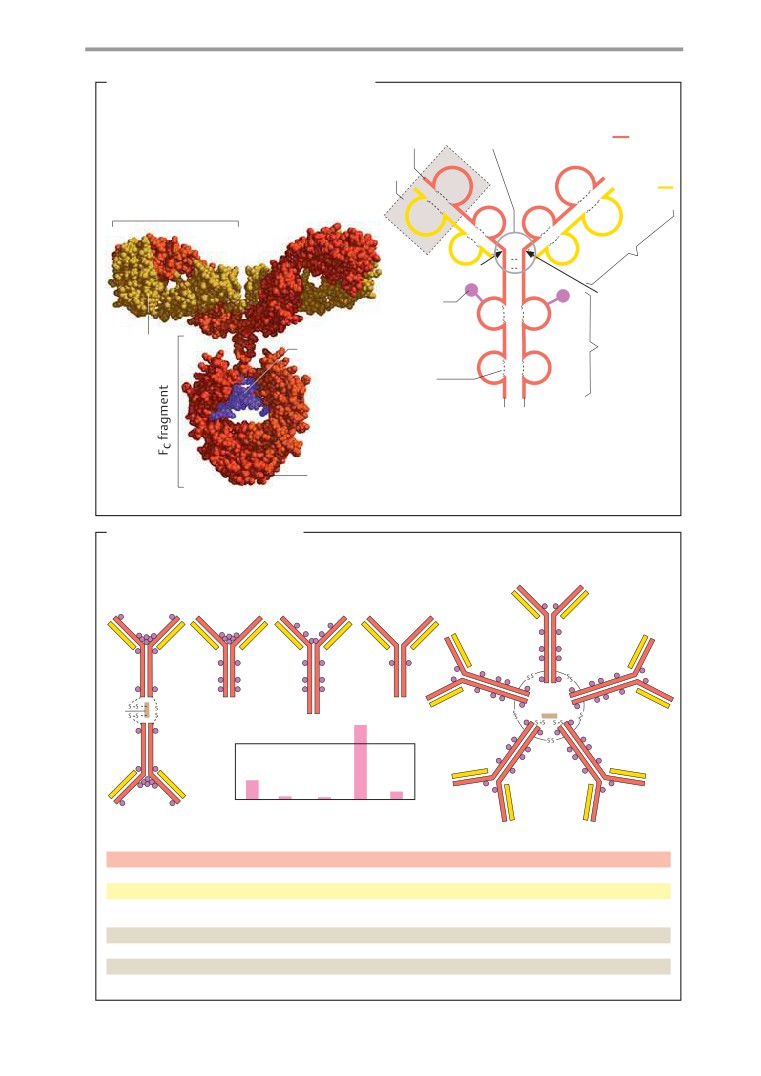
Antibodies
300
Transport proteins
220
Antibody biosynthesis
302
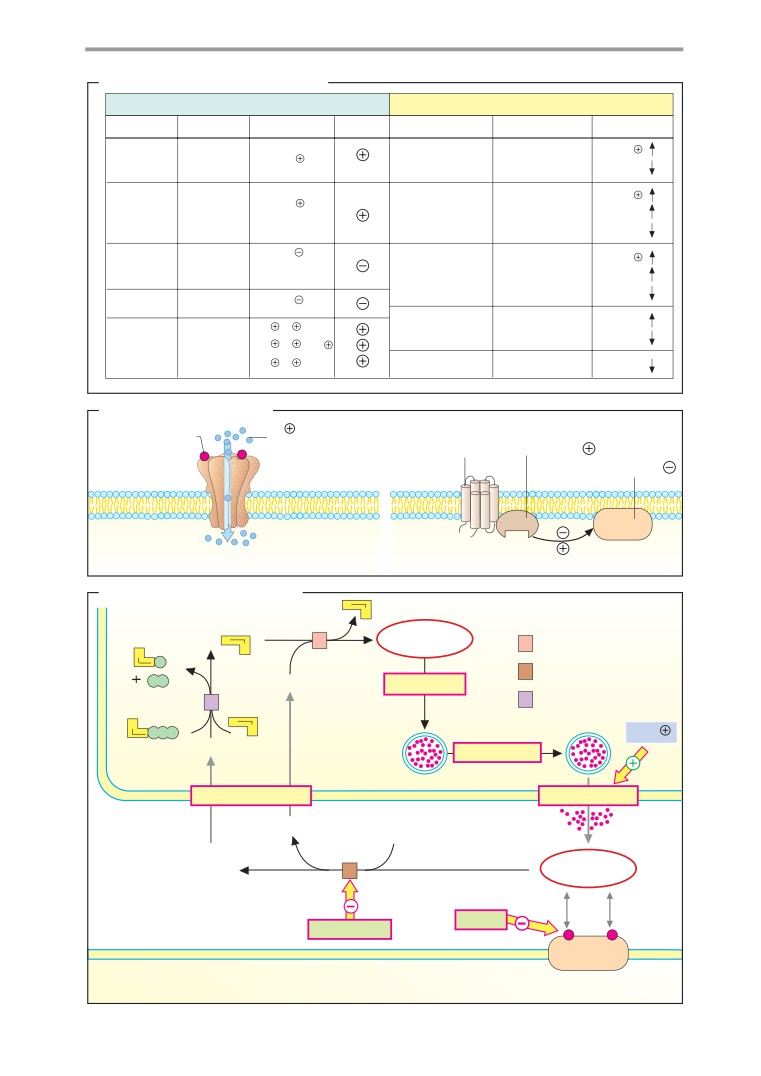
Ion channels
222
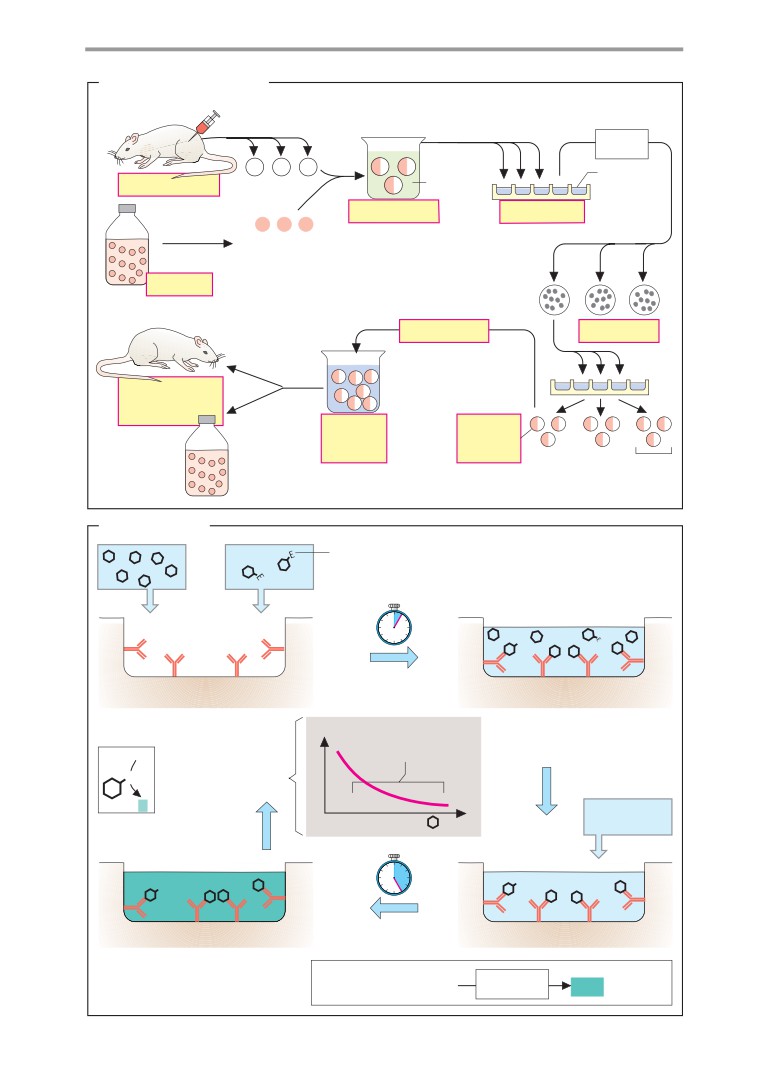
Monoclonal antibodies, immunoassay .
304
Membrane receptors
224
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
X
Contents
Liver
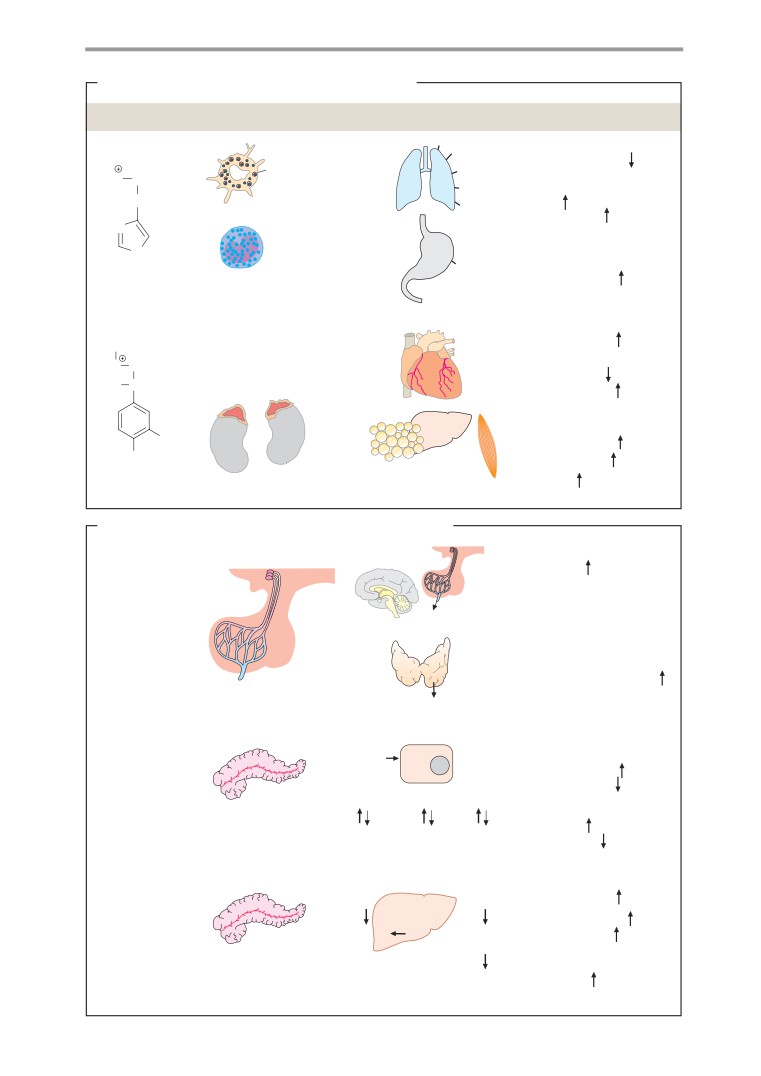
Hydrophilic hormones
380
Functions
306
Metabolism of peptide hormones
382
Buffer function in organ metabolism
308
Mechanisms of action
384
Carbohydrate metabolism
310
Second messengers
386
Lipid metabolism
312
Signal cascades
388
Bile acids
314
Other signaling substances
Biotransformations
316
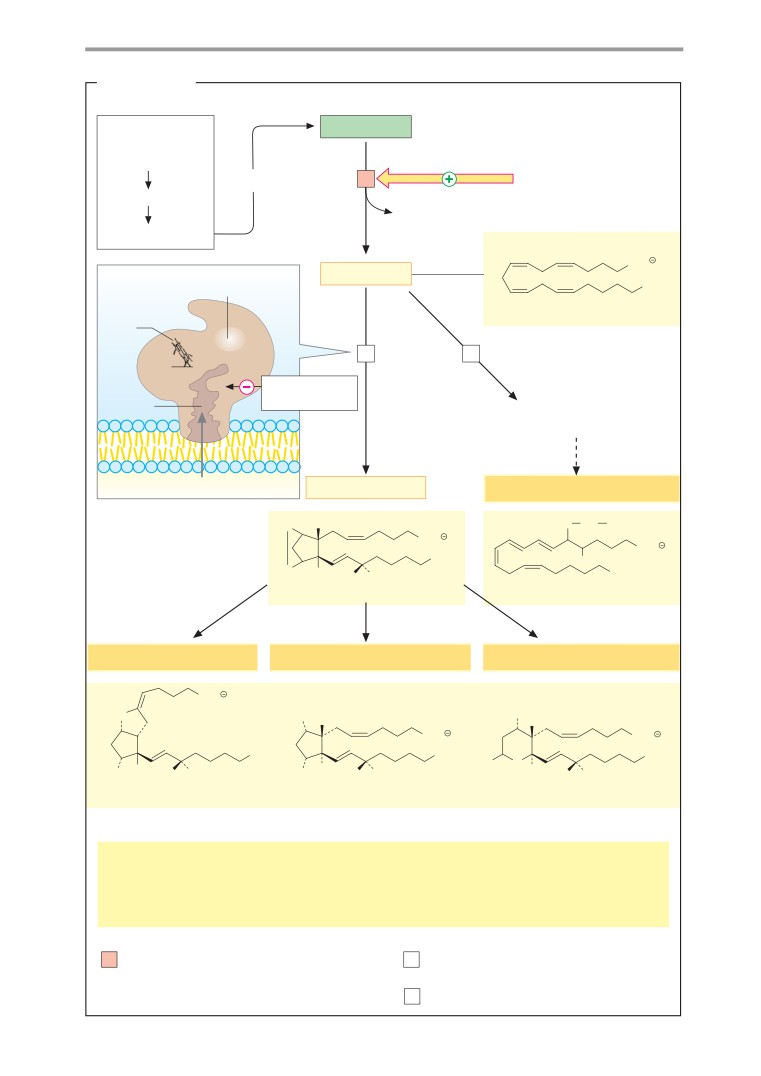
Eicosanoids
390
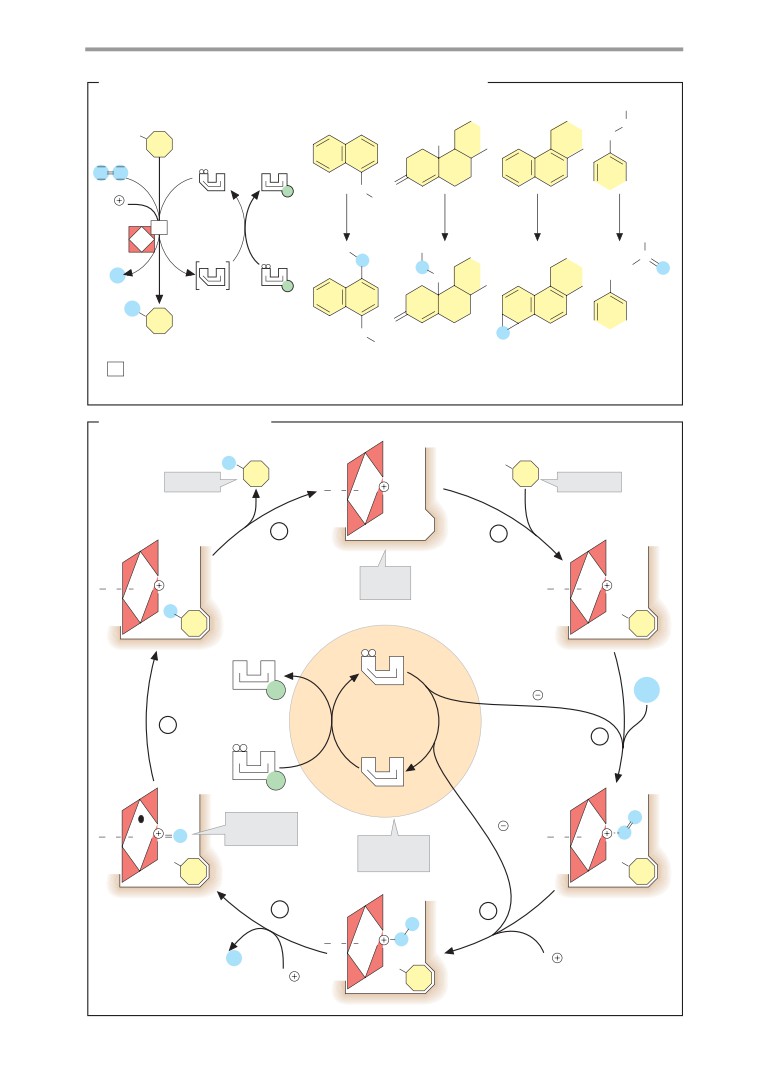
Cytochrome P450 systems
318
Cytokines
392
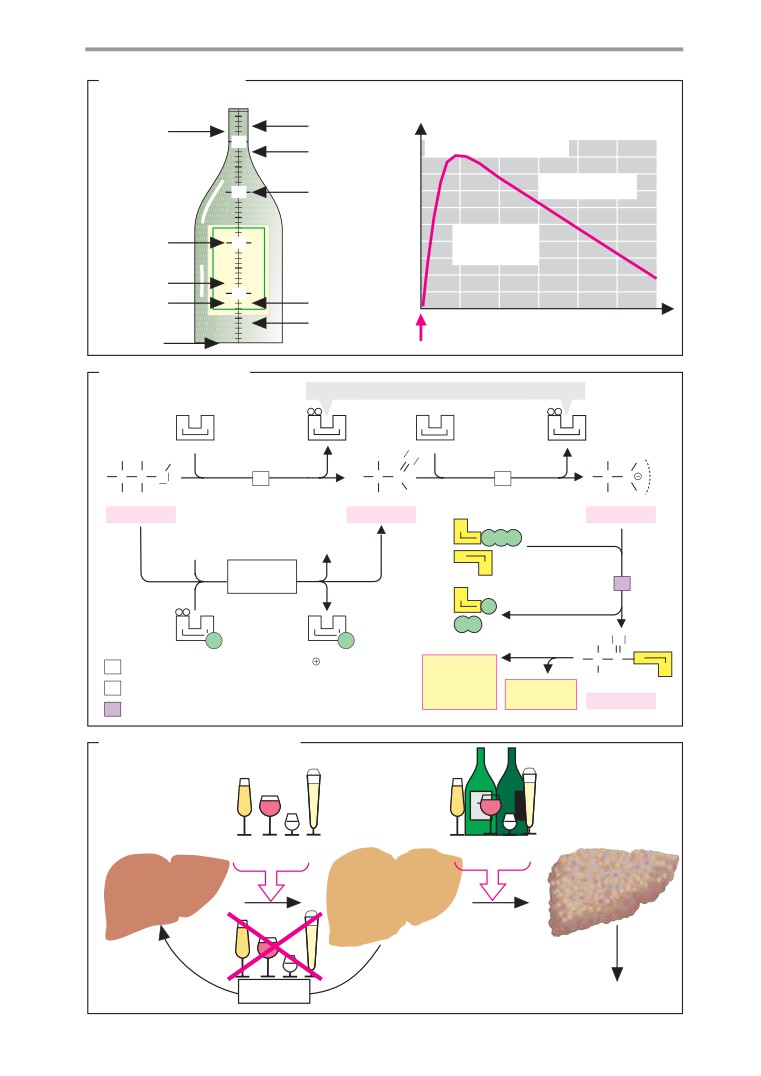
Ethanol metabolism
320
Kidney
Growth and development
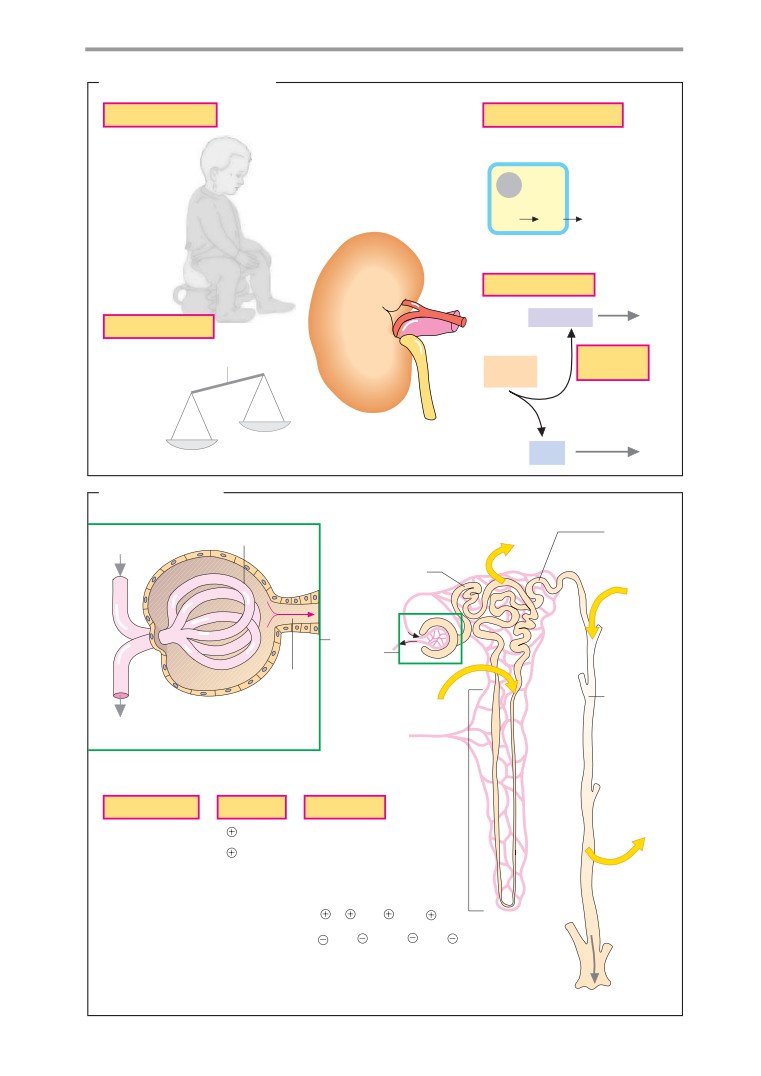
Functions
322
Cell proliferation
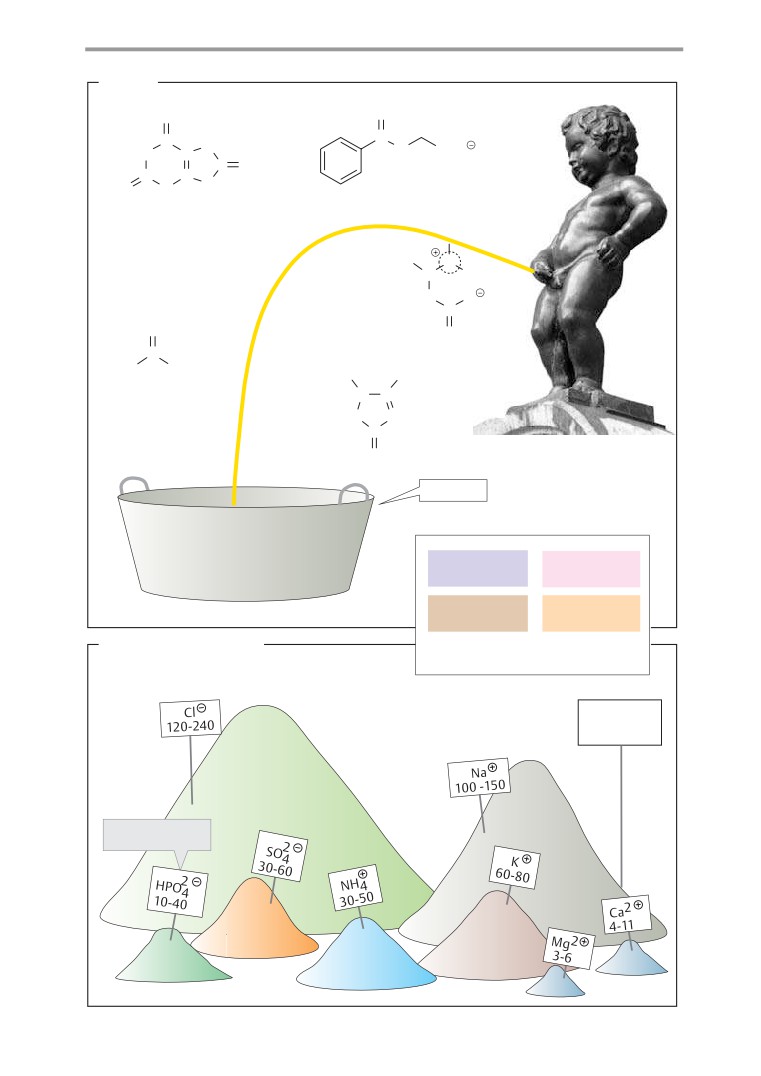
Urine
324
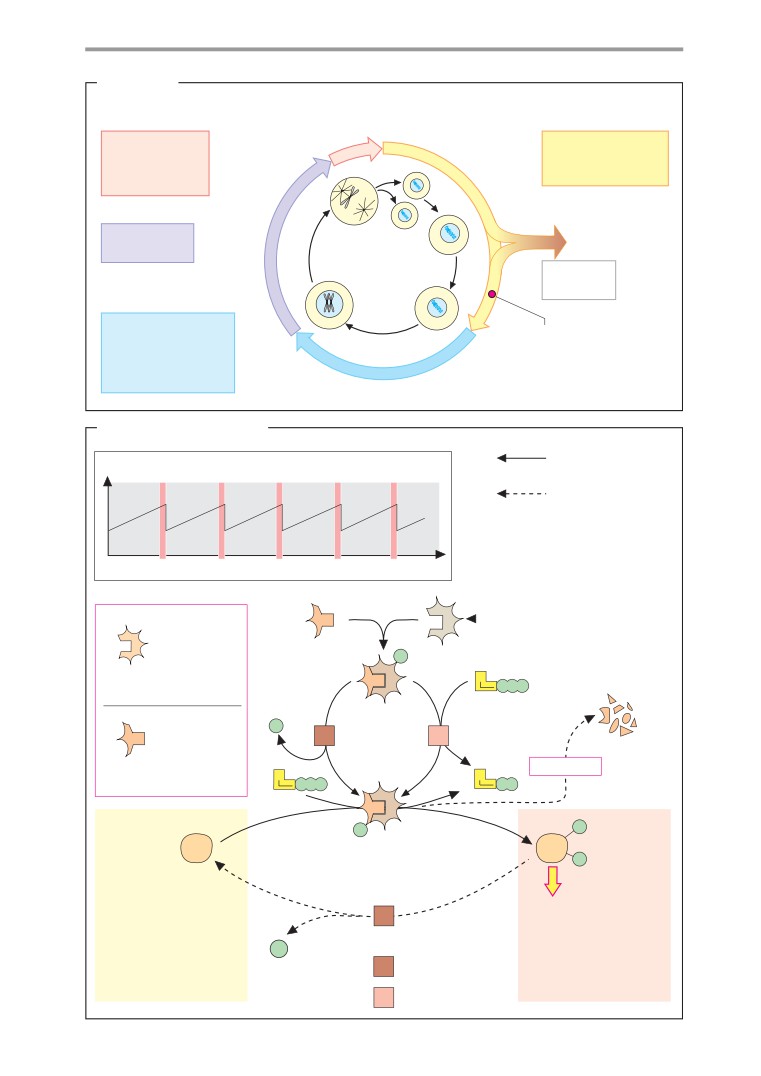
Cell cycle
394
Functions in the acid-base balance
326
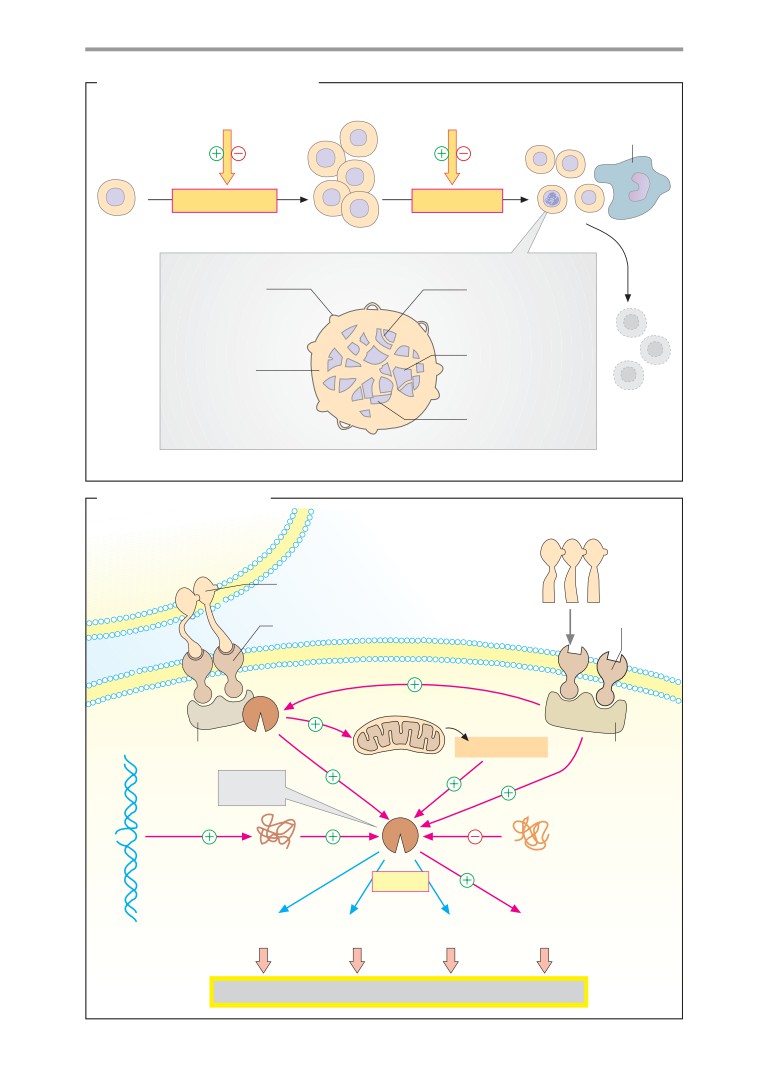
Apoptosis
396
Electrolyte and water recycling
328
Oncogenes
398
Renal hormones
330
Tumors
400
Muscle
Cytostatic drugs
402
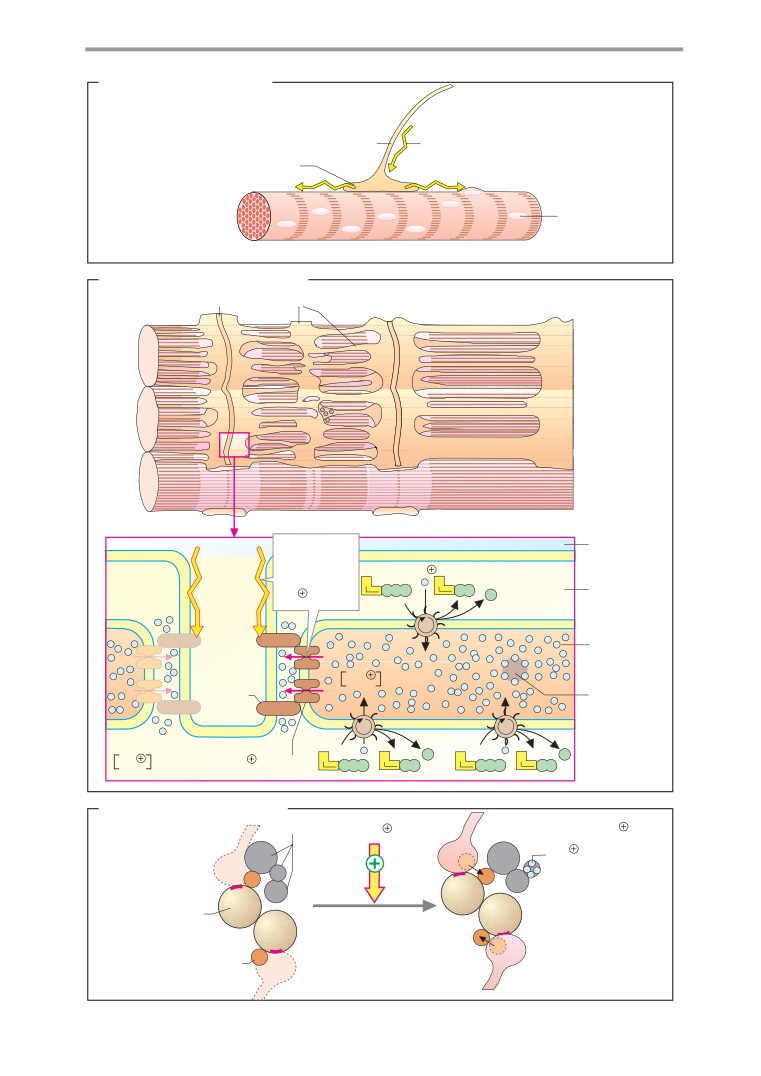
Muscle contraction
332
Viruses
404
Control of muscle contraction
334
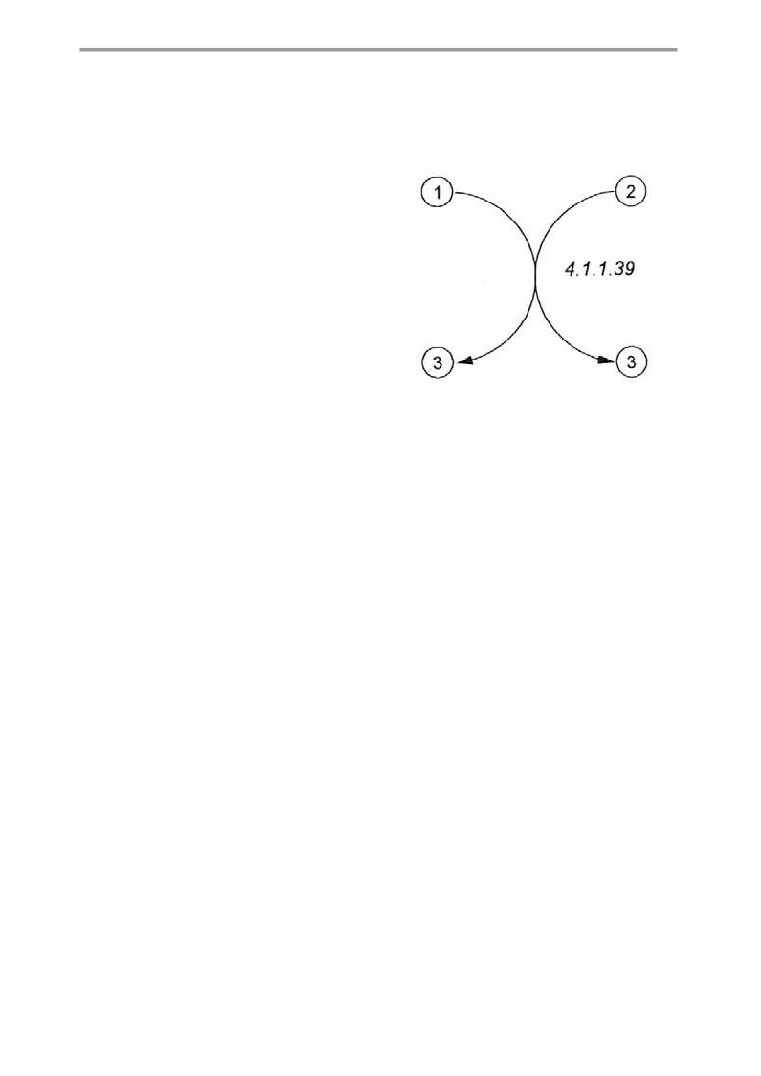
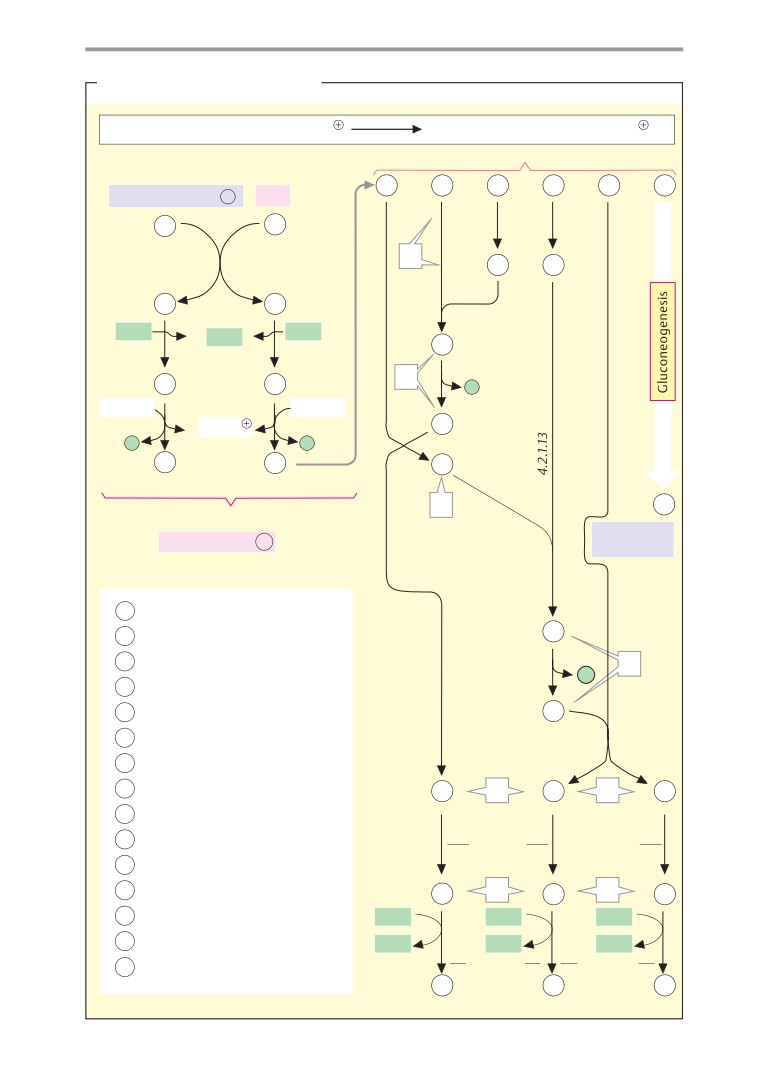
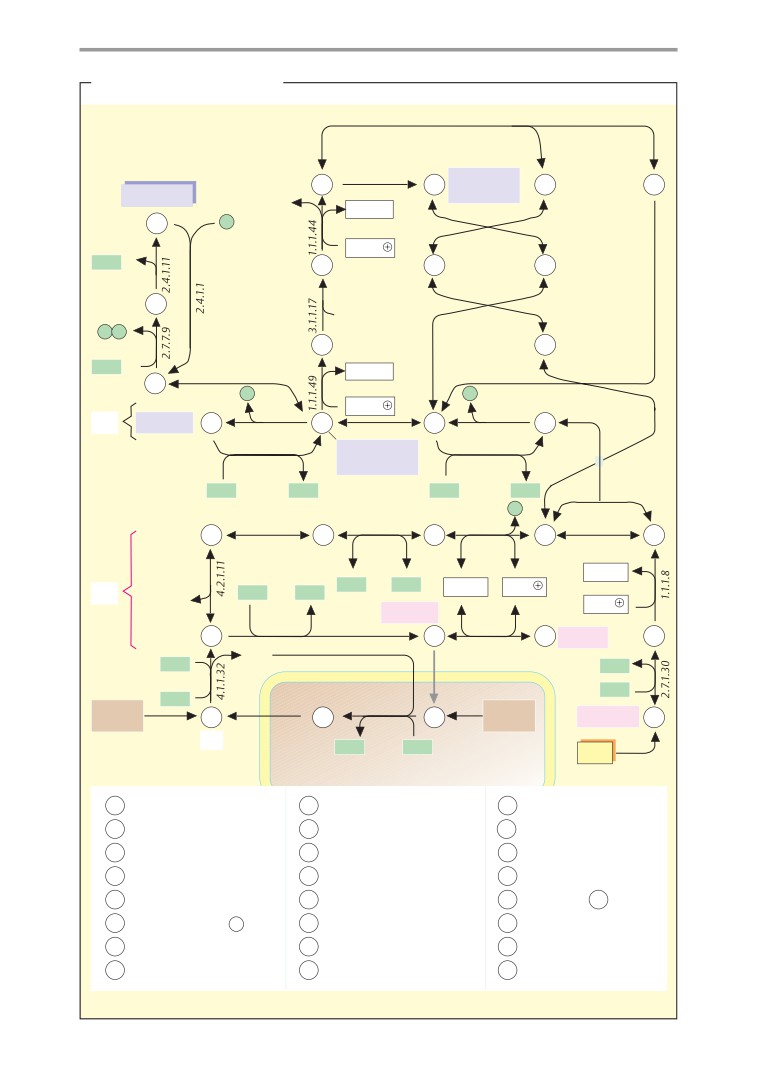
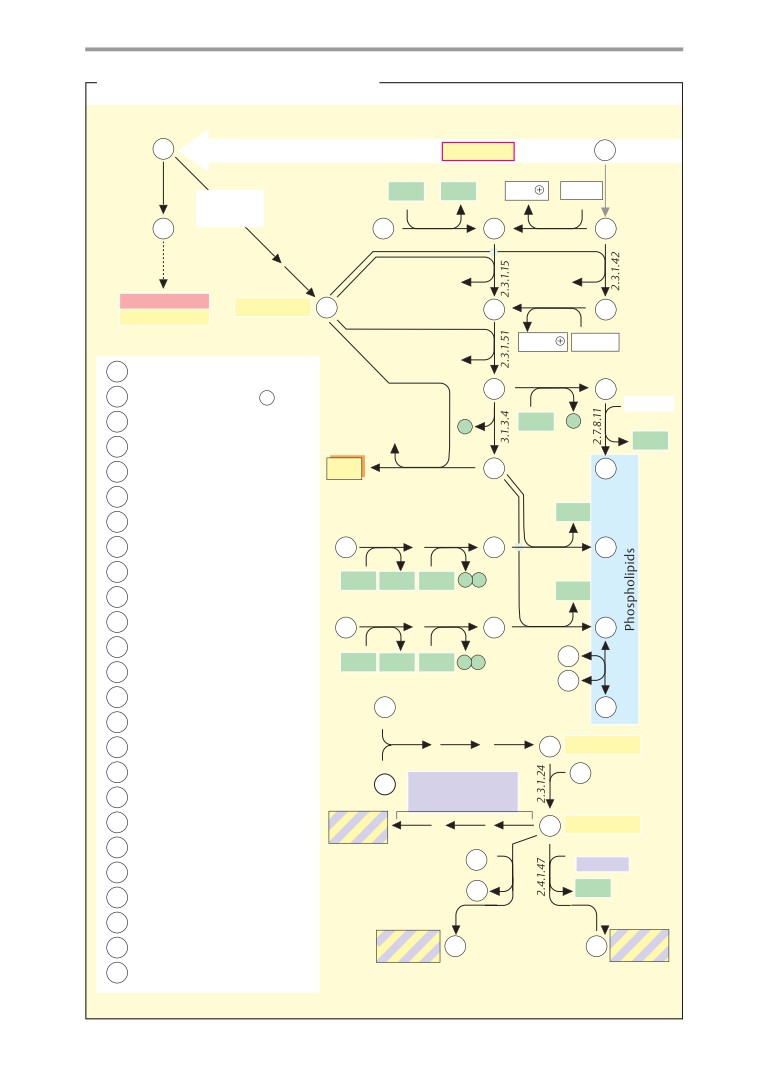
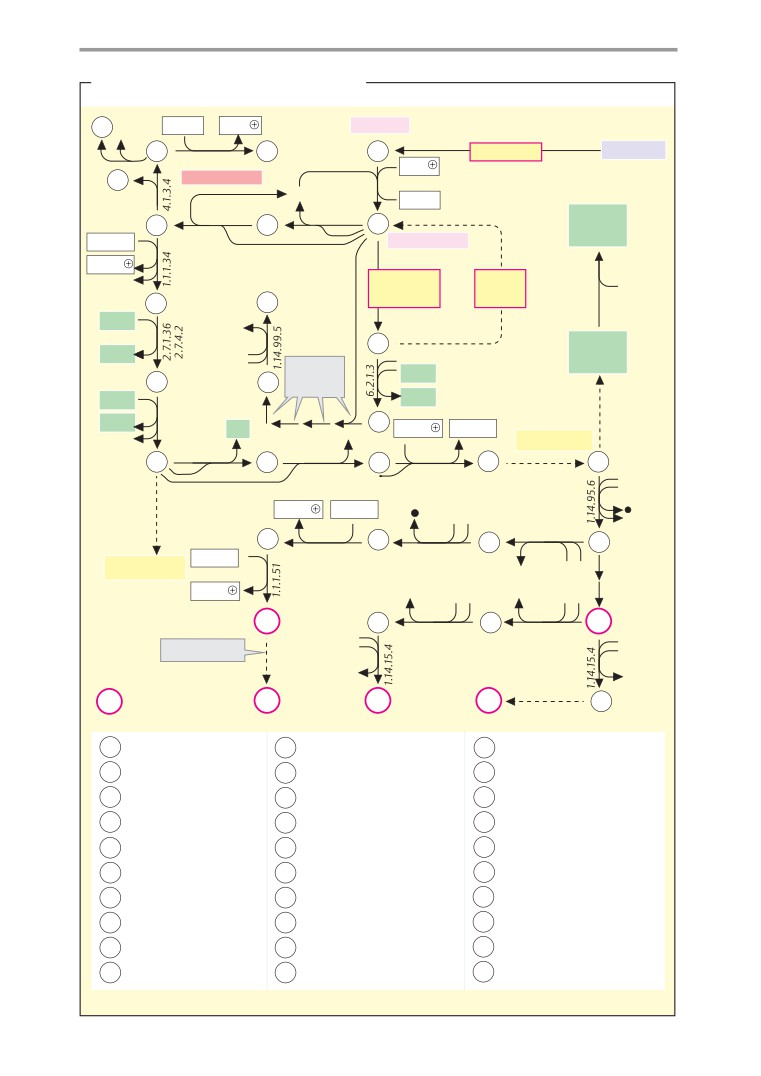
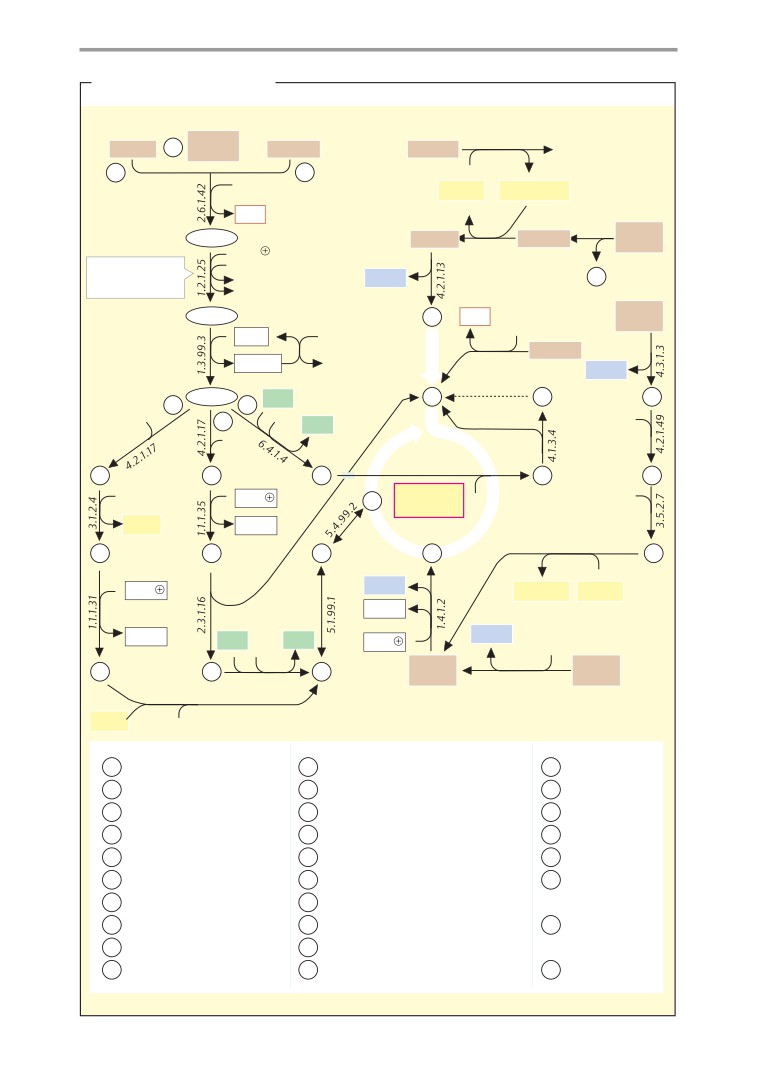
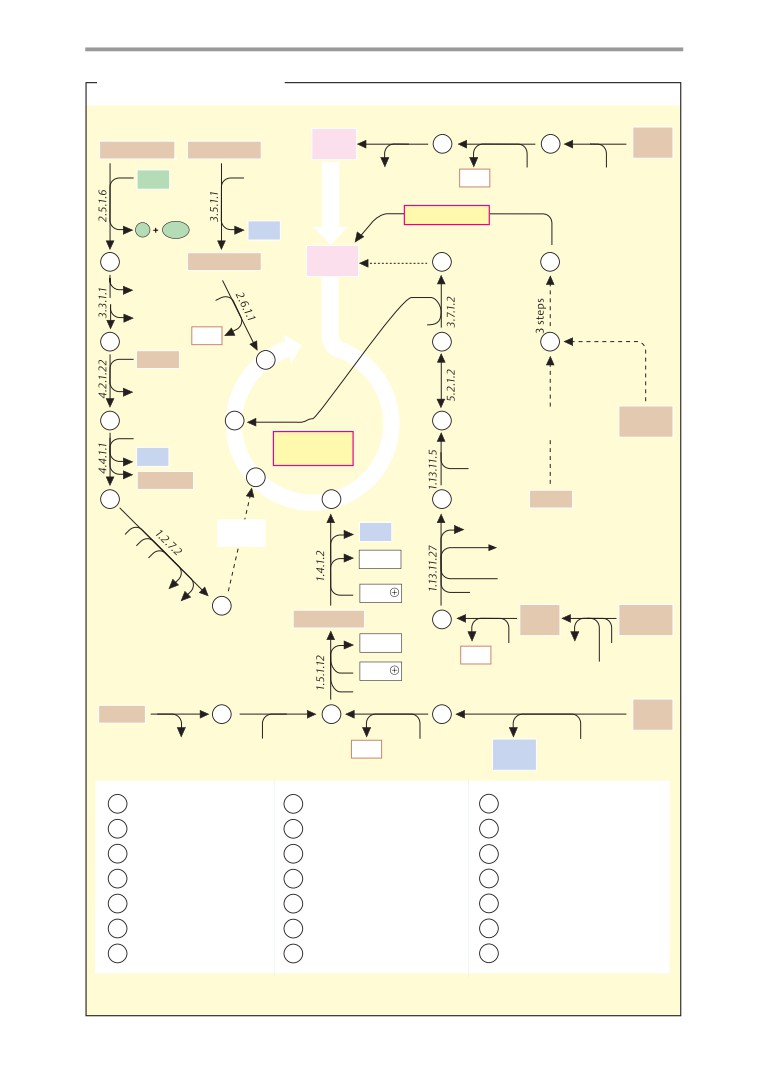
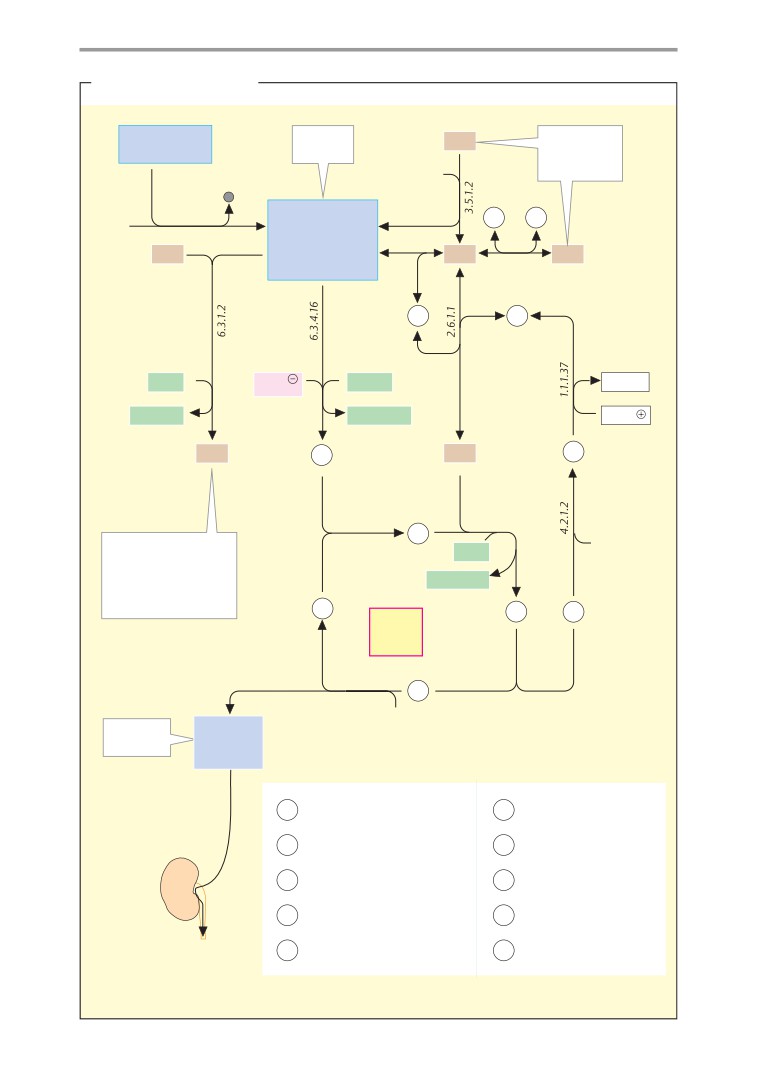
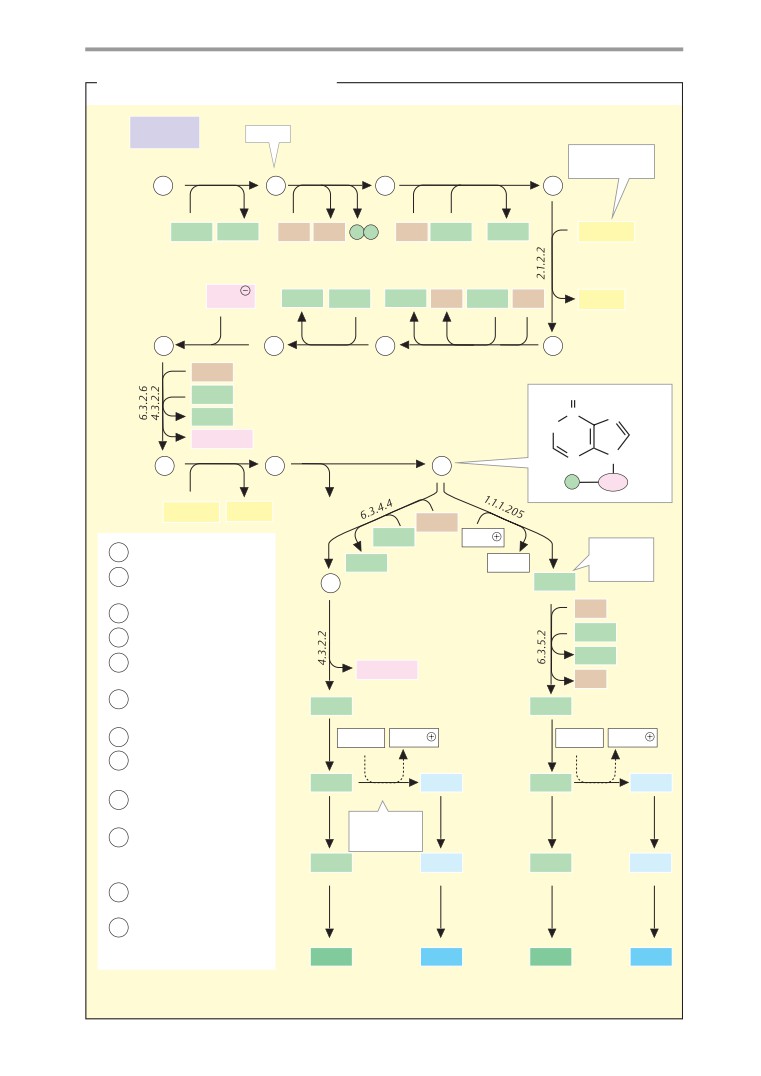
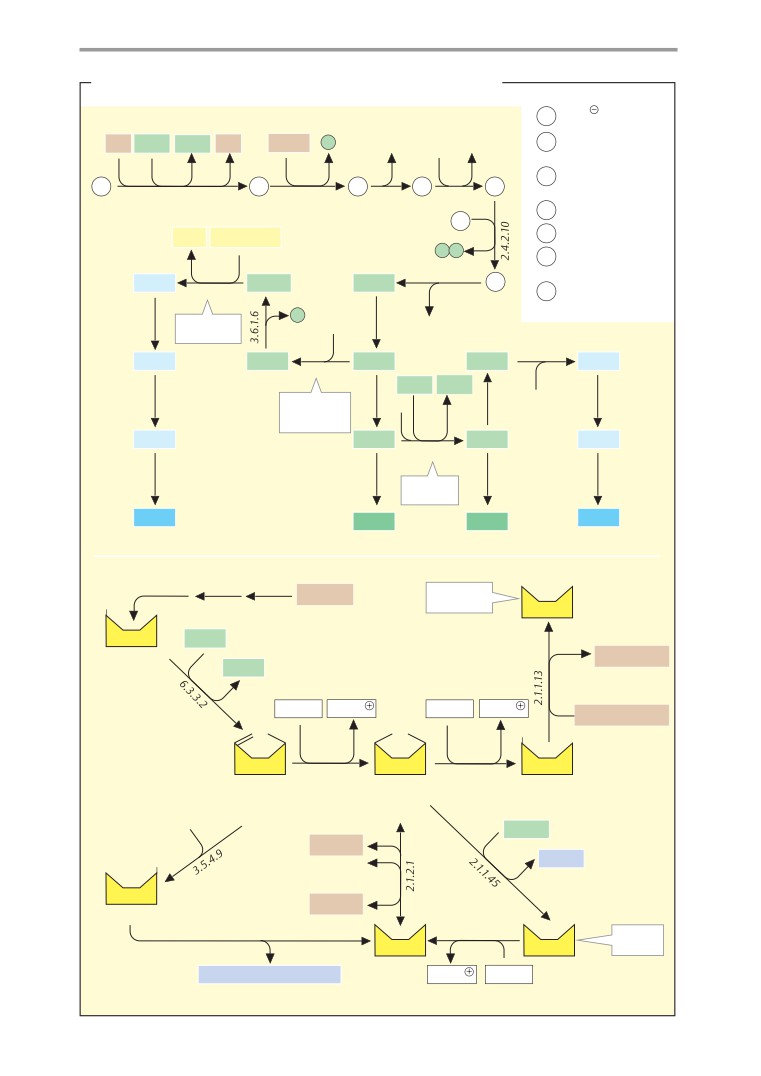
Metabolic charts
406
Muscle metabolism I
336
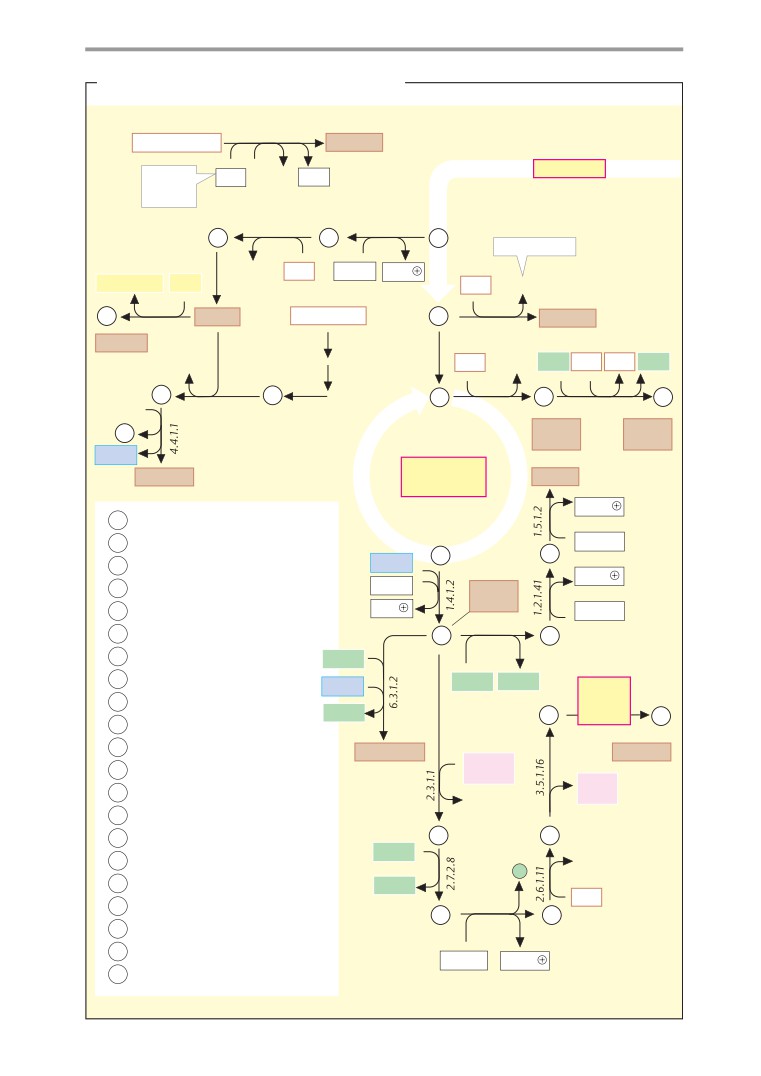
Calvin cycle
407
Muscle metabolism II
338
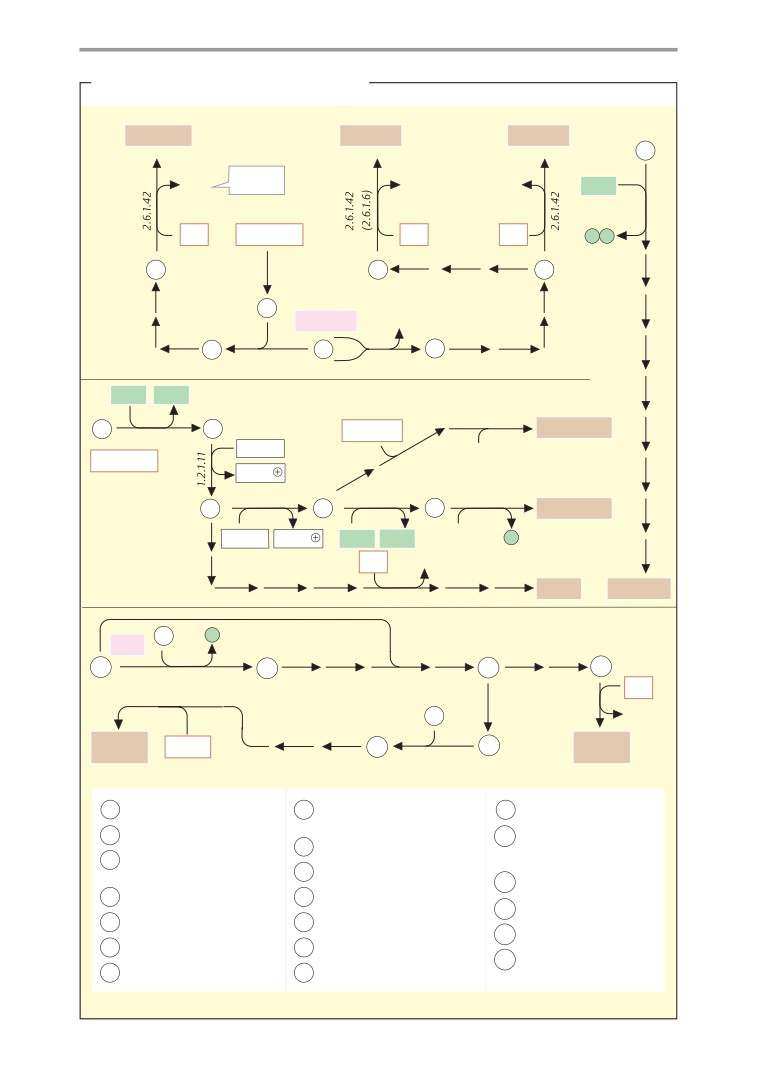
Carbohydrate metabolism
408
Connective tissue
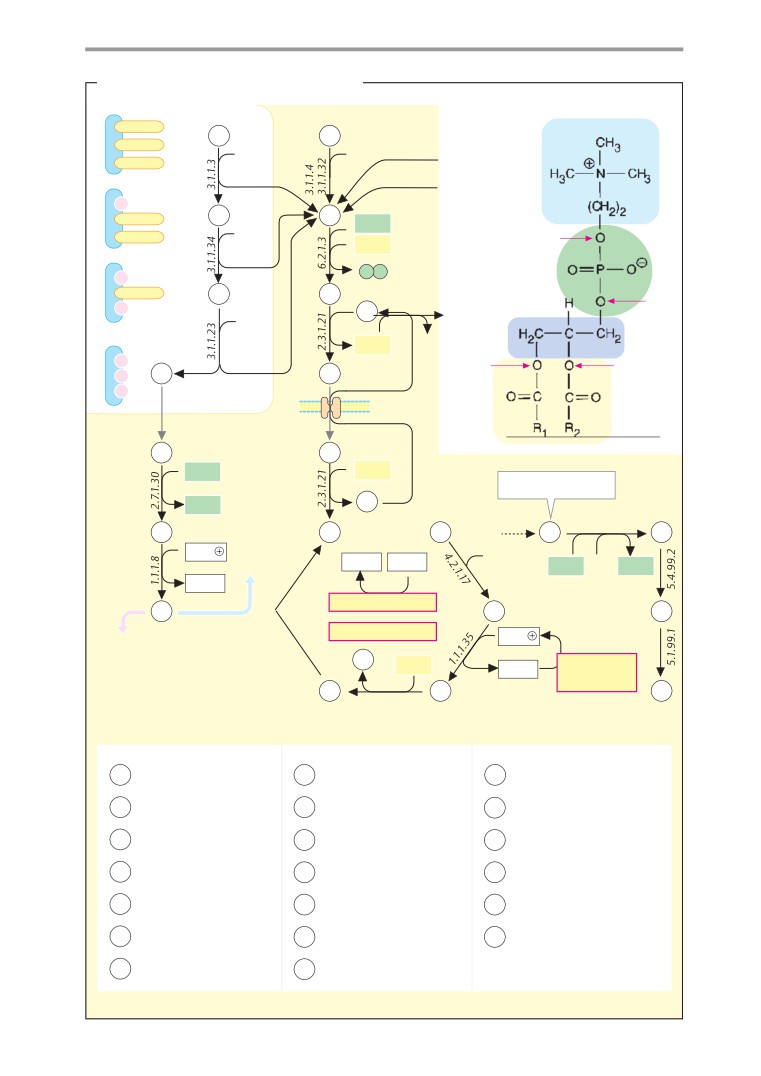
Biosynthesis of fats and
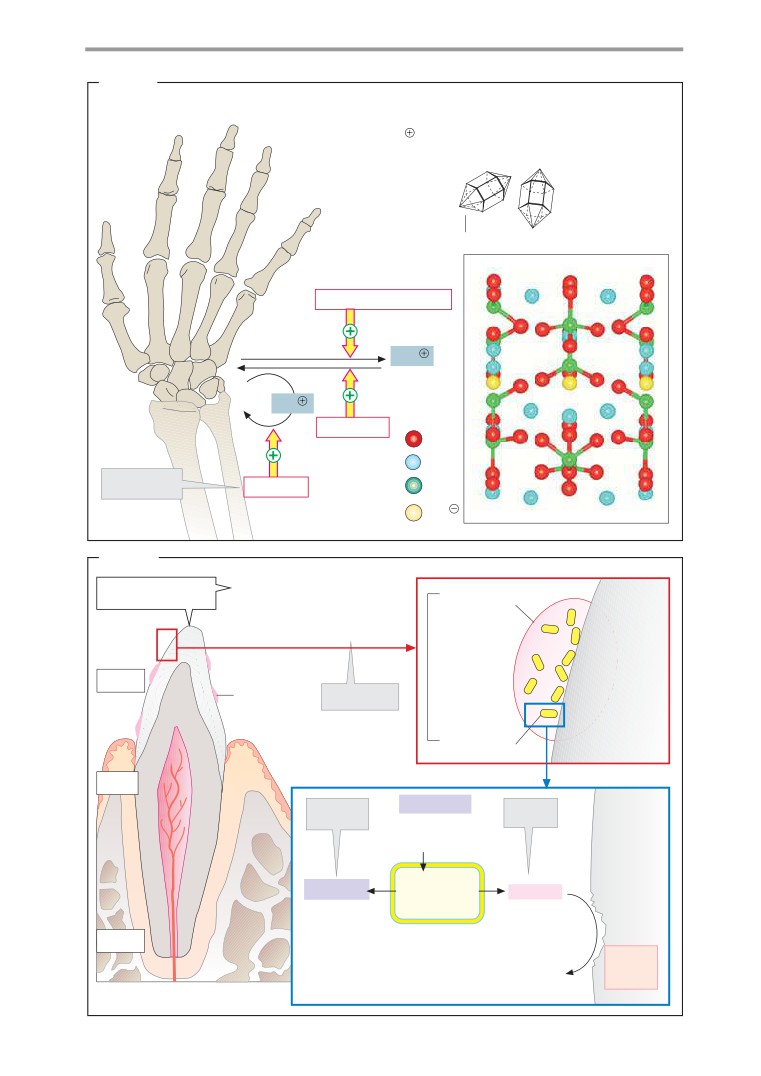
Bone and teeth
340
membrane liquids
409
Calcium metabolism
342
Synthesis of ketone bodies and steroids
410
Collagens
344
Degradation of fats and phospholipids .
411
Extracellular matrix
346
Biosynthesis of the essential
Brain and Sensory Organs
amino acids
412
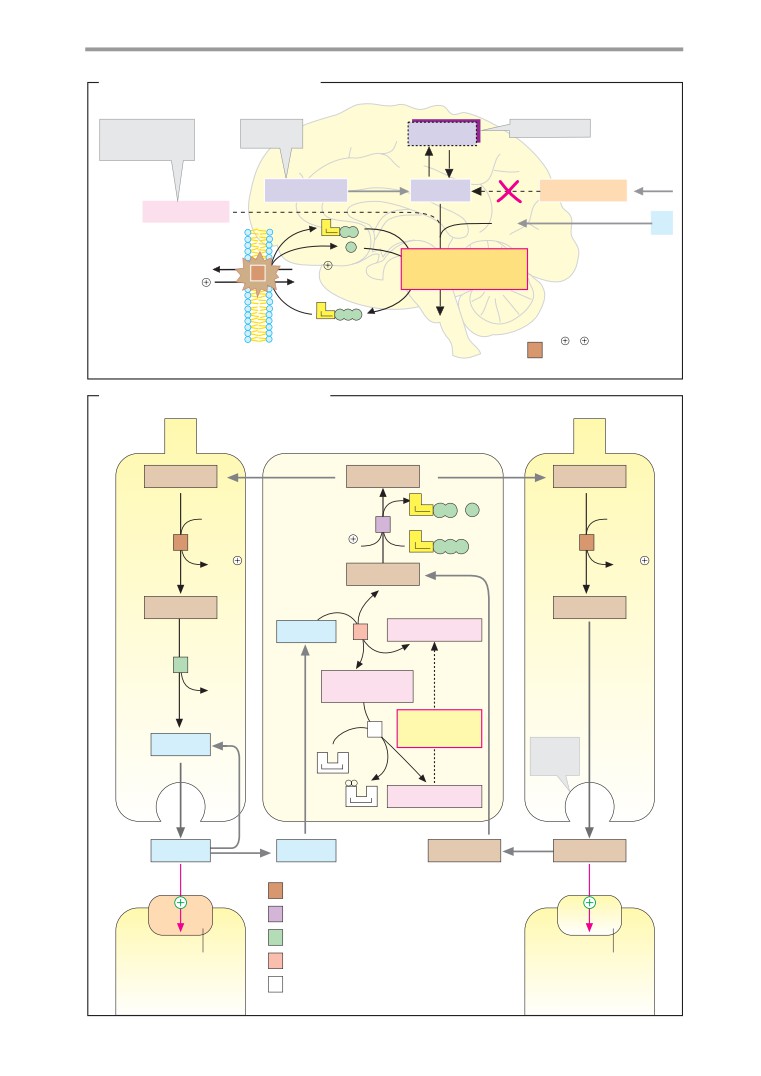
Signal transmission in the CNS
348
Biosynthesis of the non-essential
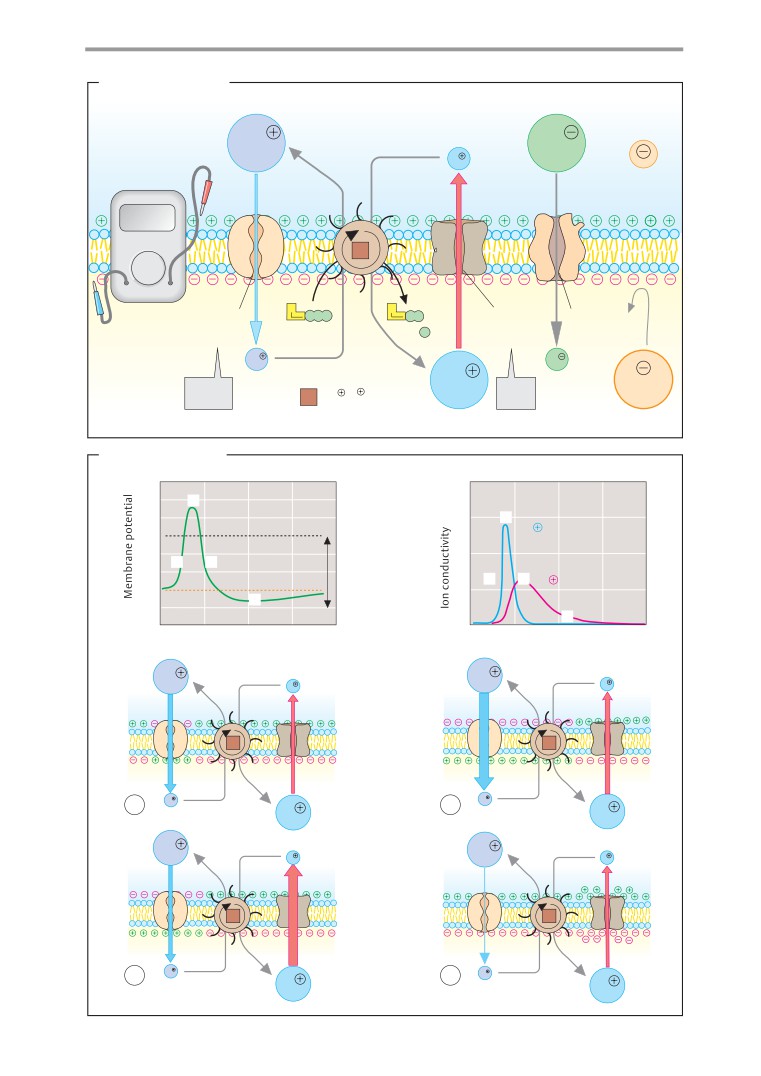
Resting potential and action potential. .
350
amino acids
413
Neurotransmitters
352
Amino acid degradation I
414
Receptors for neurotransmitters
354
Amino acid degradation II
415
Metabolism
356
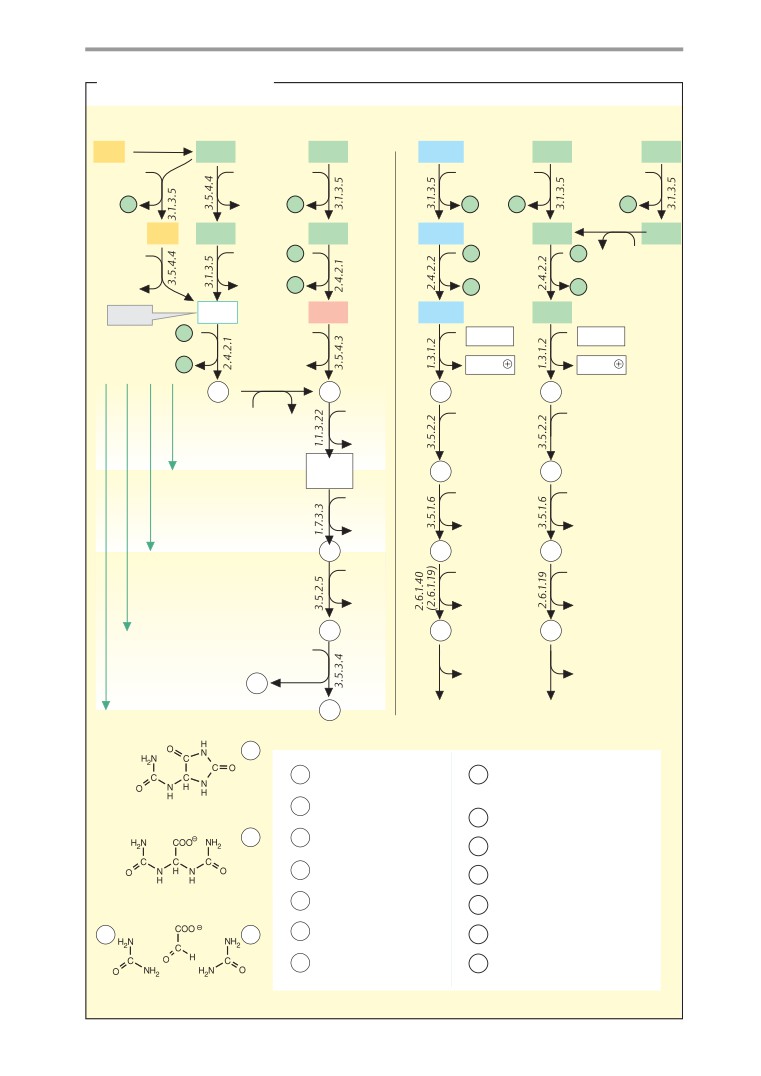
Ammonia metabolism
416
Sight
358
Biosynthesis of purine nucleotides
417
Biosynthesis of the pyrimidine nucleotides
Nutrition
and C1 metabolism
418
Nucleotide degradation
419
Nutrients
Organic substances
360
Annotated enzyme list
420
Minerals and trace elements
362
Abbreviations
431
Vitamins
Quantities and units
433
Lipid-soluble vitamins
364
Further reading
434
Water-soluble vitamins I
366
Water-soluble vitamins II
368
Source credits
435
Index
437
Hormones
Hormonal system
Basics
370
Key to color-coding:
Plasma levels and hormone hierarchy. .
372
see front and rear inside covers
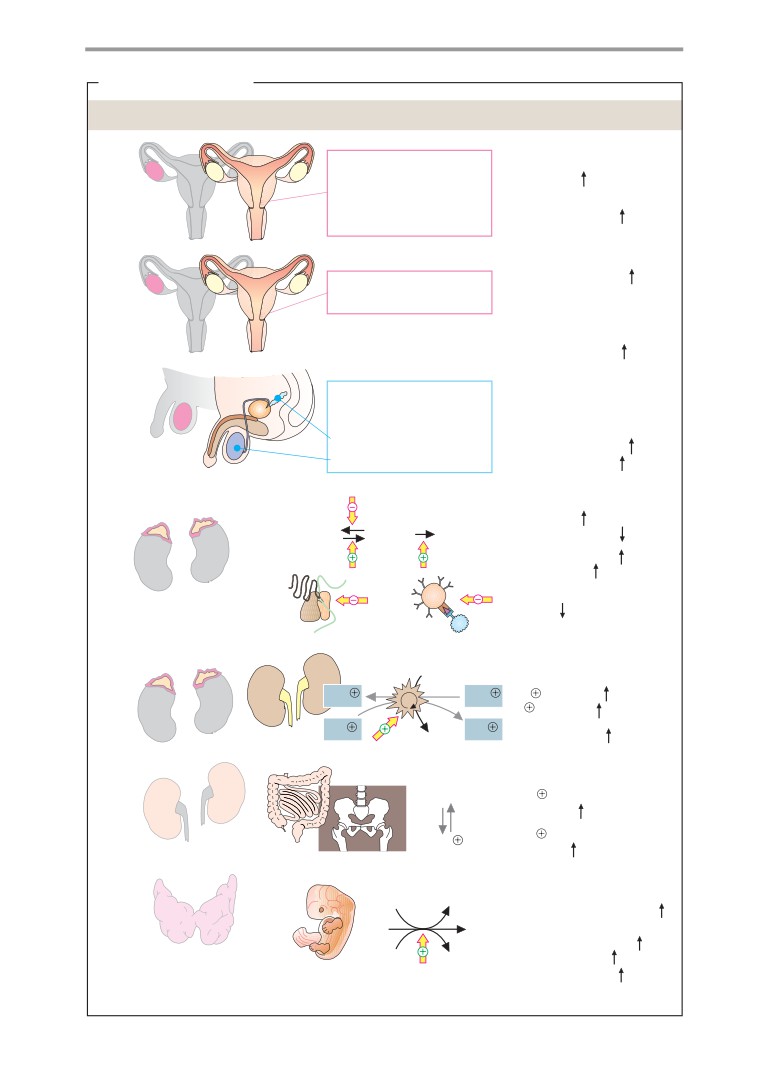
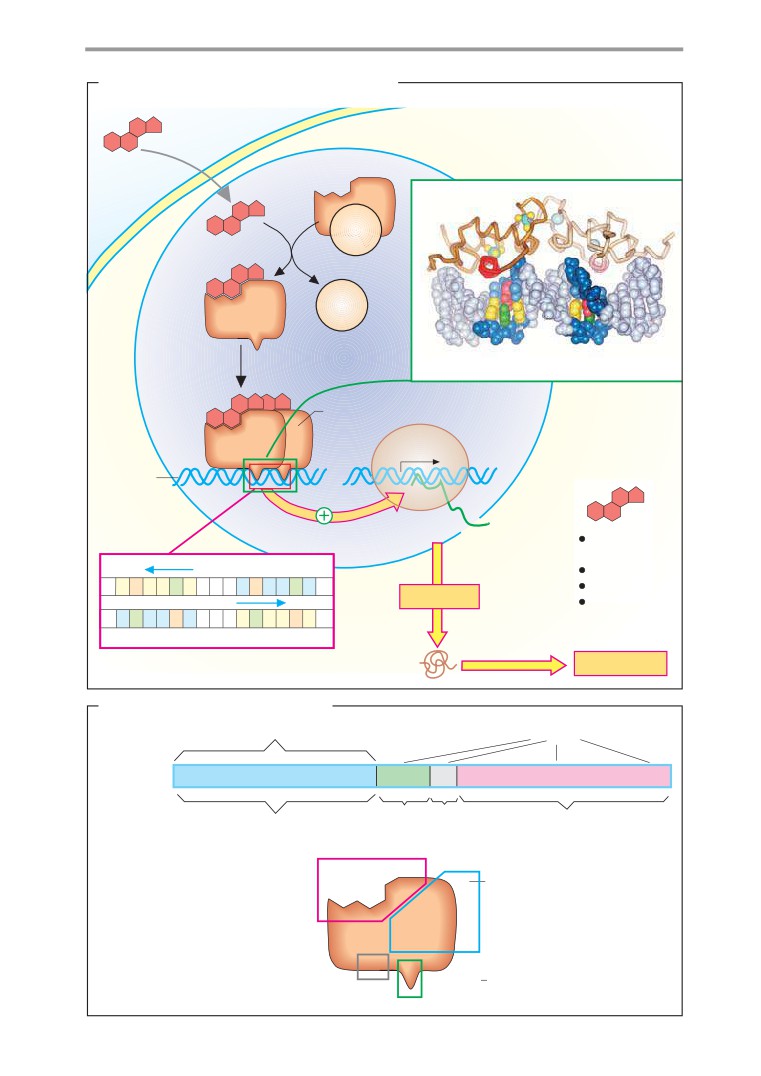
Lipophilic hormones
374
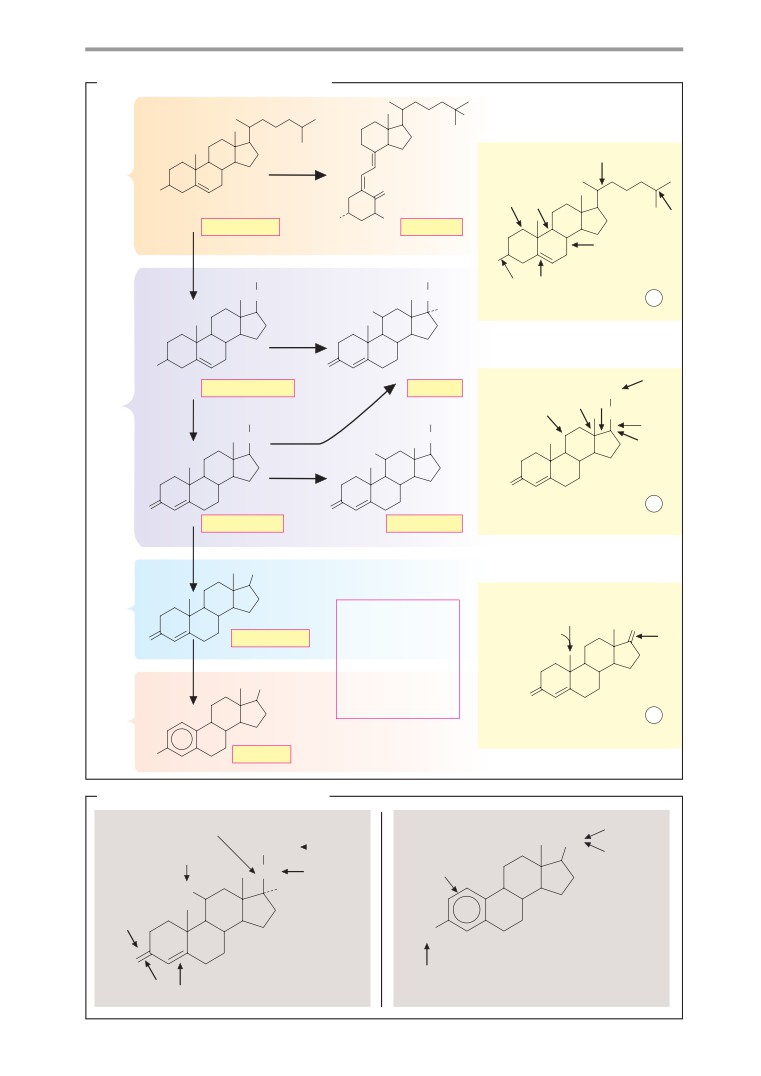
Metabolism of steroid hormones
376
Mechanism of action
378
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Chemistry
1
Introduction
This paperback atlas is intended for students
The next part presents the reactions
of medicine and the biological sciences. It
involved in the interconversion of these
provides an introduction to biochemistry,
compounds—the part of biochemistry that is
but with its modular structure it can also be
commonly referred to as metabolism
used as a reference book for more detailed
(pp. 88-195). The section starts with a dis-
information. The
216 color plates provide
cussion of the enzymes and coenzymes, and
knowledge in the field of biochemistry, ac-
discusses the mechanisms of metabolic regu-
companied by detailed information in the
lation and the so-called energy metabolism.
text on the facing page. The degree of dif -
After this, the central metabolic pathways
culty of the subject-matter is indicated by
are presented, once again arranged according
symbols in the text:
to the class of metabolite (pp. 150-195).
The second half of the book begins with a
stands for “basic biochemical knowledge”
discussion of the functional compartments
indicates
“standard biochemical knowl-
within the cell, the cellular organelles (pp.
edge”
196-235). This is followed on pp. 236-265
means “specialist biochemical knowledge.”
by the current field of molecular genetics
Some general rules used in the structure of
(molecular biology). A further extensive sec-
the illustrations are summed up in two ex-
tion is devoted to the biochemistry of
planatory plates inside the front and back
individual tissues and organs (pp. 266-359).
covers. Keywords, definitions, explanations
Here, it has only been possible to focus on the
of unfamiliar concepts and chemical formulas
most important organs and organ systems—
can be found using the index. The book starts
the digestive system, blood, liver, kidneys,
with a few basics in biochemistry (pp. 2-33).
muscles, connective and supportive tissues,
There is a brief explanation of the concepts
and the brain.
and principles of chemistry (pp. 2-15). These
Other topics include the biochemistry of
include the periodic table of the elements,
nutrition
(pp. 360-369), the structure and
chemical bonds, the general rules governing
function of important hormones
(pp.
molecular structure, and the structures of im-
370-393), and growth and development
portant classes of compounds. Several basic
(pp. 394-405).
concepts of physical chemistry are also essen-
The paperback atlas concludes with a series
tial for an understanding of biochemical
of schematic metabolic
“charts”
(pp.
processes. Pages 16-33 therefore discuss the
407-419). These plates, which are not accom-
various forms of energy and their intercon-
panied by explanatory text apart from a brief
version, reaction kinetics and catalysis, the
introduction on p. 406, show simplified ver-
properties of water, acids and bases, and re-
sions of the most important synthetic and
dox processes.
degradative pathways. The charts are mainly
These basic concepts are followed by a sec-
intended for reference, but they can also be
tion on the structure of the important biomo-
used to review previously learned material.
lecules (pp. 34-87). This part of the book is
The enzymes catalyzing the various reactions
arranged according to the different classes of
are only indicated by their EC numbers. Their
metabolites. It discusses carbohydrates, lipids,
names can be found in the systematically ar-
amino acids, peptides and proteins, nucleoti-
ranged and annotated enzyme list
(pp.
des, and nucleic acids.
420-430).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
2
Basics
tailed discussions of the subject are available
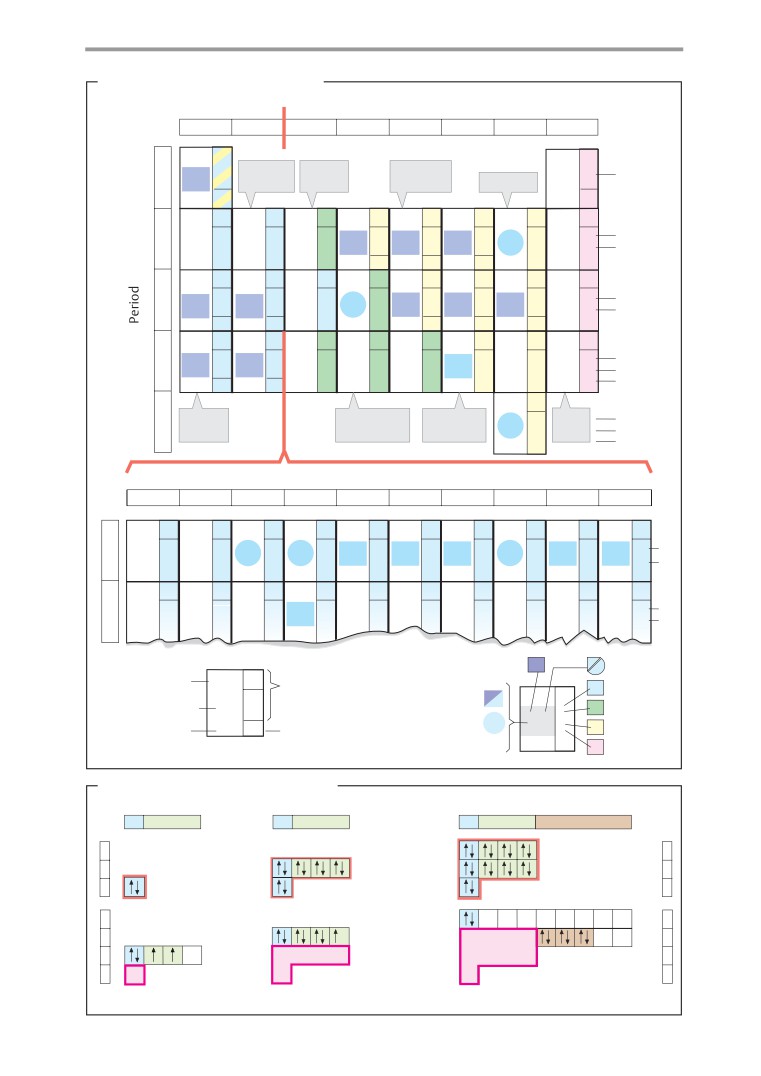
Periodic table
in chemistry textbooks.
The possible states of electrons are called
A. Biologically important elements
orbitals. These are indicated by what is
There are 81 stable elements in nature. Fifteen
known as the principal quantum number
of these are present in all living things, and a
and by a letter—s, p, or d. The orbitals are
further 8-10 are only found in particular or-
filled one by one as the number of electrons
ganisms. The illustration shows the first half
increases. Each orbital can hold a maximum of
of the periodic table, containing all of the bio-
two electrons, which must have oppositely
logically important elements. In addition to
directed “spins.” Fig. A shows the distribution
physical and chemical data, it also provides
of the electrons among the orbitals for each of
information about the distribution of the ele-
the elements. For example, the six electrons of
ments in the living world and their abun-
carbon (B1) occupy the 1s orbital, the 2s orbi-
dance in the human body. The laws of atomic
tal, and two 2p orbitals. A filled 1s orbital has
structure underlying the periodic table are
the same electron configuration as the noble
discussed in chemistry textbooks.
gas helium (He). This region of the electron
More than 99% of the atoms in animals’
shell of carbon is therefore abbreviated as
bodies are accounted for by just four ele-
“He” in Fig. A. Below this, the numbers of
ments—hydrogen (H), oxygen (O), carbon (C)
electrons in each of the other filled orbitals
and nitrogen (N). Hydrogen and oxygen are
(2s and 2p in the case of carbon) are shown on
the constituents of water, which alone makes
the right margin. For example, the electron
up 60-70 % of cell mass (see p. 196). Together
shell of chlorine (B2) consists of that of neon
with carbon and nitrogen, hydrogen and oxy-
(Ne) and seven additional electrons in 3s and
gen are also the major constituents of the
3p orbitals. In iron (B3), a transition metal of
organic compounds on which most living
the first series, electrons occupy the 4s orbital
processes depend. Many biomolecules also
even though the 3d orbitals are still partly
contain sulfur
(S) or phosphorus
(P). The
empty. Many reactions of the transition met-
above macroelements are essential for all or-
als involve empty d orbitals—e. g., redox reac-
ganisms.
tions or the formation of complexes with
A second biologically important group of
bases.
elements, which together represent only
Particularly stable electron arrangements
about 0.5% of the body mass, are present al-
arise when the outermost shell is fully occu-
most exclusively in the form of inorganic ions.
pied with eight electrons (the “octet rule”).
This group includes the alkali metals sodium
This applies, for example, to the noble gases,
(Na) and potassium (K), and the alkaline earth
as well as to ions such as Cl- (3s23p6) and Na+
metals magnesium (Mg) and calcium (Ca). The
(2s22p6). It is only in the cases of hydrogen
halogen chlorine (Cl) is also always ionized in
and helium that two electrons are already
the cell. All other elements important for life
suf cient to fill the outermost 1s orbital.
are present in such small quantities that they
are referred to as trace elements. These in-
clude transition metals such as iron (Fe), zinc
(Zn), copper (Cu), cobalt (Co) and manganese
(Mn). A few nonmetals, such as iodine (I) and
selenium (Se), can also be classed as essential
trace elements.
B. Electron configurations: examples
The chemical properties of atoms and the
types of bond they form with each other are
determined by their electron shells. The elec-
tron configurations of the elements are there-
fore also shown in Fig. A. Fig. B explains the
symbols and abbreviations used. More de-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Chemistry
3
A. Biologically important elements
Group
1
2
13
14
15
16
17
18
1.01
4.00
1
Alkaline
Boron
Nitrogen
2
1s
1
H
earths
group
group
Halogens
He
1
63
2
6.94
He
9.01
He
10.81
He
12.01
He
14.01
He
16.00
He
19.00
He
20.18
He
1
2
2
2
2
2
2
2
2s
2
i
Be
B
1
C
2
N
3
O
4
F
5
Ne
6
2p
L
3
4
5
6
9.5
7
1.4
8
25.5
9
10
22.99
Ne
24.31
Ne
26.98
Ne
28.09
Ne
30.97
Ne
32.07
Ne
35.45
Ne
39.95
Ne
3
1
2
2
2
2
2
2
2
3s
Na
Mg
l
1
Si
2
P
3
S
4
Cl
5
Ar
6
3p
A
11
0.03
12
0.01
13
14
15
0.22
16
0.05
17
0.03
18
39.10
Ar
40.08
Ar
69.72
Ar
72.61
Ar
74.92
Ar
78.96
Ar
79.90
Ar
83.80
Ar
4
1
2
10
10
10
10
10
10
3d
K
Ca
Ga
2
Ge
2
s
2
Se
2
r
2
Kr
2
4s
A
B
1
2
3
4
5
6
4p
19
0.06
20
0.31
31
32
33
34
35
36
126.9
Kr
5
Alkali
Carbon
Oxygen
10
Noble
4d
metals
group
group
I
2
gases
5s
53
5
5p
Group
3
4
5
6
7
8
9
10
11
12
44.96
Ar
47.88
Ar
50.94
Ar
52.00
Ar
54.94
Ar
55.85
Ar
58.93
Ar
58.69
Ar
63.55
Ar
65.39
Ar
1
2
3
4
5
6
7
8
9
10
3d
4
Sc
Ti
V
Cr
Mn
Fe
Co
Ni
Cu
Zn
2
2
2
2
2
2
2
2
2
2
4s
21
22
23
24
25
26
27
28
29
30
95.94
Kr
5
4
4d
Mo
2
5s
42
Macro element
Trace
element
Relative atomic
30.97
Ne
Electron
Essential for...
mass
configuration
Metal
all/most
2
Chemical symbol
organisms
P
3
Semi-metal
Percent (%) of
Atomic number
0.22
human body
for some
15
Non-metal
possibly
?
Noble gas
B. Electron configurations: examples
s
p
s
p
s
p
d
3
3
Helium
Neon
Argon
2
2
(He, Noble gas)
(Ne, Noble gas)
(Ar, Noble gas)
1
1
1s2
1s2 2s2 2p6
1s2 2s2 2p6 3s2 3p6
4
1. Carbon (C)
4
2
[He] 2s2 2p
3
3
[Ar]
2
[Ne]
2
2. Chlorine (Cl)
3. Iron (Fe)
1
[He]
1
[Ne] 3s2 3p5
[Ar] 4s2 3d6
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
4
Basics
tatable, since rotation would distort the π-
Bonds
molecular orbital. This is why all of the atoms
lie in one plane (2c); in addition, cis-trans
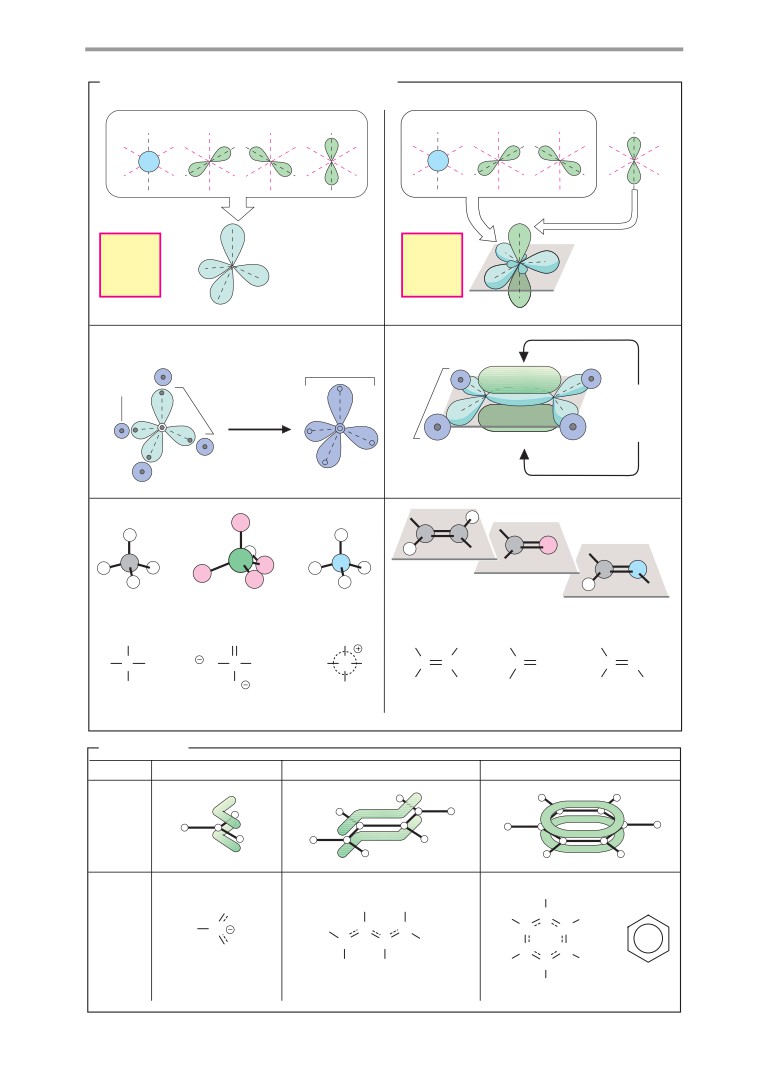
A. Orbital hybridization and chemical
isomerism arises in such cases
(see p. 8).
bonding
Double bonds that are common in biomole-
Stable, covalent bonds between nonmetal
cules are C=C and C=O. C=N double bonds are
atoms are produced when orbitals (see p. 2)
found in aldimines (Schiff bases, see p. 178).
of the two atoms form molecular orbitals that
are occupied by one electron from each of the
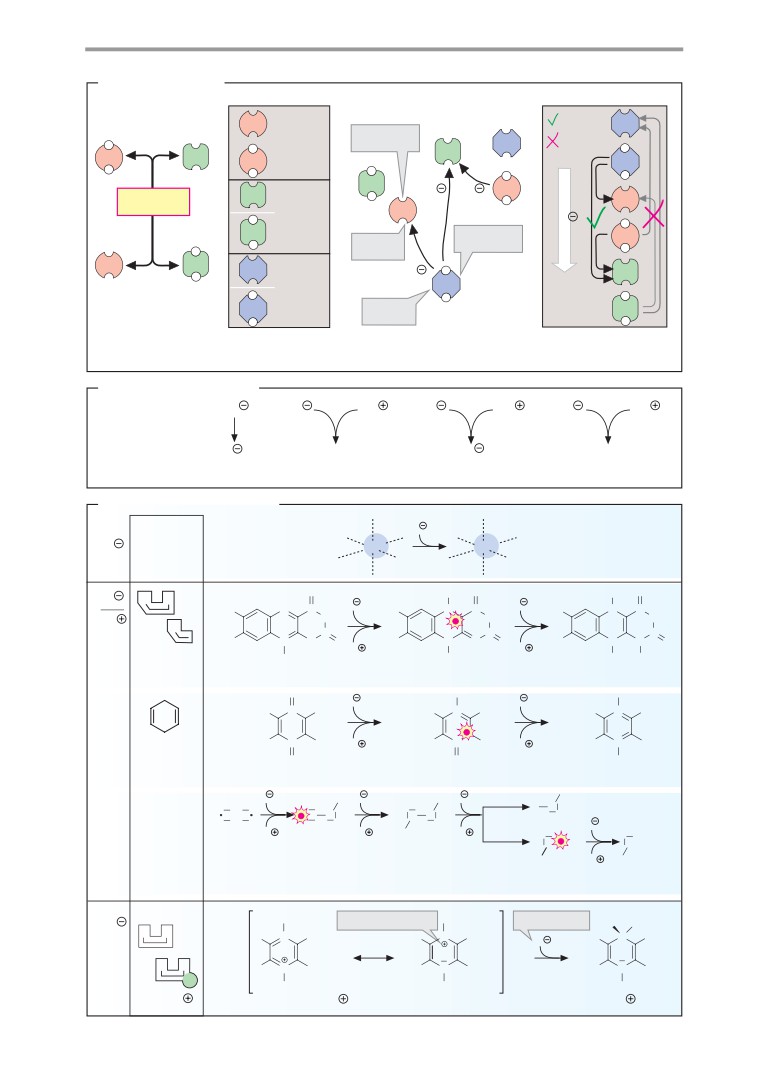
B. Resonance
atoms. Thus, the four bonding electrons of the
carbon atom occupy 2s and 2p atomic orbitals
Many molecules that have several double
(1a). The 2s orbital is spherical in shape, while
bonds are much less reactive than might be
the three 2p orbitals are shaped like dumb-
expected. The reason for this is that the
bells arranged along the x, y, and z axes. It
double bonds in these structures cannot be
might therefore be assumed that carbon
localized unequivocally. Their π orbitals are
atoms should form at least two different types
not confined to the space between the dou-
of molecular orbital. However, this is not nor-
ble-bonded atoms, but form a shared,
mally the case. The reason is an effect known
extended S-molecular orbital. Structures
as orbital hybridization. Combination of the s
with this property are referred to as reso-
orbital and the three p orbitals of carbon gives
nance hybrids, because it is impossible to de-
rise to four equivalent, tetrahedrally arranged
scribe their actual bonding structure using
sp3 atomic orbitals (sp3 hybridization). When
standard formulas. One can either use what
these overlap with the 1s orbitals of H atoms,
are known as resonance structures—i. e.,
four equivalent σ-molecular orbitals (1b) are
idealized configurations in which π electrons
formed. For this reason, carbon is capable of
are assigned to specific atoms (cf. pp. 32 and
forming four bonds—i. e., it has a valency of
66, for example)—or one can use dashed lines
four. Single bonds between nonmetal atoms
as in Fig. B to suggest the extent of the delo-
arise in the same way as the four σ or single
calized orbitals.
(Details are discussed in
bonds in methane (CH4). For example, the
chemistry textbooks.)
hydrogen phosphate ion (HPO42-) and the
Resonance-stabilized systems include car-
ammonium ion (NH4+) are also tetrahedral
boxylate groups, as in formate; aliphatic hy-
in structure (1c).
drocarbons with conjugated double bonds,
A second common type of orbital hybrid-
such as 1,3-butadiene; and the systems known
ization involves the 2s orbital and only two of
as aromatic ring systems. The best-known
the three 2p orbitals (2a). This process is
aromatic compound is benzene, which has
therefore referred to as sp2
hybridization.
six delocalized π electrons in its ring. Ex-
The result is three equivalent sp2 hybrid orbi-
tended resonance systems with 10 or more
tals lying in one plane at an angle of 120° to
π electrons absorb light within the visible
one another. The remaining 2px orbital is ori-
spectrum and are therefore colored. This
ented perpendicular to this plane. In contrast
group includes the aliphatic carotenoids (see
to their sp3
counterparts, sp2-hybridized
p. 132), for example, as well as the heme
atoms form two different types of bond
group, in which 18 π electrons occupy an ex-
when they combine into molecular orbitals
tended molecular orbital (see p. 106).
(2b). The three sp2 orbitals enter into σ bonds,
as described above. In addition, the electrons
in the two 2px orbitals, known as S electrons,
combine to give an additional, elongated π
molecular orbital, which is located above
and below the plane of the σ bonds. Bonds
of this type are called double bonds. They
consist of a σ bond and a π bond, and arise
only when both of the atoms involved are
capable of sp2 hybridization. In contrast to
single bonds, double bonds are not freely ro-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Chemistry
5
A. Orbital hybridization and chemical bonding
S
P
S
z
Py
Px
Pz
Py
Px
4 Equivalent
3 Equivalent
sp3
sp2
Hybrid-
sp3 atomic
Hybrid-
sp2 atomic
ization
orbitals
ization
orbitals
(tetrahedral)
(trigonal)
1a
2a
1s Orbital
sp3 Atomic
4 Bonding
5 Bonding
of
orbitals
σ-molecular
σ-molecular
hydro-
of
orbitals
orbitals
gen
carbon
atom
atom
Bonding
π-molecular
orbitals
C
+
4 H
CH4
1b
2b
Methane
Hydrogen phosphate Ammonium
Alkene
Carbonyl
Aldimine
Ion
compound
H
O
H
R
H
R
R
H C H
O
P
OH
H N
H
C C
C O
C N
H
R'
R'
H
R'
H
O
H
1c
2c
B. Resonance
Formate
1,3-Butadiene
Benzene
π-
Molecular
orbitals
H
H
H
O
H
C
H
Formula
H C
H
C
C
C
C
C
C
H
O
C
C
H
C
H
H H
H
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
6
Basics
Molecular structure
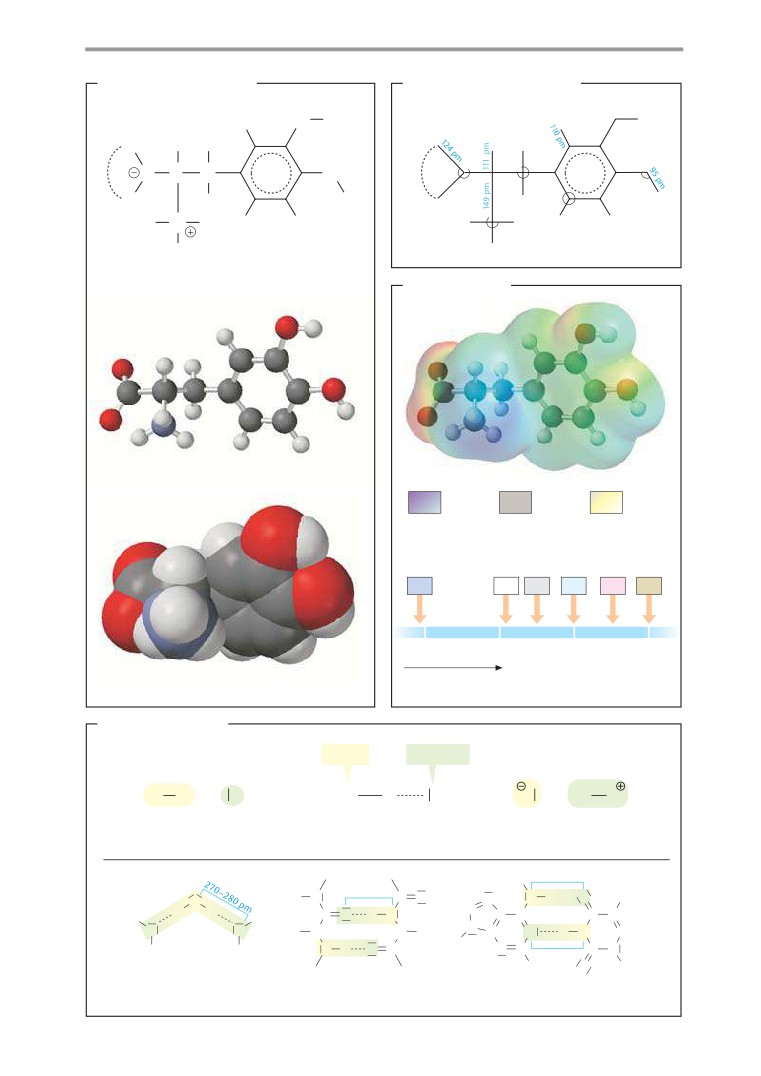
B. Bond lengths and angles
The physical and chemical behavior of mole-
Atomic radii and distances are now usually
cules is largely determined by their constitu-
expressed in picometers
(pm;
1 pm =
tion (the type and number of the atoms they
10-12 m). The old angstrom unit
(Å,
contain and their bonding). Structural formu-
Å = 100 pm) is now obsolete. The length of
las can therefore be used to predict not only
single bonds approximately corresponds to
the chemical reactivity of a molecule, but also
the sum of what are known as the covalent
its size and shape, and to some extent its
radii of the atoms involved (see inside front
conformation
(the spatial arrangement of
cover). Double bonds are around
10-20%
the atoms). Some data providing the basis
shorter than single bonds. In sp3-hybridized
for such predictions are summarized here
atoms, the angle between the individual
and on the facing page. In addition, L-dihy-
bonds is approx.
110°; in sp2-hybridized
droxyphenylalanine (L-dopa; see p. 352), is
atoms it is approx. 120°.
used as an example to show the way in which
molecules are illustrated in this book.
C. Bond polarity
A. Molecule illustrations
Depending on the position of the element in
the periodic table
(see p. 2), atoms have
In traditional two-dimensional structural
different electronegativity—i. e., a different
formulas (A1), atoms are represented as letter
tendency to take up extra electrons. The val-
symbols and electron pairs are shown as lines.
ues given in C2 are on a scale between 2 and 4.
Lines between two atomic symbols symbolize
The higher the value, the more electronega-
two bonding electrons (see p. 4), and all of the
tive the atom. When two atoms with very
other lines represent free electron pairs, such
different electronegativities are bound to
as those that occur in O and N atoms. Free
one another, the bonding electrons are drawn
electrons are usually not represented explic-
toward the more electronegative atom, and
itly (and this is the convention used in this
the bond is polarized. The atoms involved
book as well). Dashed or continuous circles or
then carry positive or negative partial
arcs are used to emphasize delocalized elec-
charges. In C1, the van der Waals surface is
trons.
colored according to the different charge con-
Ball-and-stick models (A2) are used to illus-
ditions (red = negative, blue = positive). Oxy-
trate the spatial structure of molecules. Atoms
gen is the most strongly electronegative of the
are represented as colored balls (for the color
biochemically important elements, with C=O
coding, see the inside front cover) and bonds
double bonds being especially highly polar.
(including multiple bonds) as gray cylinders.
Although the relative bond lengths and angles
correspond to actual conditions, the size at
D. Hydrogen bonds
which the atoms are represented is too small
to make the model more comprehensible.
The hydrogen bond, a special type of nonco-
Space-filling van der Waals models (A3) are
valent bond, is extremely important in bio-
useful for illustrating the actual shape and
chemistry. In this type of bond, hydrogen
size of molecules. These models represent
atoms of OH, NH, or SH groups (known as
atoms as truncated balls. Their effective ex-
hydrogen bond donors) interact with free
tent is determined by what is known as the
electrons of acceptor atoms (for example, O,
van der Waals radius. This is calculated from
N, or S). The bonding energies of hydrogen
the energetically most favorable distance be-
bonds
(10-40 kJ mol-1) are much lower
tween atoms that are not chemically bonded
than those of covalent bonds
(approx.
to one another.
400 kJ mol-1). However, as hydrogen bonds
can be very numerous in proteins and DNA,
they play a key role in the stabilization of
these molecules (see pp. 68, 84). The impor-
tance of hydrogen bonds for the properties of
water is discussed on p. 26.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Chemistry
7
A. Molecule illustrations
B. Bond lengths and angles
Chiral center
H
O
H
H H
140 pm
O
154 pm
110°
137 pm
C C C
O
120°
120°
110°
110°
108°
120°
O
H
H
120°
120°
110°
100 pm
H
N
H
H
H
110°
110°
H
1. Formula illustration
C. Bond polarity
2. Ball- and-stick model
Positive
Neutral
Negative
1. Partial charges in L-dopa
0.9
2.1
2.5
3.0
3.5
4.0
Na
H
C
N
O
F
1
2
3
4
Increasing electronegativity
3. Van der Waals model
2. Electronegativities
D. Hydrogen bonds
Dissociated Protonated
Acid
Base
Donor
Acceptor
acid
base
A H
B
A
H
B
A
H B
Initial state
Hydrogen bond
Complete reaction
1. Principle
290 pm
H
280 pm
H
N
C O
N H O
CH3
O
N
HC
H H
C
O
H
N
C C
C
C
H
H
O
O
R1
CH
HC R2
N
C
N
H N
CH
R
H
H
N H O C
N C
C N
290 pm
H
O
R
Water
Proteins
DNA
2. Examples
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
8
Basics
romolecules such as proteins or nucleic acids
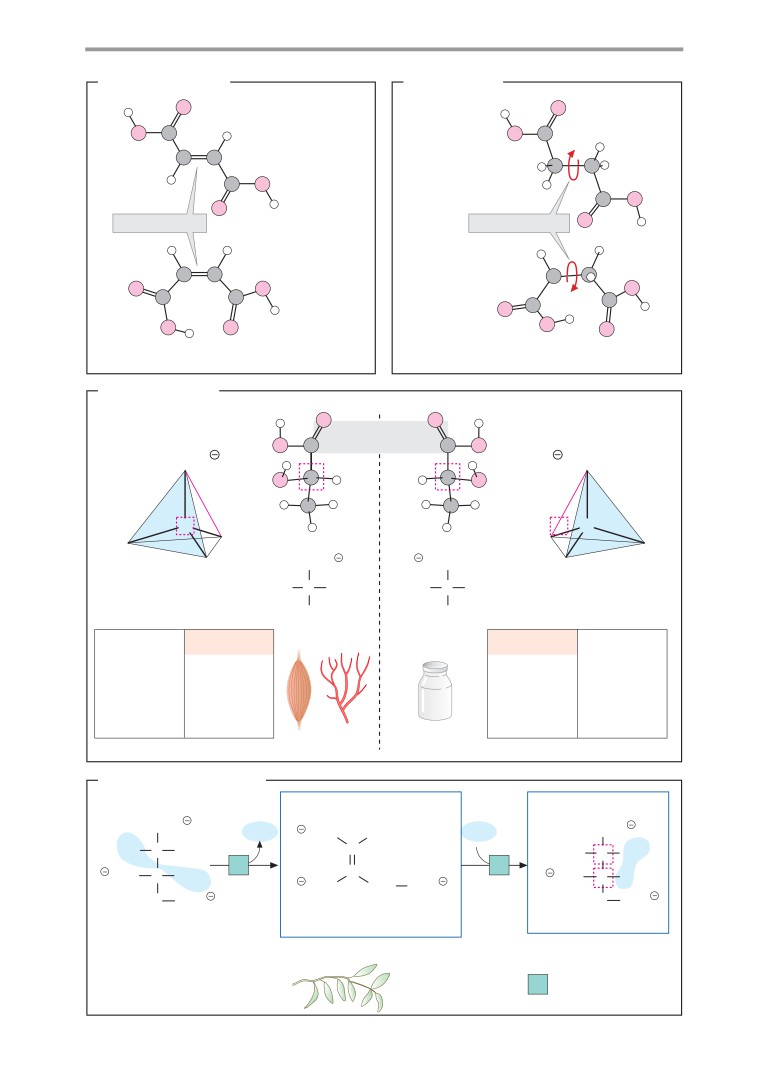
Isomerism
usually have well-defined (“native”) confor-
Isomers are molecules with the same compo-
mations, which are stabilized by interactions
sition (i. e. the same molecular formula), but
in the molecule (see p. 74).
with different chemical and physical proper-
ties. If isomers differ in the way in which their
C. Optical isomers
atoms are bonded in the molecule, they are
described as structural isomers (cf. citric acid
Another type of isomerism arises when a mol-
and isocitric acid, D). Other forms of isomer-
ecule contains a chiral center or is chiral as a
ism are based on different arrangements of
whole. Chirality (from the Greek cheir, hand)
the substituents of bonds (A, B) or on the
leads to the appearance of structures that
presence of chiral centers in the molecule (C).
behave like image and mirror-image and
that cannot be superimposed (“mirror” iso-
mers). The most frequent cause of chiral be-
A. cis-trans isomers
havior is the presence of an asymmetric C
Double bonds are not freely rotatable (see
atom—i. e., an atom with four different sub-
p. 4). If double-bonded atoms have different
stituents. Then there are two forms (enan-
substituents, there are two possible orienta-
tiomers) with different configurations. Usu-
tions for these groups. In fumaric acid, an
ally, the two enantiomers of a molecule are
intermediate of the tricarboxylic acid cycle
designated as L and D forms. Clear classifica-
(see p. 136), the carboxy groups lie on different
tion of the configuration is made possible by
sides of the double bond (trans or E position).
the R/S system (see chemistry textbooks).
In its isomer maleic acid, which is not pro-
Enantiomers have very similar chemical
duced in metabolic processes, the carboxy
properties, but they rotate polarized light in
groups lie on the same side of the bond (cis
opposite directions
(optical activity, see
or Z position). Cis-trans isomers (geometric
pp. 36, 58). The same applies to the enantiom-
isomers) have different chemical and physical
ers of lactic acid. The dextrorotatory L-lactic
properties—e. g., their melting points
(Fp.)
acid occurs in animal muscle and blood, while
and pKa values. They can only be intercon-
the D form produced by microorganisms is
verted by chemical reactions.
found in milk products, for example (see
In lipid metabolism, cis-trans isomerism is
p. 148). The Fischer projection is often used
particularly important. For example, double
to represent the formulas for chiral centers
bonds in natural fatty acids (see p. 48) usually
(cf. p. 58).
have a cis configuration. By contrast, unsatu-
rated intermediates of β oxidation have a
D. The aconitase reaction
trans configuration. This makes the break-
down of unsaturated fatty acids more compli-
Enzymes usually function stereospecifically. In
cated (see p. 166). Light-induced cis-trans iso-
chiral substrates, they only accept one of the
merization of retinal is of central importance
enantiomers, and the reaction products are
in the visual cycle (see p. 358).
usually also sterically uniform. Aconitate
hydratase (aconitase) catalyzes the conver-
sion of citric acid into the constitution isomer
B. Conformation
isocitric acid (see p. 136). Although citric acid
Molecular forms that arise as a result of rota-
is not chiral, aconitase only forms one of the
tion around freely rotatable bonds are known
four possible isomeric forms of isocitric acid
as conformers. Even small molecules can have
(2R,3S-isocitric acid). The intermediate of the
different conformations in solution. In the
reaction, the unsaturated tricarboxylic acid
two conformations of succinic acid illustrated
aconitate, only occurs in the cis form in the
opposite, the atoms are arranged in a similar
reaction. The trans form of aconitate is found
way to fumaric acid and maleic acid. Both
as a constituent of certain plants.
forms are possible, although conformation 1
is more favorable due to the greater distance
between the COOH groups and therefore oc-
curs more frequently. Biologically active mac-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Chemistry
9
A. cis-trans isomers
B. Conformers
Fumaric acid
Succinic acid
Fp. 287 °C
Conformation 1
pKa 3.0, 4.5
Not rotatable
Freely rotatable
Maleic acid
Succinic
Fp. 130 °C
acid
pKa 1.9, 6.5
Conformation 2
C. Optical isomers
Fischer projections
COO
C
CH3
HO
COO
OOC
H
L(S)
HO
C
H
H
C
HO
D(R)
CH3
CH3
L-lactic acid
D-lactic acid
Fp.
53 °C
53 °C
Fp.
pKa value
3.7
3.7
pKa value
Specific
+ 2.5˚
-2.5˚
Specific
rotation
rotation
In muscle, blood
In milk products
D. The aconitase reaction
COO
H2O
H2O
COO
OOC
H
H
C
H
C
H
C
OH
2
1
1
C
OOC
C
OH
OOC
C
H
3
OOC
CH2
COO
H2C
COO
H2C
COO
Citrate (prochiral)
cis-Aconitate (intermediate product)
(2R,3S)-Isocitrate
trans-Aconitate occurs in plants
1
Aconitase 4.2.1.3
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
10
Basics
acetals (see p. 36). The oxidation of hemiace-
Biomolecules I
tals produces carboxylic acid esters.
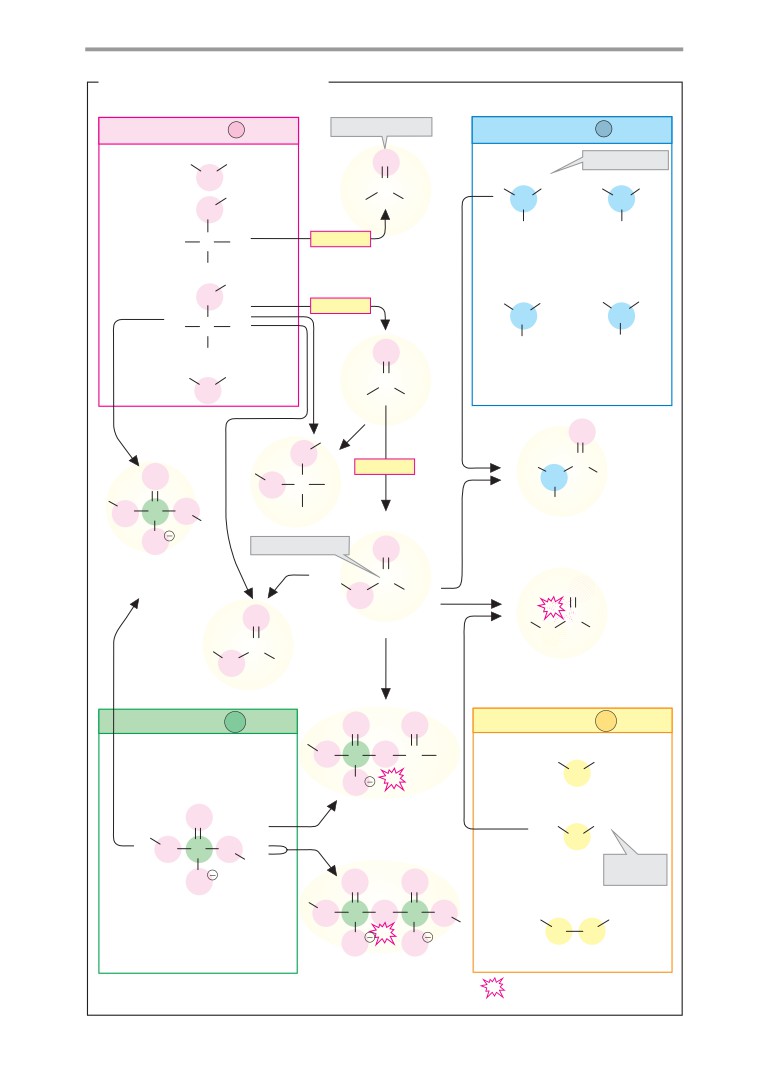
A. Important classes of compounds
Very important compounds are the carbox-
Most biomolecules are derivatives of simple
ylic acids and their derivatives, which can be
compounds of the non-metals oxygen (O),
formally obtained by exchanging the OH
hydrogen (H), nitrogen (N), sulfur (S), and
group for another group. In fact, derivatives
phosphorus (P). The biochemically important
of this type are formed by nucleophilic sub-
oxygen, nitrogen, and sulfur compounds can
stitutions of activated intermediate com-
be formally derived from their compounds
pounds and the release of water (see p. 14).
with hydrogen (i. e., H2O, NH3, and H2S). In
Carboxylic acid esters (R-O-CO-R ) arise from
biological systems, phosphorus is found al-
carboxylic acids and alcohols. This group in-
most exclusively in derivatives of phosphoric
cludes the fats, for example (see p. 48). Sim-
acid, H3PO4.
ilarly, a carboxylic acid and a thiol yield a
thioester (R-S-CO-R ). Thioesters play an ex-
If one or more of the hydrogen atoms of a
tremely important role in carboxylic acid me-
non-metal hydride are replaced formally with
tabolism. The best-known compound of this
another group, R—e. g., alkyl residues—then
type is acetyl-coenzyme A (see p. 12).
derived compounds of the type R-XHn-1,
R-XHn-2-R, etc., are obtained. In this way,
Carboxylic acids and primary amines react
alcohols (R-OH) and ethers (R-O-R) are de-
to form carboxylic acid amides (R-NH-CO-R ).
rived from water (H2O); primary amines (R-
The amino acid constituents of peptides and
NH2), secondary amines (R-NH-R) and terti-
proteins are linked by carboxylic acid amide
ary amines (R-N-R R ) amines are obtained
bonds, which are therefore also known as
from ammonia (NH3); and thiols (R-SH) and
peptide bonds (see p. 66).
thioethers (R-S-R ) arise from hydrogen sul-
fide (H2S). Polar groups such as -OH and -NH2
Phosphoric acid, H3PO4, is a tribasic (three-
are found as substituents in many organic
protic) acid—i. e., it contains three hydroxyl
compounds. As such groups are much more
groups able to donate H+ ions. At least one
reactive than the hydrocarbon structures to
of these three groups is fully dissociated
which they are attached, they are referred to
under normal physiological conditions, while
as functional groups.
the other two can react with alcohols. The
resulting products are phosphoric acid mono-
New functional groups can arise as a result
esters
(R-O-P(O)O-OH) and diesters
(R-O-
of oxidation of the compounds mentioned
P(O)O-O-R ). Phosphoric acid monoesters are
above. For example, the oxidation of a thiol
found in carbohydrate metabolism, for exam-
yields a disulfide (R-S-S-R). Double oxidation
ple
(see p. 36), whereas phosphoric acid
of a primary alcohol (R-CH2-OH) gives rise
diester bonds occur in phospholipids
(see
initially to an aldehyde (R-C(O)-H), and then
p. 50) and nucleic acids (see p. 82 ).
to a carboxylic acid (R-C(O)-OH). In contrast,
the oxidation of a secondary alcohol yields a
Compounds of one acid with another are
ketone (R-C(O)-R). The carbonyl group (C=O)
referred to as acid anhydrides. A particularly
is characteristic of aldehydes and ketones.
large amount of energy is required for the
formation of an acid—anhydride bond. Phos-
The addition of an amine to the carbonyl
phoric anhydride bonds therefore play a cen-
group of an aldehyde yields—after removal of
tral role in the storage and release of chemical
water—an aldimine (not shown; see p. 178).
energy in the cell (see p. 122). Mixed anhy-
Aldimines are intermediates in amino acid
drides between carboxylic acids and phos-
metabolism (see p. 178) and serve to bond
phoric acid are also very important “energy-
aldehydes to amino groups in proteins (see
rich metabolites” in cellular metabolism.
p. 62, for example). The addition of an alcohol
to the carbonyl group of an aldehyde yields a
hemiacetal (R-O-C(H)OH-R). The cyclic forms
of sugars are well-known examples of hemi-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Chemistry
11
A. Important classes of compounds
Carbonyl group
Oxygen
O
Nitrogen
N
Water
O
Amino group
H
H
O
R
H
H
H
C
H
N
N
R
R'
O
Secondary
Ketone
alcohol
H
H
Oxidation
R
C
R'
Primary
Ammonia
amine
H
H
Primary
O
R
R'
R
R'
Oxidation
alcohol
N
N
H
C
R'
H
R''
O
H
Aldehyde
Secondary
Tertiary
R
R'
Ether
amine
amine
C
O
H
R'
O
H
O
Oxidation
R
C
R
N
R'
O
O
C
R'
R
H
O P O
Hemiacetal
H
H
Carboxylic acid amide
O
Carboxyl group
O
Phosphoric
acid ester
H
C
O
O R'
O
R
C
Carboxylic acid
S
R'
R
C
O R'
Thioester
Carboxylic acid ester
Phosphorus
P
Sulfur
S
O
O
R
O
P
O
C
R'
H H
S
O
Hydrogen sulfide
O
Mixed anhydride
R
H
H
S
O
P
O
Sulfhydryl
H
Thiol
group
O
O
O
R
Dihydrogen phosphate
O
P
O
P
O
R
R'
H
S
S
O
O
Disulfide
Phosphoric acid anhydride
“energy-rich” bond
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
12
Basics
genic amine (see p. 62) formed by decarbox-
Biomolecules II
ylation of the amino acid cysteine.
Many biomolecules are made up of smaller
(2) The amino group of cysteamine is
units in a modular fashion, and they can be
bound to the carboxy group of another bio-
broken down into these units again. The con-
genic amine via an acid amide bond (-CO-
struction of these molecules usually takes
NH-). β-Alanine arises through decarboxyla-
place through condensation reactions involv-
tion of the amino acid aspartate, but it can
ing the removal of water. Conversely, their
also be formed by breakdown of pyrimidine
breakdown functions in a hydrolytic fash-
bases (see p. 186).
ion—i. e., as a result of water uptake. The
(3) Another acid amide bond (-CO-NH-)
page opposite illustrates this modular princi-
creates the compound for the next
ple using the example of an important coen-
constituent, pantoinate. This compound con-
zyme.
tains a chiral center and can therefore appear
in two enantiomeric forms (see p. 8). In natu-
ral coenzyme A, only one of the two forms is
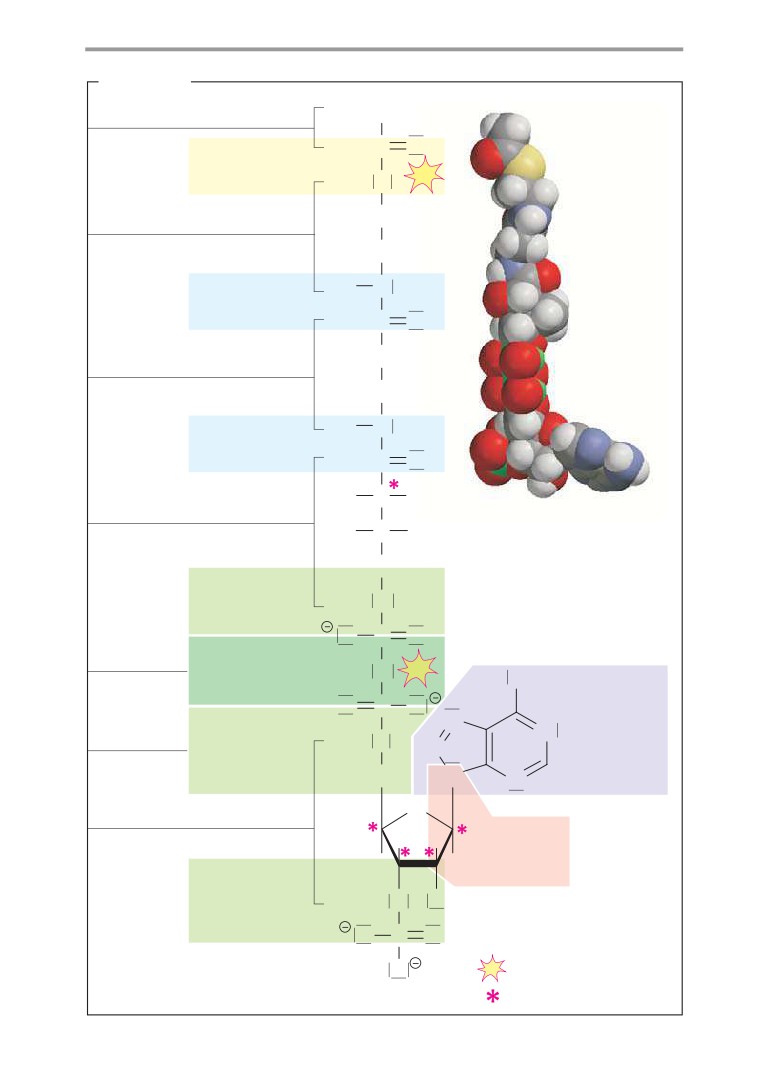
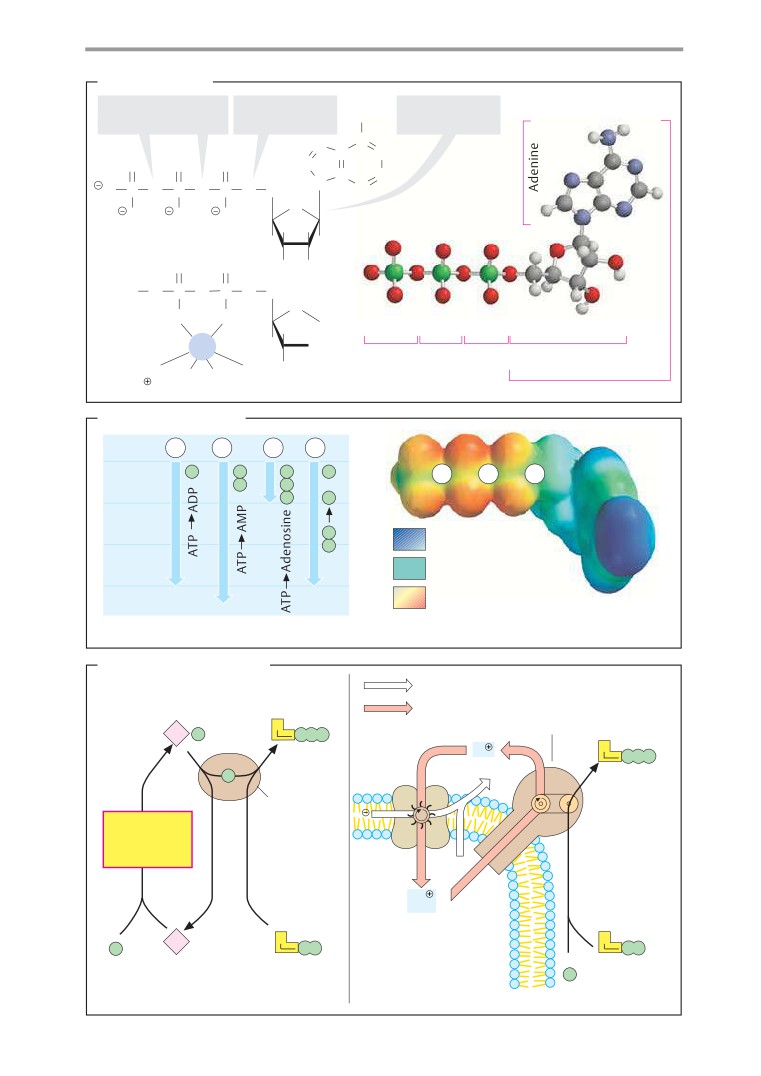
A. Acetyl CoA
found, the (R)-pantoinate. Human metabo-
Coenzyme A (see also p. 106) is a nucleotide
lism is not capable of producing pantoinate
with a complex structure (see p. 80). It serves
itself, and it therefore has to take up a
to activate residues of carboxylic acids (acyl
compound of β-alanine and pantoinate—
residues). Bonding of the carboxy group of the
pantothenate
(“pantothenic acid”)—in the
carboxylic acid with the thiol group of the
form of a vitamin in food (see p. 366).
coenzyme creates a thioester bond (-S-CO-R;
(4) The hydroxy group at C-4 of pantoinate
see p. 10) in which the acyl residue has a high
is bound to a phosphate residue by an ester
chemical potential. It can therefore be trans-
bond.
ferred to other molecules in exergonic reac-
The section of the molecule discussed so
tions. This fact plays an important role in lipid
far represents a functional unit. In the cell, it is
metabolism in particular (see pp. 162ff.), as
produced from pantothenate. The molecule
well as in two reactions of the tricarboxylic
also occurs in a protein-bound form as 4 -
acid cycle (see p. 136).
phosphopantetheine in the enzyme fatty
As discussed on p.16, the group transfer
acid synthase (see p. 168). In coenzyme A,
potential can be expressed quantitatively as
however, it is bound to 3 ,5 -adenosine di-
the change in free enthalpy (∆G) during hy-
phosphate.
drolysis of the compound concerned. This is
(5) When two phosphate residues bond,
an arbitrary determination, but it provides
they do not form an ester, but an “energy-
important indications of the chemical energy
rich” phosphoric acid anhydride bond, as
stored in such a group. In the case of acetyl-
also occurs in other nucleoside phosphates.
CoA, the reaction to be considered is:
By contrast, (6) and (7) are ester bonds again.
(8) The base adenine is bound to C-1 of
Acetyl CoA + H2O acetate + CoA
ribose by an N-glycosidic bond (see p. 36). In
addition to C-2 to C-4, C-1 of ribose also rep-
In standard conditions and at pH 7, the
resents a chiral center. The E-configuration is
change in the chemical potential G (∆G0, see
usually found in nucleotides.
p. 18) in this reaction amounts to -32 kJ
mol-1 and it is therefore as high as the ∆G0
of ATP hydrolysis (see p. 18). In addition to the
“energy-rich” thioester bond, acetyl-CoA also
has seven other hydrolyzable bonds with dif-
ferent degrees of stability. These bonds, and
the fragments that arise when they are hydro-
lyzed, will be discussed here in sequence.
(1) The reactive thiol group of coenzyme A
is located in the part of the molecule that is
derived from cysteamine. Cysteamine is a bio-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Chemistry
13
A. Acetyl CoA
CH
Acetate
3
C O
Thioester bond
S
CH2
Cysteamine
CH2
H
N
Acid-amide
bond
C O
CH2
β-Alanine
CH2
H
N
Acid-
amide bond
C O
H
C
OH
Pantoinate
H
C
C CH3
3
Van der Waals model
CH2
Phosphoric acid
ester bond
O
O
P O
Phosphate
Phosphoric acid
O
anhydride bond
NH
2
O
P O
N
N
Phosphate
Phosphoric acid
O
HC
ester bond
CH2
N
Adenine
N
Ribose
O
H
H
N-glycosidic bond
H
H
Phosphoric acid
ester bond
O
OH
O P O
O
Energy-rich bond
Phosphate
Chiral centers
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
14
Basics
hol (1b). The elimination of water from the
Chemical reactions
alcohol (2, dehydration) is also catalyzed by
Chemical reactions are processes in which
an acid and passes via the same intermediate
electrons or groups of atoms are taken up
as the addition reaction.
into molecules, exchanged between mole-
cules, or shifted within molecules. Illustrated
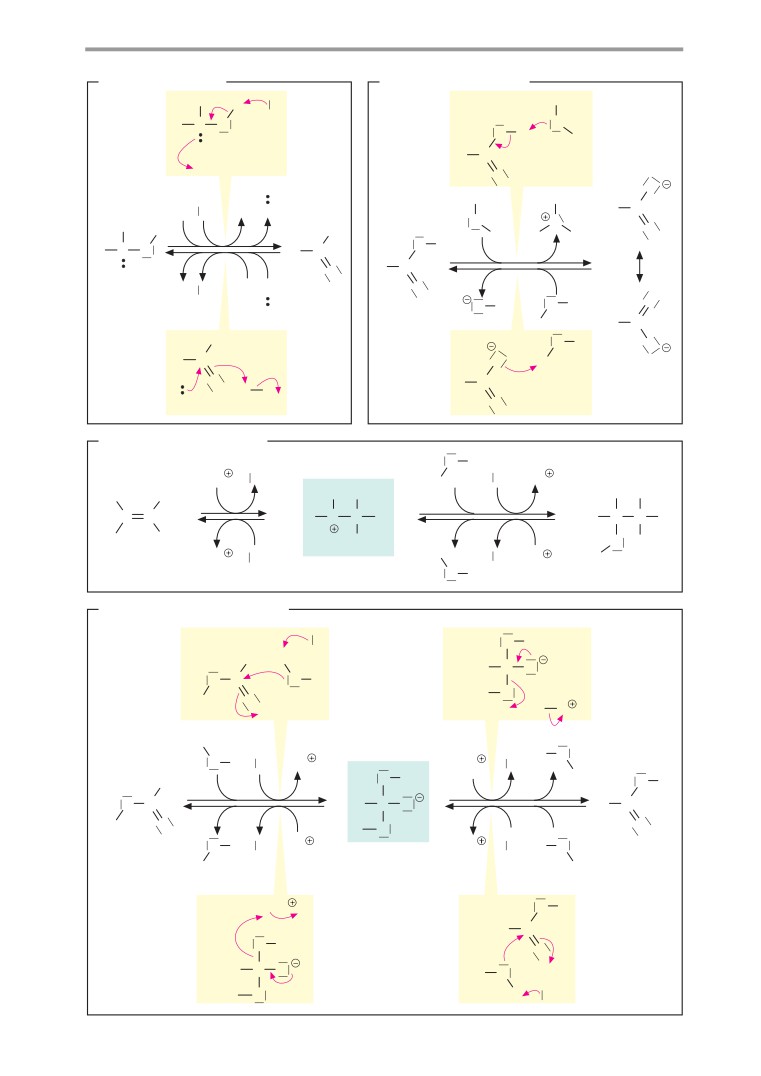
D. Nucleophilic substitutions
here are the most important types of reaction
in organic chemistry, using simple examples.
A reaction in which one functional group (see
Electron shifts are indicated by red arrows.
p. 10) is replaced by another is termed substi-
tution. Depending on the process involved, a
distinction is made between nucleophilic and
A. Redox reactions
electrophilic substitution reactions
(see
In redox reactions (see also p. 32), electrons
chemistry textbooks). Nucleophilic substitu-
are transferred from one molecule (the reduc-
tions start with the addition of one molecule
ing agent) to another (the oxidizing agent).
to another, followed by elimination of the so-
One or two protons are often also transferred
called leaving group.
in the process, but the decisive criterion for
The hydrolysis of an ester to alcohol and
the presence of a redox reaction is the elec-
acid (1) and the esterification of a carboxylic
tron transfer. The reducing agent is oxidized
acid with an alcohol (2) are shown here as an
during the reaction, and the oxidizing agent is
example of the SN2 mechanism. Both reac-
reduced.
tions are made easier by the marked polarity
Fig. A shows the oxidation of an alcohol
of the C=O double bond. In the form of ester
into an aldehyde (1) and the reduction of
hydrolysis shown here, a proton is removed
the aldehyde to alcohol (2). In the process,
from a water molecule by the catalytic effect
one hydride ion is transferred (two electrons
of the base B. The resulting strongly nucleo-
and one proton; see p. 32), which moves to
philic OH- ion attacks the positively charged
the oxidizing agent A in reaction 1. The super-
carbonyl C of the ester (1a), and an unstable
fluous proton is bound by the catalytic effect
sp3-hybridized transition state is produced.
of a base B. In the reduction of the aldehyde
From this, either water is eliminated (2b)
(2), A-H serves as the reducing agent and the
and the ester re-forms, or the alcohol ROH is
acid H-B is involved as the catalyst.
eliminated (1b) and the free acid results. In
esterification (2), the same steps take place in
reverse.
B. Acid-base reactions
In contrast to redox reactions, only proton
Further information
transfer takes place in acid-base reactions
(see also p. 30). When an acid dissociates (1),
In rearrangements
(isomerizations, not
water serves as a proton acceptor (i. e., as a
shown), groups are shifted within one and
base). Conversely, water has the function of
the same molecule. Examples of this in bio-
an acid in the protonation of a carboxylate
chemistry include the isomerization of sugar
anion (2).
phosphates (see p. 36) and of methylmalonyl-
CoA to succinyl CoA (see p. 166).
C. Additions/eliminations
A reaction in which atoms or molecules are
taken up by a multiple bond is described as
addition. The converse of addition—i. e., the
removal of groups with the formation of a
double bond, is termed elimination. When
water is added to an alkene (1a), a proton is
first transferred to the alkene. The unstable
carbenium cation that occurs as an intermedi-
ate initially takes up water (not shown), be-
fore the separation of a proton produces alco-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Chemistry
15
A. Redox reactions
B. Acid-base reactions
H
H HB
R
C
O
O
O H
H
H
R C
1
A
1
O
H
O
H
H
R C
A B H B A
O
O
H H
H
O
1
H
H
H
O
H
R
C
O
R C
1
2
R C
H
O
2
A B H B H
O
Alcohol
Aldehyde
O
H
O
O
H
A
Acid
H
R C
H
2
2
O
H
O
R C
O
H
Anion
H
O
R C
H B
A
O
C. Additions/eliminations
O
H
BH
B
B
BH
H
H
R
H
H
H H
1a
1b
C C
R C
C
R'
R
C C
R'
2a
2b
R'
H
H
O
H
H
BH B
B
BH
H
Alkene
Carbonium ion
Alcohol
O
H
D. Nucleophilic substitutions
1a
B
1b
O
H
R
H
R C
O
O C
O
H
R'
O
R'
O
H
B
H
Alcohol
O
O
H
B
BH
BH
B R'
O
H
H
R
1a
1b
O
H
O C
R C
O
R C
2b
2a
R'
O
R'
O
O
O
H
B
BH
BH
B R'
O
Ester
Transitional state
Carboxylic
H
H acid
Alcohol
2b
H B
2a
O H
R
C
O
H
O
R C O
R' O
R'
O
H
B
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
16
Basics
the amount of matter reacting
(in mol).
Energetics
Although absolute values for free enthalpy G
To obtain a better understanding of the pro-
cannot be determined, ∆G can be calculated
cesses involved in energy storage and conver-
from the equilibrium constant of the reaction
sion in living cells, it may be useful first to
(see p. 18).
recall the physical basis for these processes.
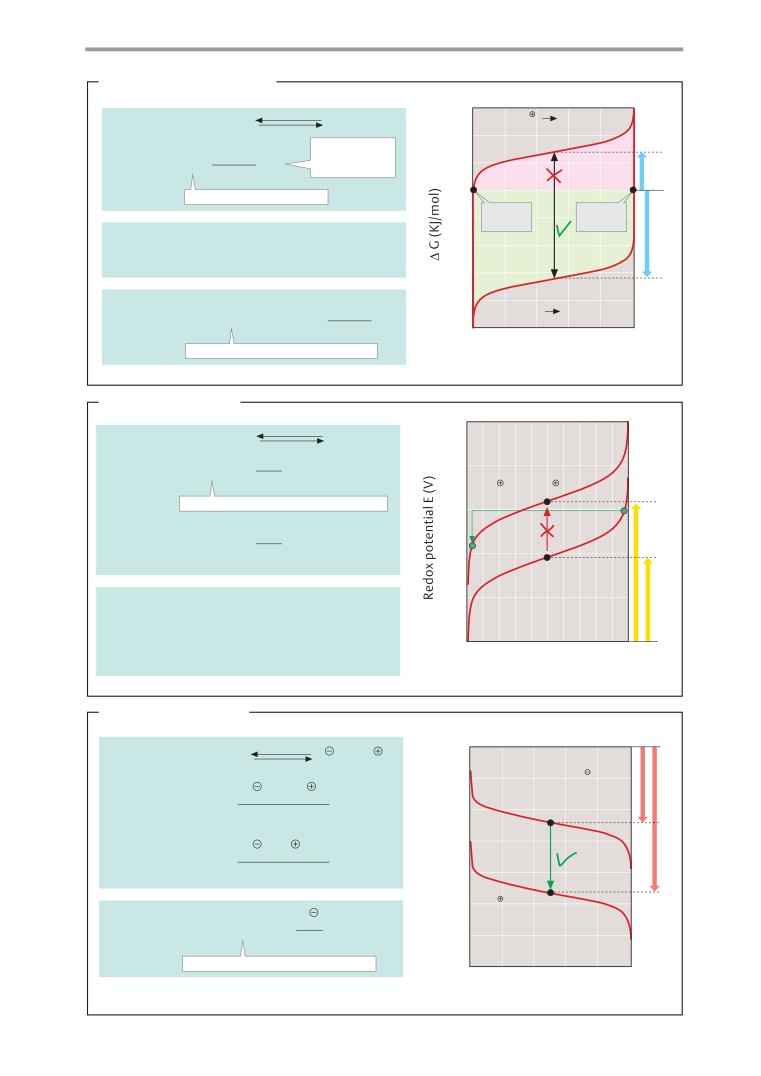
B. Energetics and the course of processes
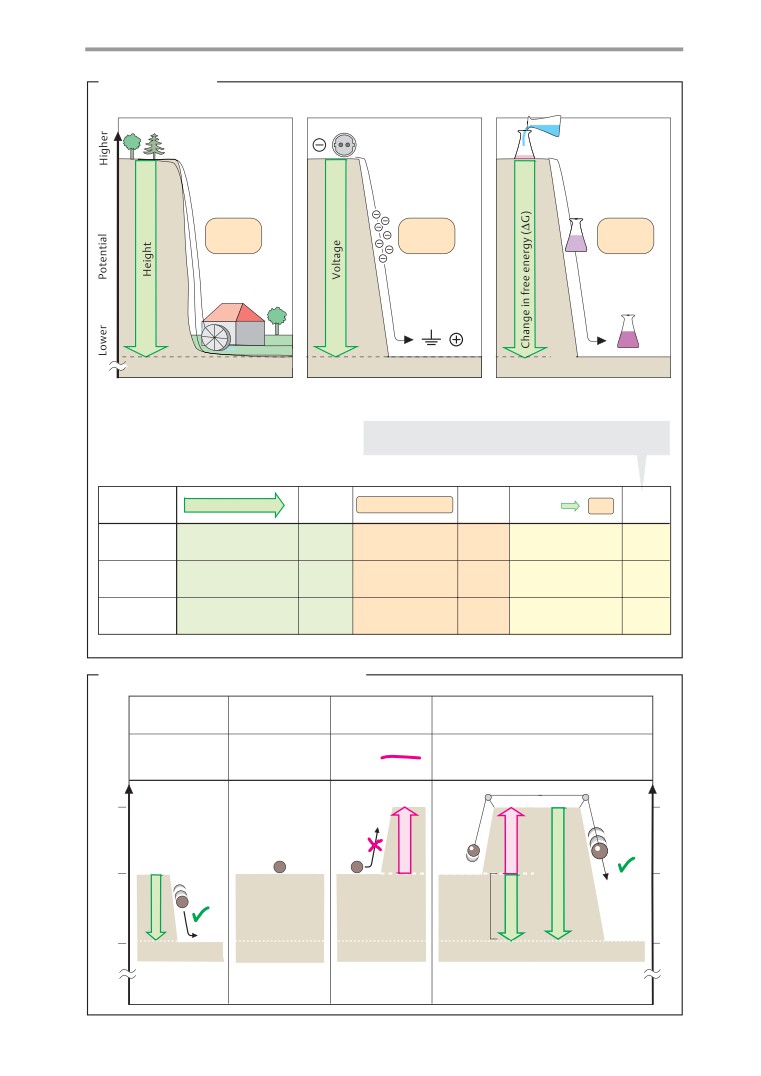
A. Forms of work
Everyday experience shows that water never
There is essentially no difference between
flows uphill spontaneously. Whether a partic-
work and energy. Both are measured in joule
ular process can occur spontaneously or not
(J = 1 N m). An outdated unit is the calorie
depends on whether the potential difference
(1 cal = 4.187 J). Energy is defined as the abil-
between the final and the initial state, ∆P =
ity of a system to perform work. There are
P2 - P1, is positive or negative. If P2 is smaller
many different forms of energy—e. g., me-
than P1, then ∆P will be negative, and the
chanical, chemical, and radiation energy.
process will take place and perform work.
A system is capable of performing work
Processes of this type are called exergonic
when matter is moving along a potential gra-
(B1). If there is no potential difference, then
dient. This abstract definition is best under-
the system is in equilibrium (B2). In the case of
stood by an example involving mechanical
endergonic processes,
∆P is positive
(B3).
work (A1). Due to the earth’s gravitational
Processes of this type do not proceed sponta-
pull, the mechanical potential energy of an
neously.
object is the greater the further the object is
Forcing endergonic processes to take place
away from the center of the earth. A potential
requires the use of the principle of energetic
difference
(∆P) therefore exists between a
coupling. This effect can be illustrated by a
higher location and a lower one. In a waterfall,
mechanical analogy (B4). When two masses
the water spontaneously follows this poten-
M1 and M2 are connected by a rope, M1 will
tial gradient and, in doing so, is able to per-
move upward even though this part of the
form work—e. g., turning a mill.
process is endergonic. The sum of the two
Work and energy consist of two quantities:
potential differences
(∆Peff = ∆P1 + ∆P2) is
an intensity factor, which is a measure of the
the determining factor in coupled processes.
potential difference—i. e., the “driving force”
When ∆Peff is negative, the entire process can
of the process—(here it is the height differ-
proceed.
ence) and a capacity factor, which is a mea-
Energetic coupling makes it possible to
sure of the quantity of the substance being
convert different forms of work and energy
transported
(here it is the weight of the
into one another. For example, in a flashlight,
water). In the case of electrical work (A2),
an exergonic chemical reaction provides an
the intensity factor is the voltage—i. e., the
electrical voltage that can then be used for
electrical potential difference between the
the endergonic generation of light energy. In
source of the electrical current and the
the luminescent organs of various animals, it
“ground,” while the capacity factor is the
is a chemical reaction that produces the light.
amount of charge that is flowing.
In the musculature (see p. 336), chemical en-
Chemical work and chemical energy are
ergy is converted into mechanical work and
defined in an analogous way. The intensity
heat energy. A form of storage for chemical
factor here is the chemical potential of a mol-
energy that is used in all forms of life is aden-
ecule or combination of molecules. This is
osine triphosphate (ATP; see p. 122). Ender-
stated as free enthalpy G
(also known as
gonic processes are usually driven by cou-
“Gibbs free energy”). When molecules spon-
pling to the strongly exergonic breakdown
taneously react with one another, the result is
of ATP (see p. 122).
products at lower potential. The difference in
the chemical potentials of the educts and
products (the change in free enthalpy, 'G) is
a measure of the “driving force” of the reac-
tion. The capacity factor in chemical work is
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
17
A. Forms of work
Elevated position
Voltage
Educts
source
Weight
Charge
Quantity
Lower position
Products
Ground
1. Mechanical work
2. Electrical work
3. Chemical work
J = Joule = N · m =1 kg · m2 · s-2, 1 cal = 4.187 J
Form of work
Intensity factor
Unit
Capacity factor
Unit
Work =
·
Unit
Mechanical
Height
m
Weight
J · m -1
Height · Weight
J
V =
Electrical
Voltage
Charge
C
Voltage · Charge
J
J · C -1
Free-enthalpy
Chemical
J · mol -1
Quantity
mol
∆G · Quantity
J
change ∆G
B. Energetics and the course of processes
∆P < 0
∆P = 0
∆P > 0
∆Peff < 0
Process occurs
Process cannot
Coupled processes can
spontaneously
occur
occur spontaneously
Potential
Potential
P3
P3
∆P
M1
∆P1
M2
P1
P1
∆P
∆ P
eff
∆P2
P2
P2
1. Exergonic
2. Equilibrium
3. Endergonic
4. Energetically coupled
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
18
Basics
redox potential of a system is, the higher the
Equilibriums
chemical potential of the transferred elec-
trons. To describe reactions between two re-
A. Group transfer reactions
dox systems, ∆Ε —the difference between the
Every chemical reaction reaches after a time a
two systems’ redox potentials—is usually
state of equilibrium in which the forward and
used instead of ∆G. ∆G and ∆E have a simple
back reactions proceed at the same speed. The
relationship, but opposite signs (below). A
law of mass action describes the concentra-
redox reaction proceeds spontaneously
tions of the educts (A, B) and products (C, D) in
when ∆E > 0, i. e. ∆G < 0.
equilibrium. The equilibrium constant K is di-
The right side of the illustration shows the
rectly related to
∆G0, the change in free
way in which the redox potential E is depen-
enthalpy G involved in the reaction
(see
dent on the composition (the proportion of
p. 16) under standard conditions (∆G0 = - R
the reduced form as a %) in two biochemically
T ln K). For any given concentrations, the
important redox systems
(pyruvate/lactate
lower equation applies. At ∆G < 0, the reac-
and NAD+/NADH+H+; see pp. 98, 104). In the
tion proceeds spontaneously for as long as it
standard state (both systems reduced to 50%),
takes for equilibrium to be reached (i. e., until
electron transfer from lactate to NAD+ is not
∆G = 0). At ∆G > 0, a spontaneous reaction is
possible, because ∆E is negative (∆E = -0.13 V,
no longer possible
(endergonic case; see
red arrow). By contrast, transfer can proceed
p. 16). In biochemistry, ∆G is usually related
successfully if the pyruvate/lactate system is
to pH 7, and this is indicated by the “prime”
reduced to 98 % and NAD+/NADH is 98% oxi-
symbol (∆G0 or ∆G ).
dized (green arrow, ∆E = +0.08 V).
As examples, we can look at two group
transfer reactions (on the right). In ATP (see
C. Acid-base reactions
p. 122), the terminal phosphate residue is at a
high chemical potential. Its transfer to water
Pairs of conjugated acids and bases are always
(reaction a, below) is therefore strongly exer-
involved in proton exchange reactions (see
gonic. The equilibrium of the reaction
p. 30). The dissociation state of an acid-base
(∆G = 0; see p. 122) is only reached when
pair depends on the H+ concentration. Usu-
more than 99.9% of the originally available
ally, it is not this concentration itself that is
ATP has been hydrolyzed. ATP and similar
expressed, but its negative decadic logarithm,
compounds have a high group transfer
the pH value. The connection between the pH
potential for phosphate residues. Quantita-
value and the dissociation state is described
tively, this is expressed as the 'G of hydrolysis
by the Henderson-Hasselbalch equation (be-
(∆G0
= -32 kJ mol-1; see p. 122).
low). As a measure of the proton transfer
In contrast, the endergonic transfer of am-
potential of an acid-base pair, its pKa value
monia (NH3) to glutamate (Glu, reaction b,
is used—the negative logarithm of the acid
∆G0 = +14 kJ mol-1) reaches equilibrium so
constant Ka (where “a” stands for acid).
quickly that only minimal amounts of the
The stronger an acid is, the lower its pKa
product glutamine (Gln) can be formed in
value. The acid of the pair with the lower pKa
this way. The synthesis of glutamine from
value (the stronger acid—in this case acetic
these preliminary stages is only possible
acid, CH3COOH) can protonate (green arrow)
through energetic coupling (see pp. 16, 124).
the base of the pair with the higher pKa (in
this case NH3), while ammonium acetate
(NH4+ and CH3COO-) only forms very little
B. Redox reactions
CH3COOH and NH3.
The course of electron transfer reactions (re-
dox reactions, see p. 14) also follows the law of
mass action. For a single redox system (see
p. 32), the Nernst equation applies (top). The
electron transfer potential of a redox system
(i. e., its tendency to give off or take up elec-
trons) is given by its redox potential E (in
standard conditions, E0 or E0 ). The lower the
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
19
A. Group transfer reactions
30
Glu + NH4
Gln + H2O
Reaction
A + B
C + D
20
Only applies
Law of
[C] · [D]
in chemical
mass action
K=
10
equilibrium
∆G°(b)
[A] · [B]
0
Equilibrium constant
Equi-
Equi-
-10
librium
librium
∆G°(a)
Relationship
∆G° = - R · T · ln K
-20
between
∆G0 and K
R = 8.314 J · mol -1 · K-1
-30
In any conditions
-40
[C] · [D]
ATP + H2O
ADP + Pi
∆G = ∆G° + R · T · ln
-50
[A] · [B]
0
20
40
60
80
100
Measure of group transfer potential
% converted
B. Redox reactions
- 0.5
Ared
Aox
For a redox
R · T
- 0.4
system
E = E° +
· ln[Aox]
a
n · F
[Ared]
NAD
/NADH+H
Measure of electron transfer potential
- 0.3
For any redox
∆E° (a)
R · T
b
reaction
∆E = ∆E° +
· ln[Box] ·[Ared]
n · F
[Bred] · [Aox]
-0.2
- 0.1
∆Eº (b)
Definition
∆E = EAcceptor - EDonor
and sizes
∆G = - n · F · ∆E
Pyruvate/lactate
0.0
n = No. of electrons transferred
F = Faraday constant
0
20
40
60
80
100
% reduced
C. Acid-base reactions
0
Standard
HA + H2O
A + H3O
reaction
CH3
COOH/CH3COO
2
pKa(a)
Law of mass
[A
] · [H3O
]
action
K =
4
[HA] · [H2O]
6
[A
] · [H
]
a
pH
pKa(b)
Simplified
Ka=
8
[HA]
10
NH4
/NH3
Henderson-
[A
]
b
Hasselbalch
pH = pKa + log
12
equation
[HA]
Measure of proton transfer potential
14
0
20
40
60
80
100
% dissociated
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
20
Basics
this is the normal state of affairs—∆S is pos-
Enthalpy and entropy
itive for this process. An increase in the order
The change in the free enthalpy of a chemical
in a system (∆S < 0) always requires an input
reaction (i. e., its ∆G) depends on a number of
of energy. Both of these statements are
factors—e. g., the concentrations of the reac-
consequences of an important natural law,
tants and the temperature (see p. 18). Two
the Second Law of Thermodynamics. The
further factors associated with molecular
connection between changes in enthalpy
changes occurring during the reaction are dis-
and entropy is described quantitatively by
cussed here.
the Gibbs-Helmholtz equation
(∆G = ∆H -
T
∆S). The following examples will help
explain these relationships.
A. Heat of reaction and calorimetry
In the knall-gas
(oxyhydrogen) reaction
All chemical reactions involve heat exchange.
(1), gaseous oxygen and gaseous hydrogen
Reactions that release heat are called
react to form liquid water. Like many redox
exothermic, and those that consume heat
reactions, this reaction is strongly exothermic
are called endothermic. Heat exchange is
(i. e.,
∆H < 0). However, during the reaction,
measured as the enthalpy change ∆H (the
the degree of order increases. The total num-
heat of reaction). This corresponds to the
ber of molecules is reduced by one-third, and
heat exchange at constant pressure. In exo-
a more highly ordered liquid is formed from
thermic reactions, the system loses heat, and
freely moving gas molecules. As a result of the
∆H is negative. When the reaction is endo-
increase in the degree of order (∆S < 0), the
thermic, the system gains heat, and ∆H be-
term -T
∆S becomes positive. However, this
comes positive.
is more than compensated for by the decrease
In many reactions, ∆H and ∆G are similar in
in enthalpy, and the reaction is still strongly
magnitude (see B1, for example). This fact is
exergonic (∆G < 0).
used to estimate the caloric content of foods.
The dissolution of salt in water (2) is endo-
In living organisms, nutrients are usually oxi-
thermic (∆H > 0)—i. e., the liquid cools. Never-
dized by oxygen to CO2 and H2O (see p. 112).
theless, the process still occurs spontane-
The maximum amount of chemical work sup-
ously, since the degree of order in the
plied by a particular foodstuff (i. e., the ∆G for
system decreases. The Na+ and Cl- ions are
the oxidation of the utilizable constituents)
initially rigidly fixed in a crystal lattice. In
can be estimated by burning a weighed
solution, they move about independently
amount in a calorimeter in an oxygen atmo-
and in random directions through the fluid.
sphere. The heat of the reaction increases the
The decrease in order (∆S > 0) leads to a
water temperature in the calorimeter. The
negative
-T
∆S term, which compensates
reaction heat can then be calculated from
for the positive
∆H term and results in a
the temperature difference ∆T.
negative ∆G term overall. Processes of this
type are described as being entropy-driven.
The folding of proteins (see p. 74) and the
B. Enthalpy and entropy
formation of ordered lipid structures in water
The reaction enthalpy ∆H and the change in
(see p. 28) are also mainly entropy-driven.
free enthalpy ∆G are not always of the same
magnitude. There are even reactions that oc-
cur spontaneously (∆G < 0) even though they
are endothermic (∆H > 0). The reason for this
is that changes in the degree of order of the
system also strongly affect the progress of a
reaction. This change is measured as the en-
tropy change ('S).
Entropy is a physical value that describes
the degree of order of a system. The lower the
degree of order, the larger the entropy. Thus,
when a process leads to increase in disor-
der—and everyday experience shows that
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
21
A. Heat of reaction and calorimetry
Thermometer
Ignition wire
to start the reaction
65
65
43
43
2
2
1
1
Temperature
12
12
34
34
insulation
5
Water
5
Water
Water
6
Combustion
6
heated
Pressurized
O2
CO2
metal
container
An enthalpy of
H2O
1kJ warms 1 l of water
Sample
by 0.24 ºC
Stirrer
B. Enthalpy and entropy
∆H: change
∆S: change of
∆G =
∆H
- T ·
∆S
of enthalpy,
entropy, i.e.
heat exchange
degree of order
Gibbs-Helmholtz equation
1 mol H2
1 mol NaCl
1/2 mol O2
(crystalline)
Low degree
High degree
of order
of order
System re-
System
leases heat,
absorbs
∆H <0
heat,
(exothermic)
∆H > 0
(endothermic)
1 mol H2O
1 mol Na
(liquid)
1 mol Cl
Lower
Higher degree
degree
of order,
of order
∆S < 0
∆S > 0
∆H = - 287 kJ · mol -1
∆H = +3.8 kJ · mol -1
-T · ∆S = +49 kJ · mol -1
-T · ∆S = - 12.8 kJ · mol -1
∆G = - 238 kJ · mol -1
∆G = - 9.0 kJ · mol -1
-200
-100
0
+100
+200
-12
-8
-4
0
+4
+8
+12
Energy
Energy
1. “Knall-gas” reaction
2. Dissolution of NaCl in water
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
22
Basics
doubles with an increase in temperature of
Reaction kinetics
10 °C.
The change in free enthalpy ∆G in a reaction
indicates whether or not the reaction can take
B. Reaction rate
place spontaneously in given conditions and
how much work it can perform (see p. 18).
The velocity v of a chemical reaction is deter-
However, it does not tell us anything about
mined experimentally by observing the
the rate of the reaction—i. e., its kinetics.
change in the concentration of an educt or
product over time. In the example shown
(again a reaction of the A B type), 3 mmol
A. Activation energy
of the educt A is converted per second and
Most organic chemical reactions (with the
3 mmol of the product B is formed per second
exception of acid-base reactions) proceed
in one liter of the solution. This corresponds
only very slowly, regardless of the value
to a rate of
of
∆G. The reason for the slow reaction rate
is that the molecules that react—the
v = 3 mM s-1 = 3
10-3 mol L-1 s-1
educts—have to have a certain minimum en-
ergy before they can enter the reaction. This is
C. Reaction order
best understood with the help of an energy
diagram (1) of the simplest possible reaction
Reaction rates are influenced not only by the
A B. The educt A and the product B are each
activation energy and the temperature, but
at a specific chemical potential (Ge and Gp,
also by the concentrations of the reactants.
respectively). The change in the free enthalpy
When there is only one educt, A (1), v is
of the reaction, ∆G, corresponds to the differ-
proportional to the concentration [A] of this
ence between these two potentials. To be
substance, and a first-order reaction is in-
converted into B, A first has to overcome a
volved. When two educts, A and B, react
potential energy barrier, the peak of which,
with one another (2), it is a second order
Ga, lies well above Ge. The potential difference
reaction (shown on the right). In this case,
Ga -Ge is the activation energy Ea of the re-
the rate v is proportional to the product of
action (in kJ mol-1).
the educt concentrations
(12 mM2 at the
The fact that A can be converted into B at all
top, 24 mM2 in the middle, and 36 mM2 at
is because the potential Ge only represents
the bottom). The proportionality factors k and
the average potential of all the molecules.
k are the rate constants of the reaction. They
Individual molecules may occasionally reach
are not dependent on the reaction concentra-
much higher potentials—e. g., due to collisions
tions, but depend on the external conditions
with other molecules. When the increase in
for the reaction, such as temperature.
energy thus gained is greater than Ea, these
In B, only the kinetics of simple irreversible
molecules can overcome the barrier and be
reactions is shown. More complicated cases,
converted into B. The energy distribution for a
such as reaction with three or more reversible
group of molecules of this type, as calculated
steps, can usually be broken down into first-
from a simple model, is shown in (2) and (3).
order or second-order partial reactions and
∆n/n is the fraction of molecules that have
described using the corresponding equations
reached or exceeded energy E (in kJ per mol).
(for an example, see the Michaelis-Menten
At 27 °C, for example, approximately 10% of
reaction, p. 92).
the molecules have energies > 6 kJ mol-1.
The typical activation energies of chemical
reactions are much higher. The course of
the energy function at energies of around
50 kJ mol-1 is shown in (3). Statistically, at
27 °C only two out of 109 molecules reach this
energy. At 37 °C, the figure is already four.
This is the basis for the long-familiar “Q10
law”—a rule of thumb that states that the
speed of biological processes approximately
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
23
A. Activation energy
10
55
Chemical
potential
8
Ga
6
Activation energy
50
Substrate A
37˚C
=
Ea
Ga - Ge
4
27 ˚C
Ge
2
27˚C
∆G = G
Product B
p - Ge
Gp
0
45
0.0
0.5
1.0
0
5
10
∆n/n
∆n/n · 109
1.
2.
3.
B. Reaction rate
[A]
= 32 mM
[A] = 29 mM
[A] = 23 mM
0
∆[A] = -3 mM
∆[A] = -9 mM
0 s
1 s
3 s
∆t = 1 s
∆t = 3 s
[B]
= 3 mM
[B] = 6 mM
[B] = 12 mM
0
∆[B] = 3 mM
∆[B] =
9 mM
mM =
1.
2.
3.
mmol · l-1
v
= -∆ [A] / ∆t = ∆ [ B] / ∆t
( mol · l -1 · s -1 )
C. Reaction order
1 Liter
A
C
A
+ B
C
0 s
1 s
0 s
1 s
[A] (mM)
v (mM · s-1)
(mM)
v (mM · s-1)
[A]
= 12
˚
5
1
[B]
= 1
1
˚
[A]
= 6
˚
10
2
[B]
= 4
2
˚
[A]
= 3
˚
1.
15
3
[B]
= 12
3
2.
˚
v = k · [A]
k, k' : Rate constants
v = k' · [A] · [B]
First-order reaction
k = 1/5 s -1
Second-order reaction
k' = 1/12 l · mmol-1· s -1
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
24
Basics
B. Catalysis of H2O2 - breakdown by iodide
Catalysis
As a simple example of a catalyzed reaction,
Catalysts are substances that accelerate
we can look at the disproportionation of hy-
chemical reactions without themselves being
drogen peroxide
(H2O2) into oxygen and
consumed in the process. Since catalysts
water. In the uncatalyzed reaction (at the
emerge from the catalyzed reaction without
top), an H2O2 molecule initially decays into
being changed, even small amounts are usu-
H2O and atomic oxygen (O), which then reacts
ally suf cient to cause a powerful acceleration
with a second H2O2 molecule to form water
of the reaction. In the cell, enzymes (see p. 88)
and molecular oxygen (O2). The activation
generally serve as catalysts. A few chemical
energy Ea required for this reaction is rela-
changes are catalyzed by special RNA mole-
tively high, at 75 kJ mol-1. In the presence of
cules, known as ribozymes (see p. 246).
iodide (I-) as a catalyst, the reaction takes a
different course (bottom). The intermediate
A. Catalysis: principle
arising in this case is hypoiodide (OI-), which
also forms H2O and O2 with another H2O2
The reason for the slow rates of most reac-
molecule. In this step, the I- ion is released
tions involving organic substances is the high
and can once again take part in the reaction.
activation energy (see p. 22) that the reacting
The lower activation energy of the reaction
molecules have to reach before they can react.
catalyzed by iodide
(Ea = 56 kJ mol-1)
In aqueous solution, a large proportion of the
causes acceleration of the reaction by a factor
activation energy is required to remove the
of 2000, as the reaction rate depends expo-
hydration shells surrounding the educts. Dur-
nentially on Ea (v ~ e-Ea/R T).
ing the course of a reaction, resonance-stabi-
Free metal ions such as iron (Fe) and plat-
lized structures (see p. 4) are often tempora-
inum (Pt) are also effective catalysts for the
rily suspended; this also requires energy. The
breakdown of H2O2. Catalase (see p. 284), an
highest point on the reaction coordinates cor-
enzyme that protects cells against the toxic
responds to an energetically unfavorable tran-
effects of hydrogen peroxide (see p. 284), is
sition state of this type (1).
much more catalytically effective still. In the
A catalyst creates a new pathway for the
enzyme-catalyzed disproportionation, H2O2
reaction (2). When all of the transition states
is bound to the enzyme’s heme group, where
arising have a lower activation energy than
it is quickly converted to atomic oxygen and
that of the uncatalyzed reaction, the reaction
water, supported by amino acid residues of
will proceed more rapidly along the alterna-
the enzyme protein. The oxygen atom is tem-
tive pathway, even when the number of in-
porarily bound to the central iron atom of the
termediates is greater. Since the starting
heme group, and then transferred from there
points and end points are the same in both
to the second H2O2 molecule. The activation
routes, the change in the enthalpy ∆G of the
energy of the enzyme-catalyzed reaction is
reaction is not influenced by the catalyst. Cat-
only 23 kJ mol-1, which in comparison with
alysts—including enzymes—are in principle
the uncatalyzed reaction leads to acceleration
not capable of altering the equilibrium state
by a factor of 1.3
109.
of the catalyzed reaction.
Catalase is one of the most ef cient en-
The often-heard statement that “a catalyst
zymes there are. A single molecule can con-
reduces the activation energy of a reaction” is
vert up to 108 (a hundred million) H2O2 mol-
not strictly correct, since a completely different
ecules per second.
reaction takes place in the presence of a cata-
lyst than in uncatalyzed conditions. However,
its activation energy is lower than in the un-
catalyzed reaction.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
25
A. Catalysis: principle
Substrates
Products
Substrates
Products
∆G
Ea
∆G
Ea1
Ea2
1. Energy profile without catalyst
2. Energy profile with catalyst
B. Catalysis of H2O2 - breakdown by iodide
H2O2
+
H2O2
O2
+
H2O
+
H2O
1. Breakdown of hydrogen peroxide
H2O
Atomic oxygen
H2O2
1b
1a
O2
Catalyst (iodide)
H2O
2
H2O
O2
2a
2b
H2O
Hypoiodide
H2O2
2. Catalyzed reaction
Relative velocity
1300000000
Uncatalyzed
Iodide
Catalase
100
H2O2
Ea
80
2100
Ea
60
Heme
40
Ea
20
1
3. Activation energies
Active center of catalase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
26
Basics
of water clusters increases until the water
Water as a solvent
begins to crystallize. Under normal atmo-
Life as we know it evolved in water and is still
spheric pressure, this occurs at 0 °C. In ice,
absolutely dependent on it. The properties of
most of the water molecules are fixed in a
water are therefore of fundamental impor-
hexagonal lattice (3). Since the distance be-
tance to all living things.
tween the individual molecules in the frozen
state is on average greater than in the liquid
state, the density of ice is lower than that of
A. Water and methane
liquid water. This fact is of immense biological
The special properties of water (H2O) become
importance—it means, for example, that in
apparent when it is compared with methane
winter, ice forms on the surface of open
(CH4). The two molecules have a similar mass
stretches of water first, and the water rarely
and size. Nevertheless, the boiling point of
freezes to the bottom.
water is more than 250 °C above that of
methane. At temperatures on the earth’s sur-
C. Hydration
face, water is liquid, whereas methane is gas-
eous. The high boiling point of water results
In contrast to most other liquids, water is an
from its high vaporization enthalpy, which in
excellent solvent for ions. In the electrical
turn is due to the fact that the density of the
field of cations and anions, the dipolar water
electrons within the molecule is unevenly
molecules arrange themselves in a regular
distributed. Two corners of the tetrahedrally-
fashion corresponding to the charge of the
shaped water molecule are occupied by un-
ion. They form hydration shells and shield
shared electrons (green), and the other two
the central ion from oppositely charged ions.
by hydrogen atoms. As a result, the H-O-H
Metal ions are therefore often present as
bond has an angled shape. In addition, the
hexahydrates ([Me(H2O)62+], on the right). In
O-H bonds are polarized due to the high elec-
the inner hydration sphere of this type of ion,
tronegativity of oxygen (see p. 6). One side of
the water molecules are practically immobi-
the molecule carries a partial charge (δ) of
lized and follow the central ion. Water has a
about -0.6 units, whereas the other is corre-
high dielectric constant of 78—i. e., the elec-
spondingly positively charged. The spatial
trostatic attraction force between ions is re-
separation of the positive and negative
duced to 1/78 by the solvent. Electrically
charges gives the molecule the properties of
charged groups in organic molecules (e. g.,
an electrical dipole. Water molecules are
carboxylate, phosphate, and ammonium
therefore attracted to one another like tiny
groups) are also well hydrated and contribute
magnets, and are also connected by hydrogen
to water solubility. Neutral molecules with
bonds (B) (see p. 6). When liquid water vapor-
several hydroxy groups, such as glycerol (on
izes, a large amount of energy has to be ex-
the left) or sugars, are also easily soluble,
pended to disrupt these interactions. By con-
because they can form H bonds with water
trast, methane molecules are not dipolar, and
molecules. The higher the proportion of polar
therefore interact with one another only
functional groups there is in a molecule, the
weakly. This is why liquid methane vaporizes
more water-soluble (hydrophilic) it is. By con-
at very low temperatures.
trast, molecules that consist exclusively or
mainly of hydrocarbons are poorly soluble or
insoluble in water. These compounds are
B. Structure of water and ice
called hydrophobic (see p. 28).
The dipolar nature of water molecules favors
the formation of hydrogen bonds (see p. 6).
Each molecule can act either as a donor or an
acceptor of H bonds, and many molecules in
liquid water are therefore connected by H
bonds (1). The bonds are in a state of constant
fluctuation. Tetrahedral networks of mole-
cules, known as water “clusters,” often arise.
As the temperature decreases, the proportion
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
27
A. Water and methane
δ
+0.3
H2O
CH
H
4
H
18 Da
Molecular mass
16 Da
+100 °C
Boiling point
-162 °C
Heat of
H
41
vaporization
8
(kJ · mol-1)
H
H
δ
Dipole moment
δ
6.2
0
H
+0.3
(10-30 C · m)
-0.6
Water (H2O)
Methane (CH4)
B. Structure of water and ice
Liquid water
density 1.00 g · cm-3
short-lived clusters
2.
3.
Ethanol
Ice
Density 0.92 g · cm-3
hexagonal lattice,
stabilized by
1.
hydrogen bonds
C. Hydration
Cation
H
O
HO
HO
Glycerol
Anion
[Me (H2
O)6]2
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
28
Basics
is therefore positive (the disorder in the water
Hydrophobic interactions
increases), and the negative term
-T
∆S
Water is an excellent solvent for ions and for
makes the separation process exergonic
substances that contain polarized bonds (see
(∆G < 0), so that it proceeds spontaneously.
p. 20). Substances of this type are referred to
as polar or hydrophilic (“water-loving”). In
C. Arrangements of amphipathic substances
contrast, substances that consist mainly of
in water
hydrocarbon structures dissolve only poorly
in water. Such substances are said to be apolar
Molecules that contain both polar and apolar
or hydrophobic.
groups are called amphipathic or amphiphilic.
This group includes soaps (see p. 48), phos-
pholipids (see p. 50), and bile acids (see p. 56).
A. Solubility of methane
As a result of the “oil drop effect” amphi-
To understand the reasons for the poor water
pathic substances in water tend to arrange
solubility of hydrocarbons, it is useful first to
themselves in such a way as to minimize the
examine the energetics (see p. 16) of the pro-
area of surface contact between the apolar
cesses involved. In (1), the individual terms of
regions of the molecule and water. On water
the Gibbs-Helmholtz equation (see p. 20) for
surfaces, they usually form single-layer films
the simplest compound of this type, methane,
(top) in which the polar “head groups” face
are shown (see p. 4). As can be seen, the tran-
toward the water. Soap bubbles (right) consist
sition from gaseous methane to water is ac-
of double films, with a thin layer of water
tually exothermic (∆H0 < 0). Nevertheless, the
enclosed between them. In water, depending
change in the free enthalpy ∆G0 is positive
on their concentration, amphipathic com-
(the process is endergonic), because the en-
pounds form micelles—i. e., spherical aggre-
tropy term T
∆S0
has a strongly positive
gates with their head groups facing toward
value. The entropy change in the process
the outside, or extended bilayered double
(∆S0) is evidently negative—i. e., a solution of
membranes. Most biological membranes are
methane in water has a higher degree of order
assembled according to this principle (see
than either water or gaseous methane. One
p. 214). Closed hollow membrane sacs are
reason for this is that the methane molecules
known as vesicles. This type of structure
are less mobile when surrounded by water.
serves to transport substances within cells
More importantly, however, the water around
and in the blood (see p. 278).
the apolar molecules forms cage-like “clath-
The separation of oil and water (B) can be
rate” structures, which—as in ice—are stabi-
prevented by adding a strongly amphipathic
lized by H bonds. This strongly increases the
substance. During shaking, a more or less
degree of order in the water—and the more so
stable emulsion then forms, in which the sur-
the larger the area of surface contact between
face of the oil drops is occupied by amphi-
the water and the apolar phase.
pathic molecules that provide it with polar
properties externally. The emulsification of
fats in food by bile acids and phospholipids
B. The “oil drop effect”
is a vital precondition for the digestion of fats
The spontaneous separation of oil and water,
(see p. 314).
a familiar observation in everyday life, is due
to the energetically unfavorable formation of
clathrate structures. When a mixture of water
and oil is firmly shaken, lots of tiny oil drops
form to begin with, but these quickly coalesce
spontaneously to form larger drops—the two
phases separate. A larger drop has a smaller
surface area than several small drops with the
same volume. Separation therefore reduces
the area of surface contact between the water
and the oil, and consequently also the extent
of clathrate formation. The ∆S for this process
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
29
A. Solubility of methane
B. The “oil drop effect”
-T · ∆S0 =
+39.6 kJ · mol-1
Oil
10 x 1 mL
Total surface
area: 48 cm2
0
Methane
∆G0 =
+26.4 kJ · mol-1
∆H0=
-13.2 kJ · mol-1
∆S > 0
Spontaneus
-T · ∆S < 0
separation
∆G < 0
1 x 10 mL
Clathrate
Surface area:
structure
22 cm2
C. Arrangements of amphipathic substances in water
Air
Surface film
Water
Air
Micelle
Soap bubble
Vesicle
Clathrate
Air
structure
Double membrane
4 - 5 nm
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
30
Basics
Acids and bases
B. pH values in the organism
pH values in the cell and in the extracellular
A. Acids and bases
fluid are kept constant within narrow limits.
In general, acids are defined as substances
In the blood, the pH value normally ranges
that can donate hydrogen ions
(protons),
only between 7.35 and 7.45 (see p. 288). This
while bases are compounds that accept pro-
corresponds to a maximum change in the H+
tons.
concentration of ca. 30%. The pH value of
Water enhances the acidic or basic proper-
cytoplasm is slightly lower than that of blood,
ties of dissolved substances, as water itself
at
7.0-7.3. In lysosomes
(see p. 234; pH
can act as either an acid or a base. For exam-
4.5-5.5), the H+ concentration is several hun-
ple, when hydrogen chloride (HCl) is in aque-
dred times higher than in the cytoplasm. In
ous solution, it donates protons to the solvent
the lumen of the gastrointestinal tract, which
(1). This results in the formation of chloride
forms part of the outside world relative to the
ions (Cl-) and protonated water molecules
organism, and in the body’s excretion prod-
(hydronium ions, H3O+, usually simply re-
ucts, the pH values are more variable. Ex-
ferred to as H+). The proton exchange be-
treme values are found in the stomach (ca.
tween HCl and water is virtually quantitative:
2) and in the small bowel (> 8). Since the
in water, HCl behaves as a very strong acid
kidney can excrete either acids or bases, de-
with a negative pKa value (see p. 18).
pending on the state of the metabolism, the
Bases such as ammonia (NH3) take over
pH of urine has a particularly wide range of
protons from water molecules. As a result of
variation (4.8-7.5).
this, hydroxyl ions
(OH-) and positively
charged ammonium ions (NH4+, 3) form. Hy-
C. Buffers
dronium and hydroxyl ions, like other ions,
exist in water in hydrated rather than free
Short-term pH changes in the organism are
form (see p. 26).
cushioned by buffer systems. These are mix-
Acid-base reactions always involve pairs of
tures of a weak acid, HB, with its conjugate
acids and the associated conjugated bases
base, B-, or of a weak base with its conjugate
(see p. 18). The stronger the acid or base, the
acid. This type of system can neutralize both
weaker the conjugate base or acid, respec-
hydronium ions and hydroxyl ions.
tively. For example, the very strongly acidic
In the first case (left), the base (B-) binds a
hydrogen chloride belongs to the very weakly
large proportion of the added protons (H+)
basic chloride ion (1). The weakly acidic am-
and HB and water are formed. If hydroxyl
monium ion is conjugated with the moder-
ions (OH-) are added, they react with HB to
ately strong base ammonia (3).
give B- and water (right). In both cases, it is
The equilibrium constant K for the acid—
primarily the [HB]/[B-] ratio that shifts, while
base reaction between H2O molecules (2) is
the pH value only changes slightly. The titra-
very small. At 25 °C,
tion curve (top) shows that buffer systems are
most effective at the pH values that corre-
K = [H+]
[OH-] / [H2O] = 2
10-16 mol L-1
spond to the pKa value of the acid. This is
where the curve is at its steepest, so that the
In pure water, the concentration
[H2O] is
pH change, ∆pH, is at its smallest with a given
practically constant at 55 mol L-1. Substitut-
increase ∆c in [H+] or [OH-]. In other words,
ing this value into the equation, it gives:
the buffer capacity ∆c/ ∆pH is highest at the
pKa value.
Kw = [H+]
[OH-] = 1
10-14 mol L-1
The product [H+]
[OH-]—the ion product of
water—is constant even when additional
acid-base pairs are dissolved in the water.
At 25 °C, pure water contains H+ and OH- at
concentrations of 1
10-7 mol L-1 each; it is
neutral and has a pH value of exactly 7.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
31
A. Acids and bases
Proton
Hydrogen
Chloride ion
exchange
chloride
Cl
Cl
Very weak base
Very strong acid
H
H
1
pKa = -7
O
O
Water
Hydronium ion
H
H
H
H
Very weak base
Very strong acid
Keq = 9 · 106 mol · l-1
Proton
Water
H
H
Hydroxyl ion
exchange
O
O
Very weak acid
Very strong base
H
2
H
pKa = 15.7
Water
O
O
Hydronium ion
H
H
H
H
Very weak base
Very strong acid
Keq = 2 · 10-16 mol · l-1
H
Proton
H
O
exchange
O
Water
Hydroxyl ion
Very weak acid
Very strong base
H
H
3
H
H
pKa = 9.2
N
H
N
H
Ammonia
Ammonium ion
H
H
Strong base
Weak acid
Keq = 6 · 10-10 mol · l-1
B. pH values in the body
C. Buffers
% B
∆ pH
pH 2
3
4
5
6
7
8
9
100
80
Base
60
pKa
Gastric juice
40
Acid
20
Lysosomes
0
∆ pH
pH
Sweat
Acid
Base
Urine
H
B
HB
OH
Cytoplasm
Blood plasma
HB
B
H2O
H2O
Buffer solution:
Small intestine
mixture of a weak acid
with the conjugate base
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
32
Basics
other (see below). Only the combination 2 e-/
Redox processes
1 H+, the hydride ion, is transferred as a unit.
A. Redox reactions
C. Biological redox systems
Redox reactions are chemical changes in
which electrons are transferred from one re-
In the cell, redox reactions are catalyzed by
action partner to another (1; see also p. 18).
enzymes, which work together with soluble
Like acid-base reactions (see p. 30), redox re-
or bound redox cofactors.
actions always involve pairs of compounds. A
Some of these factors contain metal ions as
pair of this type is referred to as a redox
redox-active components. In these cases, it is
system (2). The essential difference between
usually single electrons that are transferred,
the two components of a redox system is the
with the metal ion changing its valency. Un-
number of electrons they contain. The more
paired electrons often occur in this process,
electronrich component is called the reduced
but these are located in d orbitals (see p. 2)
form of the compound concerned, while the
and are therefore less dangerous than single
other one is referred to as the oxidized form.
electrons in non-metal atoms (“free radicals”;
The reduced form of one system (the reducing
see below).
agent) donates electrons to the oxidized form
We can only show here a few examples
of another one (the oxidizing agent). In the
from the many organic redox systems that
process, the reducing agent becomes oxidized
are found. In the complete reduction of the
and the oxidizing agent is reduced (3). Any
flavin coenzymes FMN and FAD (see p.104),
given reducing agent can reduce only certain
2 e- and 2 H+ are transferred. This occurs in
other redox systems. On the basis of this type
two separate steps, with a semiquinone radi-
of observation, redox systems can be ar-
cal appearing as an intermediate. Since or-
ranged to form what are known as redox
ganic radicals of this type can cause damage
series (4).
to biomolecules, flavin coenzymes never oc-
The position of a system within one of
cur freely in solution, but remain firmly
these series is established by its redox
bound in the interior of proteins.
potential E (see p. 18). The redox potential
In the reduction or oxidation of quinone/
has a sign; it can be more negative or more
quinol systems, free radicals also appear as
positive than a reference potential arbitrarily
intermediate steps, but these are less reactive
set at zero (the normal potential of the system
than flavin radicals. Vitamin E, another qui-
[2 H+/H2]). In addition, E depends on the con-
none-type redox system (see p. 104), even
centrations of the reactants and on the reac-
functions as a radical scavenger, by delocaliz-
tion conditions (see p. 18). In redox series (4),
ing unpaired electrons so effectively that they
the systems are arranged according to their
can no longer react with other molecules.
increasing redox potentials. Spontaneous
The pyridine nucleotides NAD+ and NADP+
electron transfers are only possible if the re-
always function in unbound form. The oxi-
dox potential of the donor is more negative
dized forms contain an aromatic nicotinamide
than that of the acceptor (see p. 18).
ring in which the positive charge is delocal-
ized. The right-hand example of the two res-
onance structures shown contains an elec-
B. Reduction equivalents
tron-poor, positively charged C atom at the
In redox reactions, protons (H+) are often
para position to nitrogen. If a hydride ion is
transferred along with electrons (e-), or pro-
added at this point (see above), the reduced
tons may be released. The combinations of
forms NADH or NADPH arise. No radical inter-
electrons and protons that occur in redox
mediate steps occur. Because a proton is re-
processes are summed up in the term reduc-
leased at the same time, the reduced pyridine
tion equivalents. For example, the combina-
nucleotide coenzymes are correctly expressed
tion 1 e-/1 H+ corresponds to a hydrogen
as NAD(P)H+H+.
atom, while 2 e- and 2 H+ together produce
a hydrogen molecule. However, this does not
mean that atomic or molecular hydrogen is
actually transferred from one molecule to the
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Physical Chemistry
33
A. Redox reactions
Possible
A red
B ox
Redox
Oxidizing
Not
system
agent
possible
A
2e
2e
Electron
Redox
exchange
system
2e
B
Reducing
becomes
agent
reduced
2e
Redox
A ox
B red
system
C
becomes
oxidized
1. Principle
2. Redox systems
3. Possible electron
4. Redox series
3. transfers
B. Reducing equivalents
Transferred
1e
1e
1H
2e
1H
2e
2H
components
e
[H]
H
[H2]
Equivalent
Electron
Hydrogen atom
Hydride ion
Hydrogen molecule
C. Biological redox systems
e
Metal
n+
1e
Me
Mem+
complexes
Oxidized
Reduced
O
H
O
H
O
1e
F
A
e
e
H3C
N
C
H3C
N
C
H3C
N
C
NH
NH
NH
1H
F
C
C
C
H
3C
N N
O
H3C
N N
O
H3C
N N
O
Flavin
H
H
R
R
R
H
Oxidized flavin
Semiquinone radical
Reduced flavin
O
OH
OH
e
e
H3CO C
H
H3CO
C H
H3CO
C H
H3CO
C
R
H3CO
C
R
H3CO
C
R
Quinone/
H
H
hydro-
O
O
OH
quinone
p-Benzoquinone
Semiquinone radical
Hydroquinone
Water H
e
H e
H
e
H O
O O
O O
O O
e
H
H
H
H
Reactive
O
O H
oxygen
Hydroperoxyl
Hydrogen
H
H
H
species
radical
peroxide
Oxygen
(ROS)
Hydroxyl radical
Water
H
Electron-poor
H
Hydride ion
H
H
1H
N
A
H
C
CONH
2
H
C CONH
2
H
C CONH
2
H
N
A
H
N H
H
N H
H
N H
P
R
R
R
NAD (P)
NAD(P)
(Resonance structures)
NAD(P)H + H
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
34
Biomolecules
The open-chained form of glucose shown
Overview
in (1) is found in neutral solution in less than
The carbohydrates are a group of naturally
0.1% of the molecules. The reason for this is an
occurring carbonyl compounds
(aldehydes
intramolecular reaction in which one of the
or ketones) that also contain several hydroxyl
OH groups of the sugar is added to the alde-
groups. The carbohydrates include single sug-
hyde group of the same molecule (2). This
ars
(monosaccharides) and their polymers,
gives rise to a cyclic hemiacetal (see p. 10). In
the oligosaccharides and polysaccharides.
aldohexoses, the hydroxy group at C-5 reacts
preferentially, and a six-membered pyran
ring is formed. Sugars that contain this ring
A. Carbohydrates: overview
are called pyranoses. By contrast, if the OH
Polymeric carbohydrates-above all starch, as
group at C-4 reacts, a five-part furan ring is
well as some disaccharides-are important
formed. In solution, pyranose forms and
(but not essential) components of food (see
furanose forms are present in equilibrium
p. 360). In the gut, they are broken down into
with each other and with the open-chained
monosaccharides and resorbed in this form
form, while in glucose polymers only the
(see p. 272). The form in which carbohydrates
pyranose form occurs.
are distributed by the blood of vertebrates is
The Haworth projection (2) is usually used
glucose (“blood sugar”). This is taken up by the
to depict sugars in the cyclic form, with the
cells and either broken down to obtain energy
ring being shown in perspective as viewed
(glycolysis) or converted into other metabo-
from above. Depending on the configuration,
lites (see pp. 150-159). Several organs (partic-
the substituents of the chiral C atoms are then
ularly the liver and muscles) store glycogen as
found above or below the ring. OH groups
a polymeric reserve carbohydrate (right; see
that lie on the right in the Fischer projection
p. 156). The glycogen molecules are covalently
(1) appear under the ring level in the Haworth
bound to a protein, glycogenin. Polysaccha-
projection, while those on the left appear
rides are used by many organisms as building
above it.
materials. For example, the cell walls of bac-
As a result of hemiacetal formation, an ad-
teria contain murein as a stabilizing compo-
ditional chiral center arises at C-1, which can
nent (see p. 40), while in plants cellulose and
be present in both possible configurations
other polysaccharides fulfill this role
(see
(anomers) (see p. 8). To emphasize this, the
p. 42). Oligomeric or polymeric carbohydrates
corresponding bonds are shown here using
are often covalently bound to lipids or pro-
wavy lines.
teins. The glycolipids and glycoproteins
The Haworth formula does not take ac-
formed in this way are found, for example,
count of the fact that the pyran ring is not
in cell membranes
(center). Glycoproteins
plain, but usually has a chair conformation. In
also occur in the blood in solute form (plasma
B3, two frequent conformations of D-glucopy-
proteins; see p. 276) and, as components of
ranose are shown as ball-and-stick models. In
proteoglycans, form important constituents of
the1C4 conformation (bottom), most of the
the intercellular substance (see p. 346).
OH groups appear vertical to the ring level, as
in the Haworth projection (axial or a posi-
tion). In the slightly more stable4C1 confor-
B. Monosaccharides: structure
mation (top), the OH groups take the equato-
The most important natural monosaccharide,
rial or e position. At room temperature, each
D-glucose, is an aliphatic aldehyde with six C
form can change into the other, as well as into
atoms, five of which carry a hydroxyl group
other conformations.
(1). Since C atoms 2 to 5 represent chiral
centers
(see p. 8), there are
15
further
isomeric aldohexoses in addition to D-glucose,
although only a few of these are important in
nature (see p. 38). Most natural monosaccha-
rides have the same configuration at C-5 as
D-glyceraldehyde-they belong to the D series.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrates
35
A. Carbohydrates: overview
Other monosaccharides
Transporter
Glycogenin
CH
2OH
H
O
OH
H
Glucose
OH H
HO
H
Mono-
H OH
Gluconeo-
saccharide
Glycolysis
genesis
Pyruvate
Amino
acids
Glycoproteins
ATP
CO2+H2O
Glycogen
Glycolipids
Peptidoglycan
(Murein)
Proteoglycans
Periplasm
Bacterium
B. Monosaccharides: structure
HOCH2
6
Open-chained
4
H
OH
O
5
form (< 0.1%)
H
O
H
4
OH
H
C
H
C
1
1
1
HO
3
2
H
C
OH
2
H
OH
HO3
C
H
4C1-conformation
H
C
OH
4
Hemiacetal formation
CH
H
C
OH
2OH
5
HOCH2
CH2OH
HO
C
H
1
6
OH
O
H
O
OH
H
4
Open-chained
OH H
OH
H
H
form of glucose
H
H
HO
H OH
H
OH
D-Gluco-
D-Gluco-
Chiral
1C4-conformation
furanose (<1%)
pyranose (99%)
center
1. Fischer projection
2. Ring forms (Haworth projection)
3. Conformations
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
36
Biomolecules
5. Esterification. The hydroxyl groups of
Chemistry of sugars
monosaccharides can form esters with acids.
In metabolism, phosphoric acid esters such as
A. Reactions of the monosaccharides
glucose 6-phosphate and glucose 1-phosphate
The sugars (monosaccharides) occur in the
(6) are particularly important.
metabolism in many forms
(derivatives).
Only a few important conversion reactions
B. Polarimetry, mutarotation
are discussed here, using D-glucose as an ex-
ample.
Sugar solutions can be analyzed by polarim-
1. Mutarotation. In the cyclic form, as op-
etry, a method based on the interaction be-
posed to the open-chain form, aldoses have a
tween chiral centers and linearly polarized
chiral center at C-1 (see p. 34). The corre-
light—i. e., light that oscillates in only one
sponding isomeric forms are called anomers.
plane. It can be produced by passing normal
In the β-anomer (center left), the OH group at
light through a special filter (a polarizer). A
C-1 (the anomeric OH group) and the CH2OH
second polarizing filter of the same type (the
group lie on the same side of the ring. In the α-
analyzer), placed behind the first, only lets the
anomer (right), they are on different sides.
polarized light pass through when the polar-
The reaction that interconverts anomers into
izer and the analyzer are in alignment. In this
each other is known as mutarotation (B).
case, the field of view appears bright when
2. Glycoside formation. When the anome-
one looks through the analyzer (1). Solutions
ric OH group of a sugar reacts with an alcohol,
of chiral substances rotate the plane of polar-
with elimination of water, it yields an
ized light by an angle α either to the left or to
O-glycoside (in the case shown, α -methylglu-
the right. When a solution of this type is
coside). The glycosidic bond is not a normal
placed between the polarizer and the ana-
ether bond, because the OH group at C-1 has a
lyzer, the field of view appears darker (2).
hemiacetal quality. Oligosaccharides and pol-
The angle of rotation, α, is determined by
ysaccharides also contain O-glycosidic bonds.
turning the analyzer until the field of view
Reaction of the anomeric OH group with an
becomes bright again (3). A solution’s optical
NH2 or NH group yields an N-glycoside (not
rotation depends on the type of chiral com-
shown). N-glycosidic bonds occur in nucleo-
pound, its concentration, and the thickness of
tides
(see p. 80) and in glycoproteins (see
the layer of the solution. This method makes it
p. 44), for example.
possible to determine the sugar content of
3. Reduction and oxidation. Reduction of
wines, for example.
the anomeric center at C-1 of glucose (2) pro-
Certain procedures make it possible to ob-
duces the sugar alcohol sorbitol. Oxidation of
tain the α and β anomers of glucose in pure
the aldehyde group at C-1 gives the intramo-
form. A 1-molar solution of α-D-glucose has a
lecular ester (lactone) of gluconic acid (a gly-
rotation value [α]D of +112°, while a corre-
conic acid). Phosphorylated gluconolactone is
sponding solution of β-D-glucose has a value
an intermediate of the pentose phosphate
of +19°. These values change spontaneously,
pathway (see p. 152). When glucose is oxi-
however, and after a certain time reach the
dized at C-6, glucuronic acid (a glycuronic
same end point of +52°. The reason for this is
acid) is formed. The strongly polar glucuronic
that, in solution, mutarotation leads to an
acid plays an important role in biotransforma-
equilibrium between the α and β forms in
tions in the liver (see pp. 194, 316).
which, independently of the starting condi-
4. Epimerization. In weakly alkaline solu-
tions, 62% of the molecules are present in the
tions, glucose is in equilibrium with the
β form and 38% in the α form.
ketohexose D-fructose and the aldohexose D-
mannose, via an enediol intermediate (not
shown). The only difference between glucose
and mannose is the configuration at C-2. Pairs
of sugars of this type are referred to as epi-
mers, and their interconversion is called epi-
merization.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrates
37
A. Reactions of the monosaccharides
Glucuronate
Gluconolactone
Sorbitol
6
COO
HOCH2
HOCH2
H
OOH
H
O
H
OH
H
H
H
1
OH
H
OH
H
O
OH
H
CH2OH
1
HO
H
HO
HO
H
OH
H
OH
H
OH
Oxidation
Oxidation
Reduction
HOCH2
HOCH2
H
OOHβ
H
O
H
H
H
1
1
OH H
Mutarotation
OH
H
HO
H
HO
OH
α
H
OH
H
OH
β-D-Glucose
α-D-Glucose
Esterification
Epimerization
Glycoside formation
O
O
P
O
6CH2
H
HOCH2
HOCH2
1
O
H
OOH
H
O
CH2OH
H
O
H
H
O
H
H
H
H
H
2
1
OH H
H
HO
OHHO
OH
H
HO
H
HO
OH
HO
OH
HO
O CH3
3
2
α
H
OH
OH
H
H
H
H
OH
Glucose 6-
D
-Fructose
α-D-Mannose
α-Methyl-
phosphate
glucoside
B. Polarimetry, mutarotation
Polarizer
Analyzer
α-D-Glucose: [ α ]
= +112°
D
)
α (˚
1.
100
a
80
+ 52°
62% β
2.
60
40
38% α
a
20
3.
0
10
20
30
40
50
Time (min)
β-D-Glucose: [α]
= +19°
D
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
38
Biomolecules
Sugar alcohols
(6) such as sorbitol and
Monosaccharides and disaccharides
mannitol do not play an important role in
animal metabolism.
A. Important monosaccharides
Only the most important of the large number
B. Disaccharides
of naturally occurring monosaccharides are
mentioned here. They are classified according
When the anomeric hydroxyl group of one
to the number of C atoms (into pentoses,
monosaccharide is bound glycosidically with
hexoses, etc.) and according to the chemical
one of the OH groups of another, a disaccha-
nature of the carbonyl function into aldoses
ride is formed. As in all glycosides, the glyco-
and ketoses.
sidic bond does not allow mutarotation. Since
The best-known aldopentose (1), D-ribose,
this type of bond is formed stereospecifically
is a component of RNA and of nucleotide
by enzymes in natural disaccharides, they are
coenzymes and is widely distributed. In these
only found in one of the possible configura-
compounds, ribose always exists in the fura-
tions (α or β).
nose form (see p. 34). Like ribose, D-xylose and
Maltose (1) occurs as a breakdown product
L-arabinose are rarely found in free form.
of the starches contained in malt
(“malt
However, large amounts of both sugars are
sugar”; see p. 148) and as an intermediate in
found as constituents of polysaccharides in
intestinal digestion. In maltose, the anomeric
the walls of plant cells (see p. 42).
OH group of one glucose molecule has an α-
The most important of the aldohexoses (1)
glycosidic bond with C-4 in a second glucose
is D-glucose. A substantial proportion of the
residue.
biomass is accounted for by glucose polymers,
Lactose (“milk sugar,” 2) is the most impor-
above all cellulose and starch. Free D-glucose
tant carbohydrate in the milk of mammals.
is found in plant juices (“grape sugar”) and as
Cow’s milk contains 4.5% lactose, while hu-
“blood sugar” in the blood of higher animals.
man milk contains up to 7.5%. In lactose, the
As a constituent of lactose (milk sugar), D-
anomeric OH group of galactose forms a β-
galactose is part of the human diet. Together
glycosidic bond with C-4 of a glucose. The
with D-mannose, galactose is also found in
lactose molecule is consequently elongated,
glycolipids and glycoproteins (see p. 44).
and both of its pyran rings lie in the same
Phosphoric acid esters of the ketopentose
plane.
D-ribulose (2) are intermediates in the pen-
Sucrose (3) serves in plants as the form in
tose phosphate pathway (see p. 152) and in
which carbohydrates are transported, and as a
photosynthesis (see p. 128). The most widely
soluble carbohydrate reserve. Humans value
distributed of the ketohexoses is D-fructose. In
it because of its intensely sweet taste. Sources
free form, it is present in fruit juices and in
used for sucrose are plants that contain par-
honey. Bound fructose is found in sucrose (B)
ticularly high amounts of it, such as sugar
and plant polysaccharides (e. g., inulin).
cane and sugar beet (cane sugar, beet sugar).
In the deoxyaldoses (3), an OH group is
Enzymatic hydrolysis of sucrose-containing
replaced by a hydrogen atom. In addition to
flower nectar in the digestive tract of bees—
2-deoxy-D-ribose, a component of DNA (see
catalyzed by the enzyme invertase—produces
p. 84) that is reduced at C-2, L-fucose is shown
honey, a mixture of glucose and fructose. In
as another example of these. Fucose, a sugar
sucrose, the two anomeric OH groups of glu-
in the λ series (see p. 34) is reduced at C-6.
cose and fructose have a glycosidic bond; su-
The acetylated amino sugars N-acetyl-D-
crose is therefore one of the non-reducing
glucosamine and N-acetyl-D-Galactosamine
sugars.
(4) are often encountered as components of
glycoproteins.
N-acetylneuraminic acid (sialic acid, 5), is a
characteristic component of glycoproteins.
Other acidic monosaccharides such as D-glu-
curonic acid, D-galacturonic acid, and liduronic
acid, are typical constituents of the glycosa-
minoglycans found in connective tissue.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrates
39
A. Important monosaccharides
1
Aldoses
2
Ketoses
D-Ribose (Rib)
D-Xylose (Xyl)
L-Arabinose (Ara)
D-Ribulose (Rub)
CH2OH
HOCH2
HOCH2
H
H
O
O
O
OH
C O
O
H H
OH
OH H OH
OH H OH
H H
H
C
OH
CH2OH
H
H
HOH2C
H
H
C
OH
OH OH
H OH
H OH
OH OH
CH2OH
Pentoses
D
-Glucose (Glc)
D-Mannose (Man)
D-Galactose (Gal)
D-Fructose (Fru)
CH2OH
HOCH2
HOCH2
HOCH2
C O
H
O
H
O
HO
O
H
H
H
HO
C
H
OH
H OH
OHHO
OH
OH H
OH
HOCH2
OH
O
H
C
OH
HO
HO
H
H HO
CH2OH
H
OH
H
H
H
OH
H
C
OH
H
Hexoses
CH2OH
OH H
3
Deoxyaldoses
4
Acetylated amino sugars
2-Deoxy-
L-Fucose (Fuc)
N-Acetyl-D-glucos-
N -Acetyl-D-galac-
D-ribose (dRib)
amine (GlcNAc)
tosamine (GalNAc)
HOCH2
H
HOCH2
HOCH2
O
H
O
H
O
HO
O
H H OH
CH
6
H
H
3
H
HO OH
OH
H
OH
OH
H
OH
H
2
HO
HO
2
H
2
OH H
OH
H
H
HN
C CH3
H
HN
C CH3
O
O
5
Acidic monosaccharides
N-Acetylneuraminic acid
6
Sugar alcohols (alditoles)
(NeuAc)
D-Sorbitol
D-Mannitol
D-Glucuronic acid
L-Iduronic acid
9
CH2OH
(GlcUA)
(IduUA)
CH2OH
CH2OH
HO
C
H
6
COO
H
H
H
C OH
HO
C H
HO
C
H
H
O
H
O
H
2
O
7
H
COO
COO
HO
C H
HO
C H
OH
H
OH
OH
H
OH
3
OH
H
6
5
H
C
OH
H
C
OH
HO
HO
H
4
H
H
C
OH
H
C
OH
H
OH
H
OH
H
HN
C CH3
CH2OH
CH2OH
O
B. Disaccharides
CH2OH
CH2OH
CH2OH
CH2OH
CH2OH
β
CH2OH
H
O
H
H
O
HO
O
H
O
H
O
H
H
H
H
H
H
H
O
OH
H
1
4
OH
H
OH
OH
H
1
O
4
OH
H
OH
OH
H
1
2
H HO
HO
α
O
H
H
HO
O
CH2OH
α
β
H
OH
H
OH
H
OH
H
OH
H
OH
OH H
1. Maltose
2. Lactose
3. Sucrose
α-D-Glucopyranosyl-
β-D-Galactopyranosyl-
α-D-Glucopyranosyl-
(1
4)-D-glucopyranose
(1
4)-D-glucopyranose
(1
2)-β-D-fructofuranoside
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
40
Biomolecules
Polysaccharides: overview
B. Important polysaccharides
Polysaccharides are ubiquitous in nature.
The table gives an overview of the composi-
They can be classified into three separate
tion and make-up both of the glycans men-
groups, based on their different functions.
tioned above and of several more.
Structural polysaccharides provide mechani-
In addition to murein, bacterial polysac-
cal stability to cells, organs, and organisms.
charides include dextrans—glucose polymers
Waterbinding polysaccharides are strongly
that are mostly α1
6-linked and α1
3-
hydrated and prevent cells and tissues from
branched. In water, dextrans form viscous
drying out. Finally, reserve polysaccharides
slimes or gels that are used for chromato-
serve as carbohydrate stores that release
graphic separation of macromolecules after
monosaccharides as required. Due to their
chemical treatment (see p. 78). Dextrans are
polymeric nature, reserve carbohydrates are
also used as components of blood plasma
osmotically less active, and they can therefore
substitutes
(plasma expanders) and food-
be stored in large quantities within the cell.
stuffs.
Carbohydrates from algae (e. g., agarose
and carrageenan) can also be used to produce
A. Polysaccharides: structure
gels. Agarose has been used in microbiology
Polysaccharides that are formed from only
for more than 100 years to reinforce culture
one type of monosaccharide are called homo-
media (“agar-agar”). Algal polysaccharides are
glycans, while those formed from different
also added to cosmetics and ready-made
sugar constituents are called heteroglycans.
foods to modify the consistency of these prod-
Both forms can exist as either linear or
ucts.
branched chains.
The starches, the most important vegetable
A section of a glycogen molecule is shown
reserve carbohydrate and polysaccharides
here as an example of a branched homogly-
from plant cell walls, are discussed in greater
can. Amylopectin, the branched component of
detail on the following page. Inulin, a fructose
vegetable starch (see p. 42), has a very similar
polymer, is used as a starch substitute in dia-
structure. Both molecules mainly consist of
betics’ dietary products (see p. 160). In addi-
α1
4-linked glucose residues. In glycogen,
tion, it serves as a test substance for measur-
on average every 8th to 10th residue car-
ing renal clearance (see p. 322).
ries —via an α1
6 bond—another 1,4-linked
Chitin, a homopolymer from β1
4-linked
chain of glucose residues. This gives rise to
N-acetylglucosamine, is the most important
branched, tree-like structures, which in ani-
structural substance in insect and crustacean
mal glycogen are covalently bound to a
shells, and is thus the most common animal
protein, glycogenin (see p. 156).
polysaccharide. It also occurs in the cell wall
The linear heteroglycan murein, a struc-
of fungi.
tural polysaccharide that stabilizes the cell
Glycogen, the reserve carbohydrate of
walls of bacteria, has a more complex struc-
higher animals, is stored in the liver and mus-
ture. Only a short segment of this thread-like
culature in particular (A, see pp. 156, 336). The
molecule is shown here. In murein, two differ-
formation and breakdown of glycogen are
ent components, both β1
4-linked, alter-
subject to complex regulation by hormones
nate: N-acetylglucosamine
(GlcNAc) and
and other factors (see p. 120).
N-acetylmuraminic acid
(MurNAc), a lactic
acid ether of N-acetylglucosamine. Peptides
are bound to the carboxyl group of the lactyl
groups, and attach the individual strands of
murein to each other to form a three-dimen-
sional network (not shown). Synthesis of the
network-forming peptides in murein is inhib-
ited by penicillin (see p. 254).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrates
41
A. Polysaccharides: structure
HOCH2
HOCH2
Glycogen -
H
O
H
H
O
H
branched
H
H
1
4
1
homopolymer
OH
H
OH H
α
O
O
α
O
H
OH
H
HO
6CH2
HOCH2
HOCH2
HOCH2
Reducing end
H
O
H
H
O
H
H
O
H
H
O
H
H
H
H
OH
H
1
4
OH H
1
4
OH H
1
4
OH
H
OH
O
O
O
O
α
α
α
H
OH
H
OH
H
OH
H
OH
HOCH2
HOCH2
HOCH2
HOCH2
β
β
β
H
O
H
O
H
O
H
O
H
H
H
H
O
OH
H
1
O
4
H
1
O
4
OH
H
1
O
4
H
O
3
2
H
H
H
H
H
NHCOCH3
H
NHCOCH3
H
NHCOCH3
H
NHCOCH3
O
O
Murein - linear
H3C C
C O
H3C
C
C O
heteropolymer
H
NH
H
NH
Peptide
B. Important polysaccharides
Poly-
Mono-
Mono-
Linkage
Branch-
Occurrence
Function
saccharide
saccharide 1
saccharide 2
ing
Bacteria
Murein
D-GlcNAc
D-MurNAc1)
β
1
4
Cell wall
SC
Dextran
D-Glc
α
1
6
α
1
3
Slime
WB
Plants
Agarose
D-Gal
L-aGal2)
β
1
4
β
1
3
Red algae (agar)
WB
Carrageenan
D-Gal
β
1
3
α
1
4
Red algae
WB
Cellulose
D-Glc
β
1
4
Cell wall
SC
Xyloglucan
D-Glc
D-Xyl (D-Gal,
β
1
4
β
1
6
Cell wall
SC
L-Fuc)
(
β
1
2)
(Hemicellulose)
SC
Arabinan
L-Ara
α
1
5
α
1
3
Cell wall (pectin)
Amylose
D-Glc
α
1
4
Amyloplasts
RC
Amylopectin
D-Glc
α
1
4
α
1
6
Amyloplasts
RC
Inulin
D-Fru
β
2
1
Storage cells
RC
Animals
Chitin
D-GlcNAc
β
1
4
Insects, crabs
SK
Glycogen
D-Glc
α
1
4
α
1
6
Liver, muscle
RK
Hyaluronic
D-GlcUA
D-GlcNAc
β
1
4
Connective tissue
SK,WB
acid
β
1
3
SC= structural carbohydrate, RC= reserve carbohydrate,
WB = water-binding carbohydrate; 1) N-acetylmuramic acid, 2) 3,6-anhydrogalactose
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
42
Biomolecules
B. Starch
Plant polysaccharides
Starch, a reserve polysaccharide widely dis-
Two glucose polymers of plant origin are of
tributed in plants, is the most important car-
special importance among the polysac-
bohydrate in the human diet. In plants, starch
charides: β1
4-linked polymer cellulose
is present in the chloroplasts in leaves, as well
and starch, which is mostly α1
4-linked.
as in fruits, seeds, and tubers. The starch con-
tent is especially high in cereal grains (up to
A. Cellulose
75% of the dry weight), potato tubers (ap-
proximately 65%), and in other plant storage
Cellulose, a linear homoglycan of β1
4-
organs.
linked glucose residues, is the most abundant
In these plant organs, starch is present in
organic substance in nature. Almost half of the
the form of microscopically small granules in
total biomass consists of cellulose. Some
special organelles known as amyloplasts.
40-50% of plant cell walls are formed by cel-
Starch granules are virtually insoluble in cold
lulose. The proportion of cellulose in cotton
water, but swell dramatically when the water
fibers, an important raw material, is 98%. Cel-
is heated. Some 15-25% of the starch goes
lulose molecules can contain more than 104
into solution in colloidal form when the mix-
glucose residues (mass 1-2
106 Da) and can
ture is subjected to prolonged boiling. This
reach lengths of 6-8 µm.
proportion is called amylose
(“soluble
Naturally occurring cellulose is extremely
starch”).
mechanically stable and is highly resistant to
Amylose consists of unbranched α1
4-
chemical and enzymatic hydrolysis. These
linked chains of 200-300 glucose residues.
properties are due to the conformation of
Due the α configuration at C-1, these chains
the molecules and their supramolecular or-
form a helix with 6-8 residues per turn (1).
ganization. The unbranched β1
4 linkage re-
The blue coloring that soluble starch takes on
sults in linear chains that are stabilized by
when iodine is added (the “iodine-starch re-
hydrogen bonds within the chain and be-
action”) is caused by the presence of these
tween neighboring chains (1). Already during
helices—the iodine atoms form chains inside
biosynthesis, 50-100 cellulose molecules as-
the amylose helix, and in this largely non-
sociate to form an elementary fibril with a
aqueous environment take on a deep blue
diameter of 4 nm. About 20 such elementary
color. Highly branched polysaccharides turn
fibrils then form a microfibril (2), which is
brown or reddishbrown in the presence of
readily visible with the electron microscope.
iodine.
Cellulose microfibrils make up the basic
Unlike amylose, amylopectin, which is
framework of the primary wall of young plant
practically insoluble, is branched. On average,
cells (3), where they form a complex network
one in 20-25 glucose residues is linked to
with other polysaccharides. The linking poly-
another chain via an α1
6 bond. This leads
saccharides include hemicellulose, which is a
to an extended tree-like structure, which—
mixture of predominantly neutral heterogly-
like amylose—contains only one anomeric
cans (xylans, xyloglucans, arabinogalactans,
OH group (a “reducing end”). Amylopectin
etc.). Hemicellulose associates with the cellu-
molecules can contain hundreds of thousands
lose fibrils via noncovalent interactions. These
of glucose residues; their mass can be more
complexes are connected by neutral and
than 108 Da.
acidic pectins, which typically contain galac-
turonic acid. Finally, a collagen-related
protein, extensin, is also involved in the for-
mation of primary walls.
In the higher animals, including humans,
cellulose is indigestible, but important as
roughage (see p. 273). Many herbivores (e. g.,
the ruminants) have symbiotic unicellular or-
ganisms in their digestive tracts that break
down cellulose and make it digestible by the
host.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrates
43
A. Cellulose
Microfibril
Pectin
Elementary
Ca2
Ca2
fibril
2.
Hemi-
cellulose
1.
6
Cellulose
β
4
Extensin
microfibril
3.
3
2
1
5
4
5
2
1
3
6
Mitochondria
Golgi apparatus
Nucleus
Endoplasmic
reticulum
Plasma membrane
B. Starch
Vacuole
Chloroplasts
Plant cell
Starch
Starch
1. Amylose 20%
2. Amylopectin 80%
OH
OH
O
O
O
O
OH
O
1
OH
O
O
HO
OH
α
6
α
O
1
O
O O
4
O
O
O
HO
1α
O
4
O O
O
O
O
HO Reducing end
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
44
Biomolecules
As an example of the carbohydrate compo-
Glycosaminoglycans and
nent of a glycoprotein, the structure of one of
glycoproteins
the oligosaccharide chains of immunoglobu-
lin G (IgG; see p. 300) is shown here. The
A. Hyaluronic acid
oligosaccharide has an N-glycosidic link to
the amide group of an asparagine residue in
As constituents of proteoglycans (see p. 346),
the Fc part of the protein. Its function is not
the glycosaminoglycans—a group of acidic
known.
heteropolysaccharides—are important struc-
Like all N-linked carbohydrates, the oligo-
tural elements of the extracellular matrix.
saccharide in IgG contains a T-shaped core
Glycosaminoglycans contain amino sugars
structure consisting of two N-acetylglucos-
as well as glucuronic acid and iduronic acid as
amines and three mannose residues (shown
characteristic components (see p. 38). In ad-
in violet). In addition, in this case the struc-
dition, most polysaccharides in this group are
ture contains two further N-acetylglucos-
esterified to varying extents by sulfuric acid,
amine residues, as well as a fucose residue
increasing their acidic quality. Glycosamino-
and a galactose residue. Glycoproteins show
glycans can be found in free form, or as com-
many different types of branching. In this
ponents of proteoglycans throughout the or-
case, we not only have β1
4 linkage, but
ganism.
also β1
2, α1
3, and α1
6 bonds.
Hyaluronic acid, an unesterified glycosami-
noglycan with a relatively simple structure,
consists of disaccharide units in which N-
C. Glycoproteins: forms
acetylglucosamine and glucuronic acid are
On the cell surface of certain glycoproteins,
alternately β1
4-linked and β1
3-linked.
O-glycosidic links are found between the car-
Due to the unusual β1
3 linkage, hyaluronic
bohydrate part and a serine or threonine res-
acid molecules-which may contain several
idue, instead of N-glycosidic links to aspara-
thousand monosaccharide residues—are
gine residues. This type of link is less common
coiled like a helix. Three disaccharide units
than the N-glycosidic one.
form each turn of the helix. The outwardfac-
There are two types of oligosaccharide
ing hydrophilic carboxylate groups of the glu-
structure with N-glycosidic links, which arise
curonic acid residues are able to bind Ca2+
through two different biosynthetic pathways.
ions. The strong hydration of these groups
During glycosylation in the ER, the protein is
enables hyaluronic acid and other glycosami-
initially linked to an oligosaccharide, which in
noglycans to bind water up to 10 000 times
addition to the core structure contains six
their own volume in gel form. This is the
further mannose residues and three terminal
function which hyaluronic acid has in the vit-
glucose residues
(see p. 230). The simpler
reous body of the eye, which contains approx-
from of oligosaccharide
(the mannose-rich
imately 1% hyaluronic acid and 98% water.
type) is produced when only the glucose res-
idues are cleaved from the primary product,
and no additional residues are added. In other
B. Oligosaccharide in immunoglobulin G
cases, the mannose residues that are located
(IgG)
outside the core structure are also removed
Many proteins on the surface of the plasma
and replaced by other sugars. This produces
membrane, and the majority of secreted pro-
oligosaccharides such as those shown on the
teins, contain oligosaccharide residues that
right (the complex type). At the external end
are post-translationally added to the endo-
of the structure, glycoproteins of the complex
plasmic reticulum and in the Golgi apparatus
type often contain N-acetylneuraminic acid
(see p. 230). By contrast, cytoplasmic proteins
residues, which give the oligosaccharide com-
are rarely glycosylated. Glycoproteins can
ponents negative charges.
contain more than 50 % carbohydrate; how-
ever, the proportion of protein is generally
much greater.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrates
45
A. Hyaluronic acid
COO
HOCH
2
COO
HOCH2
COO
HOCH2
β
β
H
O
H
O
H
O
H
O
H
O
H
O
O
H
H
O
H
H
O
H
H
O
H
O
OH H
1
3
H
1
O
OH
H
H
O
OH
H
HO
H
H HO
H
H
HO
H
H
4
H
OH
H
NHCOCH
3
H
OH
H
NHCOCH3
H
OH
H
NHCOCH3
Disaccharide unit
II
V
VI
IV
I
III
[
3)-β-D-GlcNAc-(1
4)-β-D-GlcUA-(1
4]n
B. Oligosaccharide in immunoglobulin G (IgG)
D-Gal
D-GlcNAc
HOCH
HOCH2
2
Core structure
HO
O
H
O
H
H
OH
H
O
OH H
H
H
H
H
OH
H
NHCOCH3
HOH2C
H
H
O
H
H
O
H
D-Man
H
CH3
O
OH
L-Fuc
H
HO
N-glycosidic
HOCH2
HO
O
HO
O
bond
H
O
H
H
OH
H
H
CH
HOCH2
CH2
D-GlcNAc
2
OH
H
H
O
H
O
H
O
HO
H
H
H
H
HO
O
OH H
O
OH H
HN
O
H
NHCOCH3
C
HOH2C
HO
H
H
H
H
O
H
H
H
H
NHCOCH3
H
NHCOCH3
CH2
D-Man
D-Man
D-GlcNAc
D-GlcNAc
H O
OH
C
HO
O
Asn-297
H
H
C. Glycoproteins: forms
NeuAc
NeuAc
O-linked
N-linked
NeuAc
Man
Man
Man
Gal
Gal
Gal
Man
Man
Man
GlcNAc GlcNAc
GlcNAc
Man
Man
Man
Man
NeuAc
Man
Man
Gal
GlcNAc
GlcNAc
GalNAc
NeuAc
GlcNAc
GlcNAc
Fuc
O
NH
NH
Ser
Protein
Asn
Asn
Mannose-rich type
Complex type
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
46
Biomolecules
are derivatives of the polyunsaturated fatty
Overview
acid arachidonic acid (see p. 390).
A. Classification
B. Biological roles
The lipids are a large and heterogeneous
group of substances of biological origin that
1. Fuel. Lipids are an important source of en-
are easily dissolved in organic solvents such
ergy in the diet. In quantitative terms, they
as methanol, acetone, chloroform, and ben-
represent the principal energy reserve in ani-
zene. By contrast, they are either insoluble or
mals. Neutral fats in particular are stored in
only poorly soluble in water. Their low water
specialized cells, known as adipocytes. Fatty
solubility is due to a lack of polarizing atoms
acids are released from these again as needed,
such as O, N, S, and P (see p. 6).
and these are then oxidized in the mitochon-
Lipids can be classified into substances that
dria to form water and carbon dioxide, with
are either hydrolyzable— i. e., able to undergo
oxygen being consumed. This process also
hydrolytic cleavage—or nonhydrolyzable. Only
gives rise to reduced coenzymes, which are
a few examples of the many lipids known can
used for ATP production in the respiratory
be mentioned here. The individual classes of
chain (see p. 140).
lipids are discussed in more detail in the fol-
2. Nutrients. Amphipathic lipids are used
lowing pages.
by cells to build membranes (see p. 214). Typ-
Hydrolyzable lipids (components shown in
ical membrane lipids include phospholipids,
brackets). The simple esters include the fats
glycolipids, and cholesterol. Fats are only
(triacylglycerol; one glycerol + three acyl res-
weakly amphiphilic and are therefore not
idues); the waxes (one fatty alcohol + one acyl
suitable as membrane components.
residue); and the sterol esters (one sterol + one
3. Insulation. Lipids are excellent insula-
acyl residue). The phospholipids are esters
tors. In the higher animals, neutral fats are
with more complex structures. Their charac-
found in the subcutaneous tissue and around
teristic component is a phosphate residue.
various organs, where they serve as mechan-
The phospholipids include the phosphatidic
ical and thermal insulators. As the principal
acids (one glycerol + two acyl residues + one
constituent of cell membranes, lipids also in-
phosphate) and the phosphatides (one glyc-
sulate cells from their environment mechan-
erol + two acyl residues + one phosphate +
ically and electrically. The impermeability of
one amino alcohol). In the sphingolipids, glyc-
lipid membranes to ions allows the formation
erol and one acyl residue are replaced by
of the membrane potential (see p. 126).
sphingosine. Particularly important in this
4. Special tasks. Some lipids have adopted
group are the sugar-containing glycolipids
special roles in the body. Steroids, eicosa-
(one sphingosine + one fatty acid + sugar).
noids, and some metabolites of phospholipids
The cerebrosides (one sphingosine + one fatty
have signaling functions. They serve as hor-
acid + one sugar) and gangliosides (one sphin-
mones, mediators, and second messengers
gosine + one fatty acid + several different
(see p. 370). Other lipids form anchors to at-
sugars, including neuraminic acid) are repre-
tach proteins to membranes (see p. 214). The
sentatives of this group.
lipids also produce cofactors for enzymatic re-
The components of the hydrolyzable lipids
actions—e. g., vitamin K (see p. 52) and ubiq-
are linked to one another by ester bonds. They
uinone (see p. 104). The carotenoid retinal, a
are easily broken down either enzymatically
light-sensitive lipid, is of central importance
or chemically.
in the process of vision (see p. 358).
Non-hydrolyzable lipids. The hydrocarbons
Several lipids are not formed indepen-
include the alkanes and carotenoids. The lipid
dently in the human body. These substances,
alcohols are also not hydrolyzable. They in-
as essential fatty acids and fat-soluble vita-
clude long-chained alkanols and cyclic sterols
mins, are indispensable components of nutri-
such as cholesterol, and steroids such as es-
tion (see pp. 364ff.)
tradiol and testosterone. The most important
acids among the lipids are fatty acids. The
eicosanoids also belong to this group; these
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipids
47
A. Classification
Hydrolyzable lipids
O
C O CH2
O
O
C O
CH2
C O CH
O
C O CH2
Esters
O
Fats
Non-hydrolyzable lipids
C O
Waxes
Sterol esters
O
Hydrocarbons
C O CH2
O
Alkanes
C O CH
O
Carotenoids
H2C O P
O
O
OH
Phospholipids
Phosphatidates
Alcohols
Phosphatids
Sphingolipids
Long-chain alkanols
Sterols
HO
O
Steroids
C O CH2
O
COOH
C O CH
O
CH2
H2C O P
O
N CH3
Acids
O
CH3
Fatty acids
O
Eicosanoids
C
NH
OH
Glycolipids
OH
HO
CH2
Cerebrosides
O
HO O
Gangliosides
OH
OH
B. Biological roles
Fat
Membrane
Cytoplasm
Glycerol
Fatty acid
ADP+P
ATP
i
Phospho-
lipid
O
2
Lipid
Mitochondrion
bilayer
CO2
H2O
1. Fuel
2. Building block
37 °C
Signaling
Anchor
0 °C
CoQ
Cell
Cofactor
Visual pigment
3. Thermal insulator
4. Special tasks
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
48
Biomolecules
(see p. 390). As the organism is capable of
Fatty acids and fats
elongating fatty acids by adding C2 units, but
is not able to introduce double bonds into the
A. Carboxylic acids
end sections of fatty acids (after C-9), arachi-
The naturally occurring fatty acids are carbox-
donic acid has to be supplied with the diet.
ylic acids with unbranched hydrocarbon
Linoleic and linolenic acid can be converted
chains of
4-24
carbon atoms. They are
into arachidonic acid by elongation, and they
present in all organisms as components of
can therefore replace arachidonic acid in the
fats and membrane lipids. In these com-
diet.
pounds, they are esterified with alcohols
(glycerol, sphingosine, or cholesterol). How-
B. Structure of fats
ever, fatty acids are also found in small
amounts in unesterified form. In this case,
Fats are esters of the trivalent alcohol glycerol
they are known as free fatty acids (FFAs). As
with three fatty acids. When a single fatty acid
free fatty acids have strongly amphipathic
is esterified with glycerol, the product is re-
properties (see p. 28), they are usually present
ferred to as a monoacylglycerol (fatty acid res-
in protein-bound forms.
idue = acyl residue).
The table lists the full series of aliphatic
Formally, esterification with additional
carboxylic acids that are found in plants and
fatty acids leads to diacylglycerol and ulti-
animals. In higher plants and animals, un-
mately to triacylglycerol, the actual fat (for-
branched, longchain fatty acids with either
merly termed “triglyceride”). As triacylglycer-
16 or 18 carbon atoms are the most common—
ols are uncharged, they are also referred to as
e. g., palmitic and stearic acid. The number of
neutral fats. The carbon atoms of glycerol are
carbon atoms in the longer, natural fatty acids
not usually equivalent in fats. They are distin-
is always even. This is because they are bio-
guished by their
“sn” number, where sn
synthesized from C2
building blocks
(see
stands for “stereospecific numbering.”
p. 168).
The three acyl residues of a fat molecule
Some fatty acids contain one or more
may differ in terms of their chain length and
isolated double bonds, and are therefore “un-
the number of double bonds they contain.
saturated.” Common unsaturated fatty acids
This results in a large number of possible
include oleic acid and linoleic acid. Of the two
combinations of individual fat molecules.
possible cis-trans isomers (see p. 8), usually
When extracted from biological materials,
only the cis forms are found in natural lipids.
fats always represent mixtures of very similar
Branched fatty acids only occur in bacteria. A
compounds, which differ in their fatty acid
shorthand notation with several numbers is
residues. A chiral center can arise at the mid-
used for precise characterization of the struc-
dle C atom (sn -C-2) of a triacylglycerol if the
ture of fatty acids—e g., 18:2;9,12 for linoleic
two external fatty acids are different. The
acid. The first figure stands for the number of
monoacylglycerols and diacylglycerols shown
C atoms, while the second gives the number
here are also chiral compounds. Nutritional
of double bonds. The positions of the double
fats contain palmitic, stearic, oleic acid, and
bonds follow after the semicolon. As usual,
linoleic acid particularly often. Unsaturated
numbering starts at the carbon with the high-
fatty acids are usually found at the central
est oxidation state (i. e., the carboxyl group
C atom of glycerol.
corresponds to C-1). Greek letters are also
The length of the fatty acid residues and
commonly used (α = C-2; β = C-3; ω = the
the number of their double bonds affect the
last carbon, ω-3 = the third last carbon).
melting point of the fats. The shorter the fatty
Essential fatty acids are fatty acids that
acid residues and the more double bonds they
have to be supplied in the diet. Without ex-
contain, the lower their melting points.
ception, these are all polyunsaturated fatty
acids: the C20
fatty acid arachidonic acid
(20:4;5,8,11,14) and the two C18 acids linoleic
acid
(18:2;9,12)
and linolenic acid
(18:3;9,12,15). The animal organism requires
arachidonic acid to synthesize eicosanoids
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipids
49
A. Carboxylic acids
Name
Number of carbons
Number of double bonds
Position of double bonds
Formic acid
0
1 :
Not contained
Acetic acid
2 :0
in lipids
Propionic acid
3 :0
Butyric acid
4 :0
Valerianic acid
5 :0
HOOC
CH2
CH2
CH2
CH2
CH3
Caproic acid
6 :0
Caprylic acid
8 :0
Caproic acid
Capric acid
10 :0
Lauric acid
12 :0
Myristic acid
14 :0
Palmitic acid
16 :0
Stearic acid
18 :0
Oleic acid
18 :1;
9
Linoleic acid
18 :2;
9,12
Linolenic acid
18 :3;
9,12,15
Arachidic acid
20 :0
Arachidonic acid
20 :4;
5,8,11,14
Behenic acid
22 :0
Erucic acid
22 :1;
13
Lignoceric acid
24 :0
Nervonic acid
24 :1;
15
Essential in human nutrition
B. Structure of fats
Ester bonding
Chiral center
sn number
O
O
O
C-1
HO CH2
R'
C
O
CH2
R'
C
O
CH2
R'
C
O CH2
O
O
C-2
HO
C H
HO
C H
R''
C
O
C
H
R''
C
O
C H
O
HO
CH2
HO
CH2
HO
CH2
R'''
C
O
CH2
C-3
Glycerol
Monoacylglycerol
Diacylglycerol
Triacylglycerol = Fat
Fatty acids
Glycerol
Acyl residue
Acyl residue
Van der Waals
model
Acyl residue
of tristearylglycerol
Rotatable
around C-C bond
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
50
Biomolecules
Some phospholipids carry additional
Phospholipids and glycolipids
charges, in addition to the negative charge
at the phosphate residue. In phosphatidylcho-
A. Structure of phospholipids and
line and phosphatidylethanolamine, the
glycolipids
N-atom of the amino alcohol is positively
Fats (triacylglycerol, 1) are esters of glycerol
charged. As a whole, these two phosphatides
with three fatty acids (see p. 48). Within the
therefore appear to be neutral. In contrast,
cell, they mainly occur as fat droplets. In the
phosphatidylserine—with one additional pos-
blood, they are transported in the hydropho-
itive charge and one additional negative
bic interior of lipoproteins (see p. 278).
charge in the serine residue—and phosphati-
Phospholipids
(2) are the main consti-
dylinositol (with no additional charge) have a
tuents of biological membranes
(see
negative net charge, due to the phosphate
pp. 214-217). Their common feature is a phos-
residue.
phate residue that is esterified with the hy-
Sphingolipids (3), which are found in large
droxyl group at C-3 of glycerol. Due to this
quantities in the membranes of nerve cells in
residue, phospholipids have at least one neg-
the brain and in neural tissues, have a slightly
ative charge at a neutral pH.
different structure from the other membrane
Phosphatidates (anions of the phosphatidic
lipids discussed so far. In sphingolipids, sphin-
acids), the simplest phospholipids, are phos-
gosine, an amino alcohol with an unsaturated
phate esters of diacylglycerol. They are impor-
alkyl side chain, replaces glycerol and one of
tant intermediates in the biosynthesis of fats
the acyl residues. When sphingosine forms an
and phospholipids
(see p. 170). Phosphati-
amide bond to a fatty acid, the compound is
dates can also be released from phospholipids
called ceramide (3). This is the precursor of
by phospholipases.
the sphingolipids. Sphingomyelin
(2)—the
The other phospholipids can be derived
most important sphingolipid—has an addi-
from phosphatidates
(residue
= phospha-
tional phosphate residue with a choline group
tidyl). Their phosphate residues are esterified
attached to it on the sphingosine, in addition
with the hydroxyl group of an amino alcohol
to the fatty acid.
(choline, ethanolamine, or serine) or with the
Glycolipids (3) are present in all tissues on
cyclohexane derivative myo-inositol. Phos-
the outer surface of the plasma membrane.
phatidylcholine is shown here as an example
They consist of sphingosine, a fatty acid, and
of this type of compound. When two phos-
an oligosaccharide residue, which can some-
phatidyl residues are linked with one glyc-
times be quite large. The phosphate residue
erol, the result is cardiolipin (not shown), a
typical of phospholipids is absent. Galacto-
phospholipid that is characteristic of the inner
sylceramide and glucosylceramide (known as
mitochondrial membrane. Lysophospholipids
cerebroside) are simple representatives of
arise from phospholipids by enzymatic cleav-
this group. Cerebrosides in which the sugar
age of an acyl residue. The hemolytic effect of
is esterified with sulfuric acid are known as
bee and snake venoms is due in part to this
sulfatides. Gangliosides are the most complex
reaction.
glycolipids. They constitute a large family of
Phosphatidylcholine (lecithin) is the most
membrane lipids with receptor functions that
abundant phospholipid in membranes.
are as yet largely unknown. A characteristic
Phosphatidylethanolamine (cephalin) has an
component of many gangliosides is N-acetyl-
ethanolamine residue instead of choline, and
neuraminic acid (sialic acid; see p. 38).
phosphatidylserine has a serine residue. In
phosphatidylinositol, phosphatidate is esteri-
fied with the sugarlike cyclic polyalcohol
myo-inositol. A doubly phosphorylated deriv-
ative of this phospholipid, phosphatidylinosi-
tol 4,5-bisphosphate, is a special component
of membranes, which, by enzymatic cleavage,
can give rise to two second messengers, diacyl-
glycerol (DAG) and inositol 1,4,5trisphosphate
(InsP3; see p. 386).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipids
51
A. Structure of fats, phospholipids, and glycolipids
O
Acyl residue 1
C O CH2
C O C
H
O
CH3
Acyl residue 2
O
H2C O P O
(CH2)2
N CH3
Acyl residue 3
O
CH3
Fat
Phosphatide
1. Fats
(phosphatidylcholine,
lecithin)
Acyl residue 1
Acyl residue 2
CH3
COO
HO
CH2
CH2
N CH3
HO
CH2
CH NH3
2
P
CH
3
Phosphatidate
Choline
Serine
OH OH
Acyl residue 1
H
OH
HO
CH2
CH2
NH3
H H
OH
H
Acyl residue 2
HO
H
H
OH
Amino alcohol
P
or sugar alcohol
Ethanolamine
myo-Inositol
Phosphatide
Acyl residue 1
Acyl residue 1
Sphingosine
Amino alcohol
Amino alcohol
P
P
or sugar alcohol
or sugar alcohol
Sphingophospholipid
Lysophospholipid
O
Acyl residue 1
C
NH
O
CH3
Sphingosine
H
H
C
CH2
O P O
(CH2)2
N CH3
C
C
O
CH3
P
Choline
C
H
OH
Sphingomyelin
H
2. Phospholipids
Acyl residue 1
Acyl residue 1
SO
Sphingosine
Sphingosine
3
Sugar
Ceramide
Sphingosine
Sulfatide
Acyl residue 1
Acyl residue 1
Sphingosine
Sphingosine
Sugar
Glc
Gal
GalNAc
Gal
Cerebroside
Ganglioside GM1
NeuAc
(galactosyl or glycosyl ceramide)
3. Sphingolipids
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
52
Biomolecules
tabolism, but cannot be produced by them
Isoprenoids
independently, are vitamins; this group
includes vitamins A, D, E, and K. Due to its
A. Activated acetic acid as a component of
structure and function, vitamin D is now usu-
lipids
ally classified as a steroid hormone
(see
Although the lipids found in plant and animal
pp. 56, 330).
organisms occur in many different forms,
Isoprene metabolism in plants is very com-
they are all closely related biogenetically;
plex. Plants can synthesize many types of ar-
they are all derived from acetyl-CoA, the “ac-
omatic substances and volatile oils from iso-
tivated acetic acid” (see pp. 12, 110).
prenoids. Examples include menthol (I= 2 ),
1. One major pathway leads from acetyl-
camphor (I = 2), and citronellal (I = 2). These
CoA to the activated fatty acids (acyl-CoA; for
C10 compounds are also called monoterpenes.
details, see p. 168). Fats, phospholipids, and
Similarly, compounds consisting of three iso-
glycolipids are synthesized from these, and
prene units (I = 3) are termed sesquiterpenes,
fatty acid derivatives in particular are formed.
and the steroids (I = 6) are called triterpenes.
Quantitatively, this is the most important
Isoprenoids that have hormonal and sig-
pathway in animals and most plants.
naling functions form an important group.
2. The second pathway leads from acetyl-
These include steroid hormones (I = 6) and
CoA to isopentenyl diphosphate (“active iso-
retinoate (the anion of retinoic acid; I = 3) in
prene”), the basic component for the isopren-
vertebrates, and juvenile hormone (I = 3) in
oids. Its biosynthesis is discussed in connec-
arthropods. Some plant hormones also belong
tion with biosynthesis of the isoprenoid, cho-
to the isoprenoids—e. g., the cytokinins, absci-
lesterol (see p. 172).
sic acid, and brassinosteroids.
Isoprene chains are sometimes used as
lipid anchors to fix molecules to membranes
B. Isoprenoids
(see p. 214). Chlorophyll has a phytyl residue (I
Formally, isoprenoids are derived from a sin-
= 4) as a lipid anchor. Coenzymes with iso-
gle common building block, isoprene
(2-
prenoid anchors of various lengths include
methyl-1,3-butadiene), a methyl-branched
ubiquinone (coenzyme Q; I = 6-10), plastoqui-
compound with five C atoms. Activated
none (I = 9), and menaquinone (vitamin K; I =
isoprene, isopentenyl diphosphate, is used by
4-6). Proteins can also be anchored to mem-
plants and animals to biosynthesize linear
branes by isoprenylation.
and cyclic oligomers and polymers. For the
In some cases, an isoprene residue is used
isoprenoids listed here—which only represent
as an element to modify molecules chemi-
a small selection—the number of isoprene
cally. One example of this is N'-isopentenyl-
units (I) is shown.
AMP, which occurs as a modified component
From activated isoprene, the metabolic
in tRNA.
pathway leads via dimerization to activated
geraniol (I = 2) and then to activated farnesol (I
= 3). At this point, the pathway divides into
two. Further extension of farnesol leads to
chains with increasing numbers of isoprene
units—e. g., phytol (I = 4), dolichol (I = 14-24),
and rubber (I = 700-5000). The other pathway
involves a “head-to-head” linkage between
two farnesol residues, giving rise to squalene
(I = 6), which, in turn, is converted to choles-
terol (I = 6) and the other steroids.
The ability to synthesize particular iso-
prenoids is limited to a few species of plants
and animals. For example, rubber is only
formed by a few plant species, including the
rubber tree (Hevea brasiliensis). Several iso-
prenoids that are required by animals for me-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipids
53
A. Activated acetic acid as a component of lipids
Activated acetic acid
O
Isoprene
H3C
C
S
A
2
Acetyl-CoA
1
Activated fatty acid
Active isoprene
O
C
S
O P P
A
Acyl-CoA
Isopentenyl diphosphate
Fats
Phospholipids Glycolipids
Isoprenoids
B. Isoprenoids
Biosynthesis only in plants
Metabolite
Building block of
and micro-organisms
modified
all isoprenoids
with isoprene
Camphor
Isopentenyl-AMP
Active isoprene
I = 2
I =1
I =1
CH3
CH3
O
H3C
Chain-like
Citronellol
Geraniol
isoprenoids
I = 2
I = 2
Menthol
I = 2
Juvenile hormone
Farnesol
Squalene
Cholesterol
I = 3
I = 3
I = 6
I = 6
CH2OH
Retinoic
Phytol
HO
I = 4
Carotenoids
CH2OH
(Vitamin A)
Steroid hormones
CH2OH
Bile acids
Menaquinone
I = 4 - 6
Steroid glycosides
I = 6
Dolichol
Plastoquinone
I = 14 - 24
I = 9
Tocopherol
CH3
(Vitamin E)
Cyclic
isoprenoids
HO
I = 4
Rubber
I = 700 - 5 000
H3C
O
CH3
O
CH3
Ubiquinone
H3C
O
CH3
I = 6 - 10
OH
Phylloquinone
H3C
O
CH3
(Vitamin K)
O
n = 6-10
I = 4
OH
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
54
Biomolecules
onto which the much more mobile side chain
Steroid structure
is attached.
Steroids are relatively apolar (hydropho-
A. Steroid building blocks
bic). Some polar groups—e. g., hydroxyl and
Common to all of the steroids is a molecular
oxo groups—give them amphipathic proper-
core structure consisting of four saturated
ties. This characteristic is especially pro-
rings, known as gonane. At the end of the
nounced with the bile acids (see p. 314).
steroid core, many steroids also carry a side
chain, as seen in cholestane, the basic compo-
C. Thin-layer chromatography
nent of the sterols (steroid alcohols).
Thin-layer chromatography (TLC) is a power-
ful, mainly analytic, technique for rapidly sep-
B. Spatial structure
arating lipids and other small molecules such
The four rings of the steroids are distin-
as amino acids, nucleotides, vitamins, and
guished using the letters A, B, C, and D. Due
drugs. The sample being analyzed is applied
to the tetrahedral arrangement of the single
to a plate made of glass, aluminum, or plastic,
carbon bonds, the rings are not flat, but puck-
which is covered with a thin layer of silica gel
ered. Various ring conformations are known
or other material (1). The plate is then placed
by the terms “chair,” “boat,” and “twisted”
in a chromatography chamber that contains
(not shown). The chair and boat conforma-
some solvent. Drawn by capillary forces, the
tions are common. Fivemembered rings fre-
solvent moves up the plate (2). The substan-
quently adopt a conformation referred to as
ces in the sample move with the solvent. The
an “envelope”. Some rings can be converted
speed at which they move is determined by
from one conformation to another at room
their distribution between the stationary
temperature, but with steroids this is dif cult.
phase (the hydrophilic silica), and the mobile
Substituents of the steroid core lie either
phase (the hydrophobic solvent). When the
approximately in the same plane as the ring
solvent reaches the top edge of the plate,
(e = equatorial) or nearly perpendicular to it
the chromatography is stopped. After evapo-
(a = axial). In threedimensional representa-
ration of the solvent, the separated substan-
tions, substituents pointing toward the ob-
ces can be made visible using appropriate
server are indicated by an unbroken line (β
staining methods or with physical processes
position), while bonds pointing into the plane
(e. g., ultraviolet light) (3). The movement of a
of the page are indicated by a dashed line (α
substance in a given TLC system is expressed
position). The so-called angular methyl
as its Rf value. In this way, compounds that are
groups at C-10 and C-13 of the steroids always
not known can be identified by comparison
adopt the β position.
with reference substances.
Neighboring rings can lie in the same plane
A process in which the polarity of the sta-
(trans; 2) or at an angle to one another (cis; 1).
tionary and mobile phases is reversed—i. e.,
This depends on the positions of the substitu-
the stationary phase is apolar and the solvent
ents of the shared ring carbons, which can be
is polar—is known as “reversed-phase thin-
arranged either cis or trans to the angular
layer chromatography” (RP-TLC).
methyl group at C-10. The substituents of ste-
roid that lie at the points of intersection of the
individual rings are usually in trans position.
As a whole, the core of most steroids is more
or less planar, and looks like a flat disk. The
only exceptions to this are the ecdysteroids,
bile acids (in which A:B is cis), cardiac glyco-
sides, and toad toxins.
A more realistic impression of the three-
dimensional structure of steroids is provided
by the space-filling model of cholesterol (3).
The four rings form a fairly rigid scaffolding,
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipids
55
A. Steroid building blocks
B. 3D structure
Angular
21
22
24
27
methyl groups
18
25
20
23
Gonane
17
26
111213
19
C
D
16
14
15
Methyl-branched side
1
9
chain with 8 carbons
2
10
8
A
B
3
5
7
4
HO
6
Cholesterol
Double bond
Cholestane
Hydroxyl group
in ring B: ∆5
at C-3 adopts
β conformation
β con-
CH3
R
H and CH
3 in cis position
formation,
β con-
axial
formation,
CH3
H
equatorial
CH3
H
H
HO
Cholestanol
H
H and CH3 in
α con-
H
transposition
1.
formation,
2.
axial
3.
Chair
Boat
Envelope
Ring conformations
Cholesterol (Van der Waals model)
C. Thin-layer chromatography
Thin-
Chromato-
Front
Cholesterol
layer
graphy tank
esters
plate with
Triacyl-
silica gel
glycerols
surface
a
Rf =b
Free fatty acids
a
Cholesterol
b
Running
1,3-
Diacyl-
solvent:
1,2-
glycerols
Hexane/
Diethylether/
Monoacyl-
Sample:
Formic acid
glycerols
lipid
80 : 80 : 2
mixture
(v/v/v)
Start
Phospholipids
1. Load
2. Develop
3. Make visible
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
56
Biomolecules
A and B are in cis position relative to each
Steroids: overview
other
(see p. 54). One to three hydroxyl
The three most important groups of steroids
groups (in α position) are found in the steroid
are the sterols, bile acids, and steroid hor-
core at positions 3, 7, and 12. Bile acids keep
mones. Particularly in plants, compounds
bile cholesterol in a soluble state as micelles
with steroid structures are also found that
and promote the digestion of lipids in the
are notable for their pharmacological ef-
intestine (see p. 270). Cholic acid and cheno-
fects—steroid alkaloids, digitalis glycosides,
deoxycholic acid are primary bile acids that
and saponins.
are formed by the liver. Their dehydroxylation
at C-7 by microorganisms from the intestinal
flora gives rise to the secondary bile acids
A. Sterols
lithocholic acid and deoxycholic acid.
Sterols are steroid alcohols. They have a
β-positioned hydroxyl group at C-3 and one
C. Steroid hormones
or more double bonds in ring B and in the side
chain. There are no further oxygen functions,
The conversion of cholesterol to steroid
as in the carbonyl and carboxyl groups.
hormones (see p. 376) is of minor importance
The most important sterol in animals is
quantitatively, but of major importance in
cholesterol. Plants and microorganisms have
terms of physiology. The steroid hormones
a wide variety of closely related sterols in-
are a group of lipophilic signal substances
stead of cholesterol—e. g., ergosterol, β-sitos-
that regulate metabolism, growth, and repro-
terol, and stigmasterol.
duction (see p. 374).
Cholesterol is present in all animal tissues,
Humans have six steroid hormones:
and particularly in neural tissue. It is a major
progesterone, cortisol, aldosterone, testos-
constituent of cellular membranes, in which it
terone, estradiol, and calcitriol. With the ex-
regulates fluidity (see p. 216). The storage and
ception of calcitriol, these steroids have either
transport forms of cholesterol are its esters
no side chain or only a short side one consist-
with fatty acids. In lipoproteins, cholesterol
ing of two carbons. Characteristic for most of
and its fatty acid esters are associated with
them is an oxo group at C-3, conjugated with
other lipids (see p. 278). Cholesterol is a con-
a double bond between C-4 and C-5 of ring A.
stituent of the bile and is therefore found in
Differences occur in rings C and D. Estradiol is
many gallstones. Its biosynthesis, metabo-
aromatic in ring A, and its hydroxyl group at
lism, and transport are discussed elsewhere
C-3 is therefore phenolic. Calcitriol differs
(see pp. 172, 312).
from other vertebrate steroid hormones; it
Cholesterol-rich lipoproteins of the LDL
still contains the complete carbon framework
type are particularly important in the devel-
of cholesterol, but lightdependent opening of
opment of arteriosclerosis, in which the arte-
ring B turns it into what is termed a “secoste-
rial walls are altered in connection with an
roid” (a steroid with an open ring).
excess plasma cholesterol level. In terms of
Ecdysone is the steroid hormone of the
dietary physiology, it is important that plant
arthropods. It can be regarded as an early
foodstuffs are low in cholesterol. By contrast,
form of the steroid hormones. Steroid hor-
animal foods can contain large amounts of
mones with signaling functions also occur in
cholesterol—particularly butter, egg yolk,
plants.
meat, liver, and brain.
B. Bile acids
Bile acids are synthesized from cholesterol in
the liver
(see p. 314). Their structures can
therefore be derived from that of cholesterol.
Characteristic for the bile acids is a side chain
shortened by three C atoms in which the last
carbon atom is oxidized to a carboxyl group.
The double bond in ring B is reduced and rings
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipids
57
A. Sterols
Animal
Ergosterol
Plant
sterol
sterols
HO
HO
Cholesterol
HO
Stigmasterol
β-Sitosterol
HO
B. Bile acids
O
Lithocholic acid
OH
O
OH
OH
O
HO
OH
O
HO
OH
OH
OH
H
Cholic acid
HO
OH
Cheno-
deoxycholic acid
Deoxy-
HO
cholic acid
C. Steroid hormones
CH2OH
CH2OH
C O
C O
OH
OH
OHC
HO
HO
OH
O
HO
O
O
Cortisol
Aldosterone
Testosterone
Estradiol
CH3
OH
25
C O
OH
OH
HO
OH
O
HO
CH2
Molting hormone
O
of insects,
3
1
spiders
Progesterone
Calcitriol
Ecdysone
HO
OH
and crabs
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
58
Biomolecules
mensional structure as follows: firstly, the
Amino acids: chemistry and
tetrahedron is rotated in such a way that the
properties
most oxidized group (the carboxylate group)
is at the top. Rotation is then continued until
the line connecting line COO- and R (red) is
A. Amino acids: functions
level with the page. In L-amino acids, the
The amino acids (2-aminocarboxylic acids)
NH3+ group is then on the left, while in D-
fulfill various functions in the organism.
amino acids it is on the right.
Above all, they serve as the components of
peptides and proteins. Only the 20 proteino-
C. Dissociation curve of histidine
genic amino acids (see p. 60) are included in
the genetic code and therefore regularly
All amino acids have at least two ionizable
found in proteins. Some of these amino acids
groups, and their net charge therefore de-
undergo further (post-translational) change
pends on the pH value. The COOH groups at
following their incorporation into proteins
the α-C atom have pKa values of between 1.8
(see p. 62). Amino acids or their derivatives
and 2.8 and are therefore more acidic than
are also form components of lipids—e. g., ser-
simple monocarboxylic acids. The basicity of
ine in phospholipids and glycine in bile salts.
the α-amino function also varies, with pKa
Several
amino acids function
as
values of between 8.8 and 10.6, depending
neurotransmitters themselves
(see p. 352),
on the amino acid. Acidic and basic amino
while others are precursors of neurotransmit-
acids have additional ionizable groups in their
ters, mediators, or hormones
(see p. 380).
side chain. The pKa values of these side chains
Amino acids are important (and sometimes
are listed on p. 60. The electrical charges of
essential) components of food (see p. 360).
peptides and proteins are mainly determined
Specific amino acids form precursors for other
by groups in the side chains, as most α-car-
metabolites—e. g., for glucose in gluconeogen-
boxyl and α-amino functions are linked to
esis, for purine and pyrimidine bases, for
peptide bonds (see p. 66).
heme, and for other molecules. Several non-
Histidine can be used here as an example of
proteinogenic amino acids function as inter-
the pH-dependence of the net charge of an
mediates in the synthesis and breakdown of
amino acid. In addition to the carboxyl group
proteinogenic amino acids (see p. 412) and in
and the amino group at the α-C atom with pKa
the urea cycle (see p. 182).
values of 1.8 and 9.2, respectively, histidine
also has an imidazole residue in its side chain
with a pKa value of 6.0. As the pH increases,
B. Optical activity
the net charge (the sum of the positive and
The natural amino acids are mainly α-amino
negative charges) therefore changes from +2
acids, in contrast to β-amino acids such as β-
to -1. At pH 7.6, the net charge is zero, even
alanine and taurine. Most α-amino acids have
though the molecule contains two almost
four different substituents at C-2 (Cα). The α
completely ionized groups in these condi-
atom therefore represents a chiral center—i. e.,
tions. This pH value is called the isoelectric
there are two different enantiomers (L- and
point.
D-amino acids; see p. 8). Among the proteino-
At its isoelectric point, histidine is said to
genic amino acids, only glycine is not chiral (R
be zwitterionic, as it has both anionic and
= H). In nature, it is almost exclusively
cationic properties. Most other amino acids
L-amino acids that are found. D-Amino acids
are also zwitterionic at neutral pH. Peptides
occur in bacteria—e. g., in murein
(see
and proteins also have isoelectric points,
p. 40)—and in peptide antibiotics. In animal
which can vary widely depending on the
metabolism, D-Amino acids would disturb
composition of the amino acids.
the enzymatic reactions of L-amino acids
and they are therefore broken down in the
liver by the enzyme D-amino acid oxidase.
The Fischer projection (center) is used to
present the formulas for chiral centers in bio-
molecules. It is derived from their three-di-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Amino Acids
59
A. Amino acids: functions
Components of:
Precursors of:
Peptides
Keto acids
Proteins
Biogenic amines
Phospholipids
Glucose
Nucleotides
Heme, creatine
Neurotransmitters:
Transport molecule for:
R
Glutamate
NH2 groups
Aspartate
Glycine
L-Amino acid
B. Optical activity
COO
COO
C
R
H3N
C
H
H3N
R
H
L-Amino acid
D-Amino acid
Fischer projections
(mirror image)
C. Dissociation curve of histidine
pH
COO
COOH
H3N
C
H
H3N
C
H
2
pK1
CH2
CH2
1.8
4
C
C
HN
CH
HN
CH
HC NH
HC NH
pK2
6
6.0
pH 0.5
pH 5
8
COO
COO
pK3
H3N
C
H
9.2
H2N
C
H
10
CH2
CH2
C
C
HN
CH
12
HN
CH
HC N
HC N
pH 7.6
pH 11
-1
0
+1
+2
(Isoelectric point)
Net charge
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
60
Biomolecules
The aromatic amino acids (class III) contain
Proteinogenic amino acids
resonancestabilized rings. In this group, only
phenylalanine has strongly apolar properties.
A. The proteinogenic amino acids
Tyrosine and tryptophan are moderately polar,
The amino acids that are included in the ge-
and histidine is even strongly polar. The imi-
netic code (see p. 248) are described as “pro-
dazole ring of histidine is already protonated
teinogenic.” With a few exceptions (see p. 58),
at weakly acidic pH values. Histidine, which is
only these amino acids can be incorporated
only aromatic in protonated form (see p. 58),
into proteins through translation. Only the
can therefore also be classified as a basic
side chains of the 20 proteinogenic amino
amino acid. Tyrosine and tryptophan show
acids are shown here. Their classification is
strong light absorption at wavelengths of
based on the chemical structure of the side
250-300 nm.
chains, on the one hand, and on their polarity
The neutral amino acids (class IV) have
on the other (see p. 6). The literature includes
hydroxyl groups (serine, threonine) or amide
several slightly different systems for classify-
groups (asparagine, glutamine). Despite their
ing amino acids, and details may differ from
nonionic nature, the amide groups of aspara-
those in the system used here.
gine and glutamine are markedly polar.
The carboxyl groups in the side chains of
For each amino acid, the illustration names:
the acidic amino acids aspartic acid and glu-
• Membership of structural classes I-VII (see
tamic acid (class V) are almost completely
below; e. g., III and VI for histidine)
ionized at physiological pH values. The side
• Name and abbreviation, formed from the
chains of the basic amino acids lysine and
first three letters of the name (e. g., histi-
arginine are also fully ionized—i. e., positively
dine, His)
charged—at neutral pH. Arginine, with its
• The one-letter symbol introduced to save
positively charge guanidinium group, is par-
space in the electronic processing of se-
ticularly strongly basic, and therefore ex-
quence data (H for histidine)
tremely polar.
• A quantitative value for the polarity of the
Proline (VII) is a special case. Together with
side chain (bottom left; 10.3 for histidine).
the α-C atom and the α-NH2 group, its side
The more positive this value is, the more
chain forms a fivemembered ring. Its nitrogen
polar the amino acid is.
atom is only weakly basic and is not proto-
In addition, the polarity of the side chains is
nated at physiological pH. Due to its ring
indicated by color. It increases from yellow,
structure, proline causes bending of the pep-
through light and dark green, to bluish green.
tide chain in proteins (this is important in
For ionizing side chains, the corresponding
collagen, for example; see p. 70).
pKa values are also given (red numbers).
Several proteinogenic amino acids cannot
The aliphatic amino acids (class I) include
be synthesized by the human organism, and
glycine, alanine, valine, leucine, and isoleucine.
therefore have to be supplied from the diet.
These amino acids do not contain heteroa-
These essential amino acids (see p. 360) are
toms (N, O, or S) in their side chains and do
marked with a star in the illustration. Histi-
not contain a ring system. Their side chains
dine and possibly also arginine are essential
are markedly apolar. Together with threonine
for infants and small children.
(see below), valine, leucine, and isoleucine
form the group of branched-chain amino
acids. The sulfurcontaining amino acids cys-
teine and methionine (class II), are also apolar.
However, in the case of cysteine, this only
applies to the undissociated state. Due to its
ability to form disulfide bonds, cysteine plays
an important role in the stabilization of pro-
teins (see p. 72). Two cysteine residues linked
by a disulfide bridge are referred to as cystine
(not shown).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Amino Acids
61
A. The proteinogenic amino acids
Aliphatic
Sulfur-containing
Glycine
Alanine
Valine
Leucine
Isoleucine
Cysteine
Methionine
(Gly, G)
(Ala, A)
(Val, V)
(Leu, L)
(Ile, I)
(Cys, C)
(Met, M)
H
CH3
H3C
CH
CH
H3C
C
H
CH2
CH2
2
CH3
H3C
CH
CH2
SH
CH2
8.3
CH3
CH3
S
CH3
pKa value
Polarity
-2.4
-1.9
-2.0
-2.3
-2.2
-1.2
-1.5
Aromatic
Cyclic
Neutral
Phenylalanine
Tyrosine
Tryptophan
Proline
Serine
Threonine
(Phe, F)
(Tyr, Y)
(Trp, W)
(Pro, P)
(Ser, S)
(Thr, T)
CH2
CH2
CH2
COO
CH
2
H3C
C
H
CH
OH
OH
HN
CH2
N
H2C CH2
H
Indole ring
Pyrrolidine ring
OH
10.1
+0.8
+6.1
+5.9
+6.0
+5.1
+4.9
Essential amino acids
Chiral center
Neutral
Acidic
Basic
Asparagine
Glutamine
Aspartic acid
Glutamic acid
Histidine
Lysine
Arginine
(Asn,N)
(Gln, Q)
(Asp, D)
(Glu, E)
(His, H)
(Lys,K)
(Arg, R)
CH2
CH2
CH2
CH2
CH2
CH2
CH2
CONH2
CH2
COO
CH2
CH2
CH2
4.0
HN CH
CONH2
COO
CH2
CH2
HC N
4.3
6.0
CH2
NH
Imidazole ring
NH3
C
10.8
H2N
NH2
12.5
+9.7
+9.4
+11.0
+10.2
+10.3
+15.0
+20.0
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
62
Biomolecules
The ε-amino group of lysine residues is sub-
Non-proteinogenic amino acids
ject to a particularly large number of modifi-
In addition to the 20 proteinogenic amino
cations. Its acetylation (or deacetylation) is an
acids (see p. 60), there are also many more
important mechanism for controlling genetic
compounds of the same type in nature. These
activity
(see p. 244). Many coenzymes and
arise during metabolic reactions (A) or as a
cofactors are covalently linked to lysine resi-
result of enzymatic modifications of amino
dues. These include biotin (see p. 108), lipoic
acid residues in peptides or proteins (B). The
acid
(see p. 106), and pyridoxal phosphate
“biogenic amines” (C) are synthesized from α-
(see p. 108), as well as retinal (see p. 358).
amino acids by decarboxylation.
Covalent modification with ubiquitin marks
proteins for breakdown (see p. 176). In colla-
gen, lysine and proline residues are modified
A. Rare amino acids
by hydroxylation to prepare for the formation
Only a few important representatives of the
of stable fibrils (see p. 70). Cysteine residues
non-proteinogenic amino acids are men-
form disulfide bonds with one another (see
tioned here. The basic amino acid ornithine
p. 72). Cysteine prenylation serves to anchor
is an analogue of lysine with a shortened side
proteins in membranes (see p. 214). Covalent
chain. Transfer of a carbamoyl residue to or-
bonding of a cysteine residue with heme oc-
nithine yields citrulline. Both of these amino
curs in cytochrome c. Flavins are sometimes
acids are intermediates in the urea cycle (see
covalently bound to cysteine or histidine res-
p. 182). Dopa (an acronym of 3,4-dihydroxy-
idues of enzymes. Among the modifications of
phenylalanine) is synthesized by hydroxyla-
tyrosine residues, conversion into iodinated
tion of tyrosine. It is an intermediate in the
thyroxine (see p. 374) is particularly interest-
biosynthesis of catecholamines (see p. 352)
ing.
and of melanin. It is in clinical use in the
treatment of Parkinson’s disease. Selenocys-
C. Biogenic amines
teine, a cysteine analogue, occurs as a compo-
nent of a few proteins—e. g., in the enzyme
Several amino acids are broken down by de-
glutathione peroxidase (see p. 284).
carboxylation. This reaction gives rise to what
are known as biogenic amines, which have
various functions. Some of them are compo-
B. Post-translational protein modification
nents of biomolecules, such as ethanolamine
Subsequent alteration of amino acid residues
in phospholipids (see p. 50). Cysteamine and
in finished peptides and proteins is referred
b-alanine are components of coenzyme A (see
to as post-translational modification. These re-
p. 12) and of pantetheine (see pp. 108, 168).
actions usually only involve polar amino acid
Other amines function as signaling substan-
residues, and they serve various purposes.
ces. An important neurotransmitter derived
The free α-amino group at the N-terminus
from glutamate is γ-aminobutyrate (GABA,
is blocked in many proteins by an acetyl res-
see p. 356). The transmitter dopamine is also
idue or a longer acyl residue (acylation). N-
a precursor for the catecholamines epineph-
terminal glutamate can cyclize into a pyroglu-
rine and norepinephrine (see p. 352). The bio-
tamate residue, while the C-terminal carbox-
genic amine serotonin, a substance that has
ylate group can be present in an amidated
many effects, is synthesized from tryptophan
form (see TSH, p. 380). The side chains of ser-
via the intermediate 5-hydroxytryptophan.
ine and asparagine residues are often linked
Monamines are inactivated into aldehydes
to oligosaccharides (glycosylation, see p. 230).
by amine oxidase
(monoamine oxidase,
Phosphorylation of proteins mainly affects
“MAO”) with deamination and simultaneous
serine and tyrosine residues. These reactions
oxidation. MAO inhibitors therefore play an
have mainly regulatory functions (see p. 114).
important role in pharmacological interven-
Aspartate and histidine residues of enzymes
tions in neurotransmitter metabolism.
are sometimes phosphorylated, too. A special
modification of glutamate residues, J-carbox-
ylation, is found in coagulation factors. It is
essential for blood coagulation (see p. 290).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Amino Acids
63
A. Rare amino acids
COO
COO
COO
COO
H3N
C
H
H3N
C
H
H3N
C
H
H3N
C
H
CH2
CH2
CH2
CH2
CH2
CH2
Se H
CH2
CH2
OH
NH3
HN C NH2
OH
O
Seleno-
Ornithine
Citrulline
L-Dopa
cysteine
B. Post-translational protein modification
D: Pyroglutamyl-
B: Oligo-
B: Oligo-
D: Phospho-
D: Acetyl-
B: Pyridoxal-
Acetyl-
saccharide
saccharide
Methyl-
Methyl-
Liponat
(O-glyco-
(N-glyco-
Formyl-
γ-Carboxy-
γ-Hydroxy-
Biotin
sylation)
sylation)
Myristoyl-
(Glu)
Retinal
Ubiquitin
OH
CONH
COO
NH3
2
H3N
Ser, Thr
Asn, Gln
Asp, Glu
Lys
D: derivative B: bonds with
Pro
Tyr
Phe
His
Cys
OOC
D: 3-Hydroxy-
SH
HN N
4-Hydroxy-
OH
D: Amido-
D: Phospho-
D: 4-Hydroxy-
D: Phospho-
D: Disulfide
(CONH2)
Iodo-
(Tyrosine)
Methyl-
Prenyl-
Sulfato-
B: Flavin
B: Heme
Adenyl-
Flavin
C. Biogenic amines
Amino acid
Amine
Function
Amino acid
Amine
Function
Serine
Ethanol-
Glutamate
Glutamate
γ-Amino-
Neurotrans-
amine
butyrate
mitter (GABA)
Cysteine
Cysteamine
Component of
Histidine
Histamine
Mediator, neuro-
coenzyme A
transmitter
Threonine
Amino-
Component of
Dopa
Dopamine
Neurotransmitter
propanol
vitamin B12
Aspartate
β-Alanine
Component of
5-Hydroxy-
Serotonin
Mediator, neuro-
coenzyme A
tryptophan
transmitter
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
64
Biomolecules
hormone somatotropin and its receptor is
Peptides and proteins: overview
shown here as an example (middle). Here,
the extracellular domains of two receptor
A. Proteins
molecules here bind one molecule of the hor-
When amino acids are linked together by
mone. This binding activates the cytoplasmic
acid-amide bonds, linear macromolecules
domains of the complex, leading to further
(peptides) are produced. Those containing
conduction of the signal to the interior of
more than ca. 100 amino acid residues are
the cell
(see p. 384). The small peptide
described as proteins (polypeptides). Every
hormone insulin is discussed in detail else-
organism contains thousands of different pro-
where (see pp. 76, 160). DNA-binding proteins
teins, which have a variety of functions. At a
(transcription factors; see p. 118) are decisively
magnification of ca. 1.5 million, the semi-
involved in regulating the metabolism and in
schematic illustration shows the structures
differentiation processes. The structure and
of a few intra and extracellular proteins, giv-
function of the catabolite activator protein
ing an impression of their variety. The func-
(top left) and similar bacterial transcription
tions of proteins can be classified as follows.
factors have been particularly well investi-
Establishment and maintenance of struc-
gated.
ture. Structural proteins are responsible for
Catalysis. Enzymes, with more than 2000
the shape and stability of cells and tissues. A
known representatives, are the largest group
small part of a collagen molecule is shown as
of proteins in terms of numbers (see p. 88).
an example (right; see p. 70). The complete
The smallest enzymes have molecular masses
molecule is 1.5
300 nm in size, and at the
of
10-15 kDa. Intermediatesized enzymes,
magnification used here it would be as long as
such as alcohol dehydrogenase (top left) are
three pages of the book. Histones are also
around
100-200 kDa, and the largest—
structural proteins. They organize the ar-
including glutamine synthetase with its 12
rangement of DNA in chromatin. The basic
monomers (top right)—can reach more than
components of chromatin, the nucleosomes
500 kDa.
(top right; see p. 218) consist of an octameric
Movement. The interaction between actin
complex of histones, around which the DNA is
and myosin is responsible for muscle contrac-
coiled.
tion and cell movement (see p. 332). Myosin
Transport. A wellknown transport protein
(right), with a length of over
150 nm, is
is hemoglobin in the erythrocytes (bottom
among the largest proteins there are. Actin
left). It is responsible for the transport of oxy-
filaments (F-actin) arise due to the polymer-
gen and carbon dioxide between the lungs
ization of relatively small protein subunits (G-
and tissues (see p. 282). The blood plasma
actin). Along with other proteins, tropomyo-
also contains many other proteins with trans-
sin, which is associated with F-actin, controls
port functions. Prealbumin
(transthyretin;
contraction.
middle), for example, transports the thyroid
Storage. Plants contain special storage pro-
hormones thyroxin and triiodothyronine. Ion
teins, which are also important for human
channels and other integral membrane pro-
nutrition
(not shown). In animals, muscle
teins (see p. 220) facilitate the transport of
proteins constitute a nutrient reserve that
ions and metabolites across biological mem-
can be mobilized in emergencies.
branes.
Protection and defense. The immune sys-
tem protects the body from pathogens and
foreign substances. An important component
of this system is immunoglobulin G (bottom
left; see p. 300). The molecule shown here is
bound to an erythrocyte by complex forma-
tion with surface glycolipids (see p. 292).
Control and regulation. In biochemical sig-
nal chains, proteins function as signaling sub-
stances (hormones) and as hormone recep-
tors. The complex between the growth
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Peptides and Proteins
65
A. Proteins
Catabolite activator
DNA
Nucleosome
Nucleus
protein
Cyto-
plasm
Alcohol
Glutamine
dehydrogenase
synthetase
Ion channel
Somatotropin
receptor
(dimer)
Blood
Somatotropin
Water
Prealbumin
Cholesterol
Glucose
Insulin
Collagen
triple helix
Immuno-
Erythrocyte
Tropo-
globulin G
myosin
F-Actin
Hemoglobin
Myosin
10 nm
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
66
Biomolecules
example, the peptide hormone angiotensin II
Peptide bonds
(see p. 330) has the sequence Asp-Arg-Val-
Tyr-Ile-His-Pro-Phe, or DRVYIHPF.
A. Peptide bond
The amino acid components of peptides and
D. Conformational space of the peptide
proteins are linked together by amide bonds
chain
(see p. 60) between α-carboxyl and α-amino
groups. This type of bonding is therefore also
With the exception of the terminal residues,
known as peptide bonding. In the dipeptide
every amino acid in a peptide is involved in
shown here, the serine residue has a free
two peptide bonds (one with the preceding
ammonium group, while the carboxylate
residue and one with the following one). Due
group in alanine is free. Since the amino acid
to the restricted rotation around the C-N
with the free NH3+ group is named first, the
bond, rotations are only possible around the
peptide is known as seryl alanine, or in abbre-
N-Cα and Cα-C bonds (2). As mentioned
viated form Ser-Ala or SA.
above, these rotations are described by the
dihedral angles φ (phi) and ψ (psi). The angle
describes rotation around the N-Cα bond; ψ
B. Resonance
describes rotation around Cα-C—i. e., the po-
Like all acid-amide bonds, the peptide bond is
sition of the subsequent bond.
stabilized by resonance (see p. 4). In the con-
For steric reasons, only specific combina-
ventional notation (top right) it is represented
tions of the dihedral angles are possible.
as a combination of a C=O double bond with a
These relationships can be illustrated clearly
C-N single bond. However, a C=N double bond
by a so-called φ/ ψ diagram (1). Most combi-
with charges at O and N could also be written
nations of φ and ψ are sterically “forbidden”
(middle). Both of these are only extreme cases
(red areas). For example, the combination φ =
of electron distribution, known as resonance
0° and ψ = 180° (4) would place the two
structures. In reality, the π electrons are
carbonyl oxygen atoms less than
115 pm
delocalized throughout all the atoms (bot-
apart—i. e., at a distance much smaller than
tom). As a mesomeric system, the peptide
the sum of their van der Waals radii (see p. 6).
bond is planar. Rotation around the C-N
Similarly, in the case of φ = 180° and ψ = 0° (5),
bond would only be possible at the expense
the two NH hydrogen atoms would collide.
of large amounts of energy, and the bond is
The combinations located within the green
therefore not freely rotatable. Rotations are
areas are the only ones that are sterically
only possible around the single bonds marked
feasible (e. g., 2 and 3). The important secon-
with arrows. The state of these is expressed
dary structures that are discussed in the fol-
using the angles φ and ψ (see D). The plane in
lowing pages are also located in these areas.
which the atoms of the peptide bond lie is
The conformations located in the yellow areas
highlighted in light blue here and on the fol-
are energetically less favorable, but still pos-
lowing pages.
sible.
The φ/ ψ diagram (also known as a Rama-
chandran plot) was developed from modeling
C. Peptide nomenclature
studies of small peptides. However, the con-
Peptide chains have a direction and therefore
formations of most of the amino acids in pro-
two different ends. The amino terminus (N
teins are also located in the permitted areas.
terminus) of a peptide has a free ammonium
The corresponding data for the small protein,
group, while the carboxy terminus (C termi-
insulin (see p. 76), are represented by black
nus) is formed by the carboxylate group of the
dots in 1.
last amino acid. In peptides and proteins, the
amino acid components are usually linked in
linear fashion. To express the sequence of a
peptide, it is therefore suf cient to combine
the three-letter or single-letter abbreviations
for the amino acid residues (see p. 60). This
sequence always starts at the N terminus. For
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Peptides and Proteins
67
A. Peptide bonds
B. Resonance
H
C N
O
Resonance
structures
H
C N
O
ϕ ψ
H
C N
Seryl alanine
O
(Ser-Ala, H3N-Ser-Ala-COO
, SA)
Mesomeric
structure
C. Peptide nomenclature
Residue 1
Residue 2
Residue 3
Residue 4
Residue
R1
H
O R3
H O
Rn
Amino
H
H
Carboxy-
terminus
C
N
C
C
N
C
C
terminus
(N terminus)
H3N
C
C
N
C
C
N
COO
(C terminus)
H
H
H
O R2
H O R4
H
D. Conformation space of the peptide chain
ϕ =
0°
3
ϕ (Phi): Rotation about
ψ = 180°
R
ϕ
j
(Phi): N - Cα
2
R
ϕ
ψ (Psi): Rotation about
2
ψ
y (Psi): Cα - C
ψ
3
ϕ = -139°
180
1
1
ψ = -135°
B-23
C
βa
120
d = 115 pm
4.
2.
β
p
60
αl
Allowed
Forbidden
ψ 0
αr
ϕ = 180°
3
-60
ψ=
0°
ϕ = -57°
R
ψ= -47°
d =
-120
ϕ
A-9
B-20
155 pm
2
B-8
R
ψ
ψ
2
ϕ
-180
-120
-60
0
60
120
180
1.
ϕ
1
3
1
5.
Pleated sheet (antiparallel)
βa
α Helix (right-handed)
3.
αr
Pleated sheet (parallel)
βp
α Helix (left-handed)
C
Collagen helix
αl
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
68
Biomolecules
B. Collagen helix
Secondary structures
Another type of helix occurs in the collagens,
In proteins, specific combinations of the dihe-
which are important constituents of the con-
dral angles φ and ψ (see p. 66) are much more
nectivetissue matrix (see pp. 70, 344). The
common than others. When several succes-
collagen helix is left-handed, and with a pitch
sive residues adopt one of these conforma-
of 0.96 nm and 3.3 residues per turn, it is
tions, defined secondary structures arise,
steeper than the α-helix. In contrast to the
which are stabilized by hydrogen bonds ei-
α-helix, H bonds are not possible within the
ther within the peptide chain or between
collagen helix. However, the conformation is
neighboring chains. When a large part of a
stabilized by the association of three helices
protein takes on a defined secondary struc-
to form a righthanded collagen triple helix
ture, the protein often forms mechanically
(see p. 70).
stable filaments or fibers. Structural proteins
of this type (see p. 70) usually have character-
istic amino acid compositions.
C. Pleated-sheet structures
The most important secondary structural
Two additional, almost stretched, conforma-
elements of proteins are discussed here first.
tions of the peptide chain are known as E
The illustrations only show the course of the
pleated sheets, as the peptide planes are ar-
peptide chain; the side chains are omitted. To
ranged like a regularly folded sheet of paper.
make the course of the chains clearer, the
Again, H bonds can only form between neigh-
levels of the peptide bonds are shown as
boring chains (“strands”) in pleated sheets.
blue planes. The dihedral angles of the struc-
When the two strands run in opposite direc-
tures shown here are also marked in diagram
tions (1), the structure is referred to as an
D1 on p. 67.
antiparallel pleated sheet (βa). When they
run in the same direction (2), it is a parallel
A. D-Helix
pleated sheet (βp). In both cases, the α-C
atoms occupy the highest and lowest points
The right-handed α-helix (αR) is one of the
in the structure, and the side chains point
most common secondary structures. In this
alternately straight up or straight down (see
conformation, the peptide chain is wound
p. 71 C). The βa structure, with its almost lin-
like a screw. Each turn of the screw (the screw
ear H bonds, is energetically more favorable.
axis in shown in orange) covers approxi-
In extended pleated sheets, the individual
mately 3.6 amino acid residues. The pitch of
strands of the sheet are usually not parallel,
the screw (i. e., the smallest distance between
but twisted relative to one another (see p. 74).
two equivalent points) is 0.54 nm. α-Helices
are stabilized by almost linear hydrogen bonds
between the NH and CO groups of residues,
D. E Turns
which are four positions apart from each an-
E Turns are often found at sites where the
other in the sequence (indicated by red dots;
peptide chain changes direction. These are
see p. 6). In longer helices, most amino acid
sections in which four amino acid residues
residues thus enter into two H bonds. Apolar
are arranged in such a way that the course
or amphipathic α-helices with five to seven
of the chain reverses by about 180° into the
turns often serve to anchor proteins in bio-
opposite direction. The two turns shown
logical membranes (transmembrane helices;
(types I and II) are particularly frequent.
see p. 214).
Both are stabilized by hydrogen bonds be-
The mirror image of the αR helix, the left-
tween residues 1 and 4. β Turns are often
handed D-helix (αL), is rarely found in nature,
located between the individual strands of
although it would be energetically “permissi-
antiparallel pleated sheets, or between
ble.”
strands of pleated sheets and α helices.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Peptides and Proteins
69
A. α Helix
B. Collagen helix
C
C
N
N
ϕ = -57˚
-80˚ < ϕ < -50˚
ψ = -47˚
+130˚ < ψ < +155˚
C. Pleated-sheet structures
ϕ = -139˚
ϕ = -119˚
1. Antiparallel
ψ = +135˚
2. Parallel
ψ = +113˚
D. β Turns
1. Type I
N
C
2. Type II
N
C
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
70
Biomolecules
two latter amino acids are only formed during
Structural proteins
collagen biosynthesis as a result of posttrans-
The structural proteins give extracellular
lational modification (see p. 344).
structures mechanical stability, and are in-
The triplet Gly-X-Y (2) is constantly re-
volved in the structure of the cytoskeleton
peated in the sequence of collagen, with the
(see p. 204). Most of these proteins contain a
X position often being occupied by Pro and
high percentage of specific secondary struc-
the Y position by Hyp. The reason for this is
tures (see p. 68). For this reason, the amino
that collagen is largely present as a triple helix
acid composition of many structural proteins
made up of three individual collagen helices
is also characteristic (see below).
(1). In triple helices, every third residue lies
on the inside of the molecule, where for steric
reasons there is only room for glycine resi-
A. D Keratin
dues (3; the glycine residues are shown in
D-Keratin is a structural protein that predom-
yellow). Only a small section of a triple helix
inantly consists of α helices. Hair
(wool),
is illustrated here. The complete collagen
feathers, nails, claws and the hooves of ani-
molecule is approximately 300 nm long.
mals consist largely of keratin. It is also an
important component of the cytoskeleton
C. Silk fibroin
(cytokeratin), where it appears in intermedi-
ate filaments (see p. 204).
Silk is produced from the spun threads from
In the keratins, large parts of the peptide
silkworms (the larvae of the moth Bombyx
chain show right-handed α-helical coiling.
mori and related species). The main protein
Two chains each form a left-handed super-
in silk, fibroin, consists of antiparallel pleated
helix, as is also seen in myosin (see p. 65).
sheet structures arranged one on top of the
The superhelical keratin dimers join to form
other in numerous layers (1). Since the amino
tetramers, and these aggregate further to
acid side chains in pleated sheets point either
form protofilaments, with a diameter of
straight up or straight down (see p. 68), only
3 nm. Finally, eight protofilaments then
compact side chains fit between the layers. In
form an intermediate filament, with a diam-
fact, more than 80% of fibroin consists of gly-
eter of 10 nm (see p. 204).
cine, alanine, and serine, the three amino
Similar keratin filaments are found in hair.
acids with the shortest side chains. A typical
In a single wool fiber with a diameter of about
repetitive amino acid sequence is (Gly-Ala-
20 µm, millions of filaments are bundled to-
Gly-Ala-Gly-Ser). The individual pleated sheet
gether within dead cells. The individual kera-
layers in fibroin are found to lie alternately
tin helices are cross-linked and stabilized by
0.35 nm and 0.57 nm apart. In the first case,
numerous disulfide bonds (see p. 72). This fact
only glycine residues (R = H) are opposed to
is exploited in the perming of hair. Initially,
one another. The slightly greater distance of
the disulfide bonds of hair keratin are dis-
0.57 nm results from repulsion forces be-
rupted by reduction with thiol compounds
tween the side chains of alanine and serine
(see p. 8). The hair is then styled in the desired
residues (2).
shape and heat-dried. In the process, new
disulfide bonds are formed by oxidation,
which maintain the hairstyle for some time.
B. Collagen
Collagen is the quantitatively most important
protein in mammals, making up about 25% of
the total protein. There are many different
types of collagen, particularly in connective
tissue. Collagen has an unusual amino acid
composition. Approximately one-third of the
amino acids are glycine (Gly), about 10 % pro-
line (Pro), and 10% hydroxyproline (Hyp). The
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Peptides and Proteins
71
A. α-Keratin
B. Collagen
Right-handed
α helix
Left-handed
3 nm
superhelix
10 nm
Intermediary
filament
Protofilament
C. Silk fibroin
1. Triple helix (section)
1. Spatial illustration
Gly
Arg
Hyp
Gly
Gln
Arg
Gly
Pro
Hyp
Gly
Gly
Gly
Pro
Gln
Ala
Ala
Gly
Ala
Arg
Gly
X
Y
Ala
Ser
2. Typical sequence
Gly
Gly
Gly
Gly
Gly
Ala
Ala
3. Triple helix
2. Front view
(view from above)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
72
Biomolecules
tively cleaved again. The small plant protein
Globular proteins
crambin (46 amino acids) contains three di-
Soluble proteins have a more complex struc-
sulfide bonds and is therefore very stable. The
ture than the fibrous, completely insoluble
high degree of stability of insulin (see p. 76)
structural proteins. The shape of soluble pro-
has a similar reason.
teins is more or less spherical (globular). In
their biologically active form, globular
C. Protein dynamics
proteins have a defined spatial structure
(the native conformation). If this structure is
The conformations of globular proteins are
destroyed (denaturation; see p. 74), not only
not rigid, but can change dramatically on
does the biological effect disappear, but the
binding of ligands or in contact with other
protein also usually precipitates in insoluble
proteins. For example, the enzyme adenylate
form. This happens, for example, when eggs
kinase (see p. 336) has a mobile domain (do-
are boiled; the proteins dissolved in the egg
main = independently folded partial struc-
white are denatured by the heat and produce
ture), which folds shut after binding of the
the solid egg white.
substrate (yellow). The larger domain (bot-
To illustrate protein conformations in a
tom) also markedly alters its conformation.
clear (but extremely simplified) way, Richard-
There are large numbers of allosteric
son diagrams are often used. In these
proteins of this type. This group includes, for
diagrams, α-helices are symbolized by red
example, hemoglobin (see p. 280), calmodulin
cylinders or spirals and strands of pleated
(see p. 386), and many allosteric enzymes
sheets by green arrows. Less structured areas
such as aspartate carbamoyltransferase (see
of the chain, including the β-turns, are shown
p. 116).
as sections of gray tubing.
D. Folding patterns
A. Conformation-stabilizing interactions
The globular proteins show a high degree of
The native conformation of proteins is stabi-
variability in folding of their peptide chains.
lized by a number of different interactions.
Only a few examples are shown here. Purely
Among these, only the disulfide bonds (B)
helically folded proteins such as myoglobin (1;
represent covalent bonds. Hydrogen bonds,
see p. 74, heme yellow) are rare. In general,
which can form inside secondary structures,
pleated sheet and helical elements exist
as well as between more distant residues, are
alongside each other. In the hormone-binding
involved in all proteins (see p. 6). Many pro-
domain of the estrogen receptor (2; see p. 378),
teins are also stabilized by complex formation
a small, two-stranded pleated sheet functions
with metal ions (see pp. 76, 342, and 378, for
as a “cover” for the hormone binding site
example). The hydrophobic effect is particu-
(estradiol yellow). In flavodoxin, a small flavo-
larly important for protein stability. In glob-
protein with a redox function (3; FMN yel-
ular proteins, most hydrophobic amino acid
low), a fan-shaped, pleated sheet made up of
residues are arranged in the interior of the
five parallel strands forms the core of the
structure in the native conformation, while
molecule. The conformation of the β subunit
the polar amino acids are mainly found on
of the G-protein transducin (4; see pp. 224,
the surface (see pp. 28, 76).
358) is very unusual. Seven pleated sheets
form a large, symmetrical “β propeller.” The
N-terminal section of the protein contains one
B. Disulfide bonds
long and one short helix.
Disulfide bonds arise when the SH groups of
two cysteine residues are covalently linked as
a dithiol by oxidation. Bonds of this type are
only found (with a few exceptions) in extra-
cellular proteins, because in the interior of the
cell glutathione (see p. 284) and other reduc-
ing compounds are present in such high con-
centrations that disulfides would be reduc-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Peptides and Proteins
73
A. Conformation-stabilizing interactions
D. Folding patterns
Polar surface
Disulfide
bond
Apolar core
1. Myoglobin
Metal
complex
Hydrogen bond
B. Disulfide bonds
2. Estrogen receptor (domain)
H
N
S
S
N H
HC
CH2
CH2
CH
O
C
C O
3. Flavodoxin
C. Protein dynamics
Mobile domain
S
Substrate
Adenylate kinase
4. Transducin (β subunit)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
74
Biomolecules
When the urea and thiol are removed by
Protein folding
dialysis
(see p. 78), secondary and tertiary
Information about the biologically active (na-
structures develop again spontaneously. The
tive) conformation of proteins is already en-
cysteine residues thus return to a suf ciently
coded in their amino acid sequences. The na-
close spatial vicinity that disulfide bonds can
tive forms of many proteins arise spontane-
once again form under the oxidative effect of
ously in the test tube and within a few mi-
atmospheric oxygen. The active center also
nutes. Nevertheless, there are special
reestablishes itself. In comparison with the
auxiliary proteins (chaperonines) that sup-
denatured protein, the native form is aston-
port the folding of other proteins in the con-
ishingly compact, at 4.5
2.5 nm. In this state,
ditions present within the cell (see p. 232). An
the apolar side chains (yellow) predominate
important goal of biochemistry is to under-
in the interior of the protein, while the polar
stand the laws governing protein folding. This
residues are mainly found on the surface. This
would make it possible to predict the confor-
distribution is due to the “hydrophobic effect”
mation of a protein from the easily accessible
(see p. 28), and it makes a vital contribution to
DNA sequence (see p. 260).
the stability of the native conformation (B).
A. Folding and denaturation of
B. Energetics of protein folding
ribonuclease A
The energetics of protein folding are not at
The folding of proteins to the native form is
present satisfactorily understood. Only a sim-
favored under physiological conditions. The
plified model is discussed here. The confor-
native conformation is lost, as the result of
mation of a molecule is stable in any given
denaturation, at extreme pH values, at high
conditions if the change in its free enthalpy
temperatures, and in the presence of organic
during folding (∆Gfold) is negative (see p. 16).
solvents, detergents, and other denaturing
The magnitude of the folding enthalpy is af-
substances, such as urea.
fected by several factors. The main factor
The fact that a denatured protein can spon-
working against folding is the strong increase
taneously return to its native conformation
in the ordering of the molecule involved. As
was demonstrated for the first time with ri-
discussed on p. 20, this leads to a negative
bonuclease, a digestive enzyme (see p. 266)
change in entropy of ∆Sconf and therefore to
consisting of 124 amino acids. In the native
a strongly positive entropy term -T
∆S (violet
form (top right), there are extensive pleated
arrow). By contrast, the covalent and nonco-
sheet structures and three α helices. The eight
valent bonds in the interior of the protein
cysteine residues of the protein are forming
have a stabilizing influence. For this reason,
four disulfide bonds. Residues His-12, Lys-41
the change in folding enthalpy ∆Hfold is neg-
and His-119 (pink) are particularly important
ative (red arrow). A third factor is the change
for catalysis. Together with additional amino
in the system’s entropy due to the hydropho-
acids, they form the enzyme’s active center.
bic effect. During folding, the degree of order
The disulfide bonds can be reductively
in the surrounding water decreases—i. e.,
cleaved by thiols
(e. g., mercaptoethanol,
∆Swater is positive and therefore
-T
∆S is
HO-CH2-CH2-SH). If urea at a high concentra-
negative (blue arrow). When the sum of these
tion is also added, the protein unfolds com-
effects is negative (green arrow), the protein
pletely. In this form (left), it is up to 35 nm
folds spontaneously into its native conforma-
long. Polar (green) and apolar (yellow) side
tion.
chains are distributed randomly. The dena-
tured enzyme is completely inactive, because
the catalytically important amino acids (pink)
are too far away from each other to be able to
interact with each other and with the sub-
strate.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Peptides and Proteins
75
A. Folding and denaturation of ribonuclease A
N terminus
N terminus
58-110
26-84
His -12
40-95
26
His -12
65-72
Lys-41
His-119
C terminus
40
Lys-41
Folding
Denaturation
58
65
B. Energetics of protein folding
+
72
Increased order
in the molecule
Interactions
in the
molecule
Reduced
order
in water
84
95
Free enthalpy
of folding
-
(∆Gfold)
110
His-119
C terminus
∆Gfold = ∆Hfold -T·∆Swater -T·∆Sconf
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
76
Biomolecules
as the tertiary structure. In insulin, it is com-
Molecular models: insulin
pact and wedge-shaped (B). The tip of the
The opposite page presents models of insulin,
wedge is formed by the B chain, which
a small protein. The biosynthesis and function
changes its direction at this point.
of this important hormone are discussed else-
Quaternary structure. Due to non-covalent
where in this book (pp. 160, 388). Monomeric
interactions, many proteins assemble to form
insulin consists of 51 amino acids, and with a
symmetrical complexes (oligomers). The indi-
molecular mass of 5.5 kDa it is only half the
vidual components of oligomeric proteins
size of the smallest enzymes. Nevertheless, it
(usually 2-12) are termed subunits or mono-
has the typical properties of a globular pro-
mers. Insulin also forms quaternary struc-
tein.
tures. In the blood, it is partly present as a
Large quantities of pure insulin are re-
dimer. In addition, there are also hexamers
quired for the treatment of diabetes mellitus
stabilized by Zn2+ ions (light blue) (3), which
(see p. 160). The annual requirement for insu-
represent the form in which insulin is stored
lin is over 500 kg in a country the size of
in the pancreas (see p. 160).
Germany. Formerly, the hormone had to be
obtained from the pancreas of slaughtered
B. Insulin (monomer)
animals in a complicated and expensive
procedure. Human insulin, which is produced
The van der Waals model of monomeric in-
by overexpression in genetically engineered
sulin (1) once again shows the wedge-shaped
bacteria, is now mainly used (see p. 262).
tertiary structure formed by the two chains
together. In the second model (3, bottom), the
side chains of polar amino acids are shown in
A. Structure of insulin
blue, while apolar residues are yellow or pink.
There are various different structural levels in
This model emphasizes the importance of the
proteins, and these can be briefly discussed
“hydrophobic effect” for protein folding (see
again here using the example of insulin.
p. 74). In insulin as well, most hydrophobic
The primary structure of a protein is its
side chains are located on the inside of the
amino acid sequence. During the biosynthesis
molecule, while the hydrophilic residues are
of insulin in the pancreas, a continuous pep-
located on the surface. Apparently in contra-
tide chain with 84 residues is first synthesi-
diction to this rule, several apolar side chains
zed—proinsulin (see p. 160). After folding of
(pink) are found on the surface. However, all
the molecule, the three disulfide bonds are
of these residues are involved in hydrophobic
first formed, and residues 31 to 63 are then
interactions that stabilize the dimeric and
proteolytically cleaved releasing the so-called
hexameric forms of insulin.
C peptide. The molecule that is left over (1)
In the third model (2, right), the colored
now consists of two peptide chains, the A
residues are those that are located on the
chain (21 residues, shown in yellow) and the
surface and occur invariably (red) or almost
B chain (30 residues, orange). One of the di-
invariably (orange) in all known insulins. It is
sulfide bonds is located inside the A chain,
assumed that amino acid residues that are not
and the two others link the two chains to-
replaced by other residues during the course
gether.
of evolution are essential for the protein’s
Secondary structures are regions of the
function. In the case of insulin, almost all of
peptide chain with a defined conformation
these residues are located on one side of the
(see p. 68) that are stabilized by H-bonds. In
molecule. They are probably involved in the
insulin (2), the α-helical areas are predomi-
binding of the hormone to its receptor (see
nant, making up 57% of the molecule; 6%
p. 224).
consists of β-pleated-sheet structures, and
10% of β-turns, while the remainder (27%)
cannot be assigned to any of the secondary
structures.
The three-dimensional conformation of a
protein, made up of secondary structural ele-
ments and unordered sections, is referred to
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Peptides and Proteins
77
A. Structure of insulin
C Peptide
OOC
30
29
28
27
26
25
24
23
22
E R R
A K P T Y F F G
R
B chain
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
E
F
V
N
Q
H
L
C
G
S
H L
V E
A L
Y L
V
C
G
21
NH
3
1
2
3
4
5
6
12
13
14
15
16
17
18
19
20
21
Q
7
11
K R
G
I
V
E
Q
C
S L
Y
Q L
E
N
Y C
N
A chain
C
C
NH3
OOC
T S I
1. Primary structure
Disulfide bonds
8
9
10
A chain
NH3
OOC
NH3
C peptide
OOC
B chain
2. Secondary and tertiary structure
3. Quaternary structure
B. Insulin (monomer)
A-chain
B-chain
Invariant
residue
1.
2.
Apolar
side chain
Involved
in subunit
interactions
Polar
side chain
3.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
78
Biomolecules
spherical gel particles (diameter 10-500 µm)
Isolation and analysis of proteins
of polymeric material (shown schematically
Purified proteins are nowadays required for a
in 1a). The insides of the particles are tra-
wide variety of applications in research, med-
versed by channels that have defined diame-
icine, and biotechnology. Since the globular
ters. A protein mixture is then introduced at
proteins in particular are very unstable (see
the upper end of the column (1b) and elution
p. 72), purification is carried out at low tem-
is carried out by passing a buffer solution
peratures
(0-5 °C) and particularly gentle
through the column. Large protein molecules
separation processes are used. A few of the
(red) are unable to penetrate the particles,
methods of purifying and characterizing pro-
and therefore pass through the column
teins are discussed on this page.
quickly. Medium-sized (green) and small par-
ticles (blue) are delayed for longer or shorter
periods (1c). The proteins can be collected
A. Salt precipitation
separately from the ef uent (eluate) (2). Their
The solubility of proteins is strongly depen-
elution volume Ve depends mainly on their
dent on the salt concentration (ionic strength)
molecular mass (3).
of the medium. Proteins are usually poorly
soluble in pure water. Their solubility in-
D. SDS gel electrophoresis
creases as the ionic strength increases, be-
cause more and more of the well-hydrated
The most commonly used procedure for
anorganic ions (blue circles) are bound to
checking the purity of proteins is sodium do-
the protein’s surface, preventing aggregation
decyl sulfate polyacrylamide gel electropho-
of the molecules (salting in). At very high
resis
(SDS-PAGE). In electrophoresis, mole-
ionic strengths, the salt withdraws the hy-
cules move in an electrical field (see p. 276).
drate water from the proteins and thus leads
Normally, the speed of their movement de-
to aggregation and precipitation of the mole-
pends on three factors—their size, their shape,
cules (salting out). For this reason, adding
and their electrical charge.
salts such as ammonium sulfate (NH4)2SO4
In SDS-PAGE, the protein mixture is treated
makes it possible to separate proteins from a
in such a way that only the molecules’ mass
mixture according to their degree of solubility
affects their movement. This is achieved by
(fractionation).
adding sodium dodecyl sulfate
(C12H25-
OSO3Na), the sulfuric acid ester of lauryl alco-
hol (dodecyl alcohol). SDS is a detergent with
B. Dialysis
strongly amphipathic properties (see p. 28). It
Dialysis is used to remove lower-molecular
separates oligomeric proteins into their sub-
components from protein solutions, or to ex-
units and denatures them. SDS molecules
change the medium. Dialysis is based on the
bind to the unfolded peptide chains
(ca.
fact that due to their size, protein molecules
0.4 g SDS / g protein) and give them a strongly
are unable to pass through the pores of a
negative charge. To achieve complete denatu-
semipermeable membrane, while lower-mo-
ration, thiols are also added in order to cleave
lecular substances distribute themselves
the disulfide bonds (1).
evenly between the inner and outer spaces
Following electrophoresis, which is carried
over time. After repeated exchanging of the
out in a vertically arranged gel of polymeric
external solution, the conditions inside the
acrylamide (2), the separated proteins are
dialysis tube
(salt concentration, pH, etc.)
made visible by staining. In example (3), the
will be the same as in the surrounding solu-
following were separated: a) a cell extract
tion.
with hundreds of different proteins, b) a pro-
tein purified from this, and c) a mixture of
proteins with known masses.
C. Gel filtration
Gel permeation chromatography (“gel filtra-
tion”) separates proteins according to their
size and shape. This is done using a chroma-
tography column, which is filled with
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Peptides and Proteins
79
A. Salt precipitation
B. Dialysis
Solubility
Ion
Protein
Protein
Dialysis
solution
tube
Hydrate
shell
Salting in
Buffer
Salting out
Stirrer
solution
Salt concentration
C. Gel filtration
D. SDS gel electrophoresis
Gel particle
SDS
(cross-section)
R - SH Thiol
Channel
S
Pump
S
Denatured
Folded
1. Principle
Cathode
Probe
Plastic
Elution
casing
buffer
Upper buffer
Gel
reservoir
particle
Polyacrylamide
Column
gel
Anode
a
b
c
Lower buffer
reservoir
1. Principle
2. Apparatus
Ve (mL)
22
Exclusion
volume
20
18
16
14
12
8
12
16
20
24
10
20
40 6080
a
b c
20
40 6080
Eluted volume (mL)
Mass (kDa)
Mass (kDa)
2. Elution diagram
3. Analysis
3. Stained gel
4. Analysis
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
80
Biomolecules
residues, it yields nucleoside diphosphates
Bases and nucleotides
and triphosphates—e. g., ADP and ATP, which
The nucleic acids play a central role in the
are important coenzymes in energy metabo-
storage and expression of genetic information
lism (see p. 106). All of these nucleoside phos-
(see p. 236). They are divided into two major
phates are classified as nucleotides.
classes: deoxyribonucleic acid
(DNA) func-
In nucleosides and nucleotides, the pentose
tions solely in information storage, while ri-
residues are present in the furanose form (see
bonucleic acids (RNAs) are involved in most
p. 34). The sugars and bases are linked by an
steps of gene expression and protein biosyn-
N-glycosidic bond between the C-1 of the
thesis. All nucleic acids are made up from
sugar and either the N-9 of the purine ring
nucleotide components, which in turn consist
or N-1 of the pyrimidine ring. This bond al-
of a base, a sugar, and a phosphate residue.
ways adopts the β-configuration.
DNA and RNA differ from one another in the
type of the sugar and in one of the bases that
C. Oligonucleotides, polynucleotides
they contain.
Phosphoric acid molecules can form acid-an-
hydride bonds with each other. It is therefore
A. Nucleic acid bases
possible for two nucleotides to be linked via
The bases that occur in nucleic acids are
the phosphate residues. This gives rise to di-
aromatic heterocyclic compounds derived
nucleotides with a phosphoric acid-anhydride
from either pyrimidine or purine. Five of these
structure. This group includes the coenzymes
bases are the main components of nucleic
NAD(P)+ and CoA, as well as the flavin
acids in all living creatures. The purine bases
derivative FAD (1; see p. 104).
adenine (abbreviation Ade, not “A”!) and gua-
If the phosphate residue of a nucleotide
nine (Gua) and the pyrimidine base cytosine
reacts with the 3 -OH group of a second nu-
(Cyt) are present in both RNA and DNA. In
cleotide, the result is a dinucleotide with a
contrast, uracil (Ura) is only found in RNA. In
phosphoric acid diester structure. Dinucleo-
DNA, uracil is replaced by thymine (Thy), the
tides of this type have a free phosphate resi-
5-methyl derivative of uracil. 5-methylcyto-
due at the 5 end and a free OH group at the 3
sine also occurs in small amounts in the DNA
end. They can therefore be extended with
of the higher animals. A large number of other
additional mononucleotides by adding fur-
modified bases occur in tRNA (see p. 82) and
ther phosphoric acid diester bonds. This is
in other types of RNA.
the way in which oligonucleotides, and ulti-
mately polynucleotides, are synthesized.
Polynucleotides consisting of ribonucleo-
B. Nucleosides, nucleotides
tide components are called ribonucleic acid
When a nucleic acid base is N-glycosidically
(RNA), while those consisting of deoxyribonu-
linked to ribose or 2-deoxyribose (see p. 38), it
cleotide monomers are called deoxyribonu-
yields a nucleoside. The nucleoside adenosine
cleic acid (DNA; see p. 84). To describe the
(abbreviation: A) is formed in this way from
structure of polynucleotides, the abbrevia-
adenine and ribose, for example. The corre-
tions for the nucleoside components are writ-
sponding derivatives of the other bases are
ten from left to right in the 5
3
direction.
called guanosine (G), uridine (U), thymidine
The position of the phosphate residue is also
(T) and cytidine (C). When the sugar compo-
sometimes indicated by a “p”. In this way, the
nent is
2-deoxyribose, the product is a
structure of the RNA segment shown Fig. 2
deoxyribonucleoside—e. g.,
2 -deoxyadeno-
can be abbreviated as ..pUpG.. or simply as
sine (dA, not shown). In the cell, the 5 OH
..UG .. .
group of the sugar component of the nucleo-
side is usually esterified with phosphoric acid.
2 -Deoxythymidine (dT) therefore gives rise
to
2 -deoxythymidine-5 -monophosphate
(dTMP), one of the components of DNA (2).
If the 5 phosphate residue is linked via an
acid-anhydride bond to additional phosphate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nucleotides and Nucleic Acids
81
A. Nucleic acid bases
O
O
Pyrimidine bases
NH
2
H
C
C
C
CH3
C
N
3
4
5
CH
HN
CH
HN
C
N
CH
2
6
HC
1
CH
C
CH
C
CH
C
CH
O
N
N
O
N
O
N
H
H
H
Pyrimidine
Uracil (Ura)
Thymine (Thy)
Cytosine (Cyt)
Purine bases
H
NH
O
2
C
N
6
N
1
5
C
7
C
C
N
N
8
CH
N
C
HN
C
2
4
9
HC
3
C
CH
CH
N
N
HC
C
C
C
H
N
N
N
N
H2N
H
H
Purine
Adenine (Ade)
Guanine (Gua)
B. Nucleosides, nucleotides
O
NH2
T
P
CH3
C
C
N
HN
C
C
N
HC
C
CH
5'
CH
O
N
C
O
N
HO
CH2
N
5'
O
P
O
CH2
O
β
β
1'
O
H H
O
1'
H H
H
H
H
H
OH OH
OH H
1. Adenosine (Ado)
2. 2'-Deoxythymidine 5'-monophosphate (dtMP)
C. Oligonucleotides, polynucleotides
O
Flavin
To the 5' end
C
H
O
F
A
HC
NH
U
O
H3C
C
N
C
H
HC
C
C
C
C
N
Phosphoric acid-
N
O
O
P
O
CH2
anhydride bond
C
C
C
C
O
O
O
H3C
C
N
N
O
H
H
H
CH2
H
H
C
H
H
N
C
NH
FH
C
OH
A
N
O
OH
HC
G
C
C
H
C
OH
C
N
N
N
O
CH2
O
P
NH2
Ribitol
C
N
O
H
C
OH
O
O
H
C
O
C
C
H H
N
H2C
O
P
O
P
O
CH2
N
H
H
H
O
O
O
Phosphoric acid
O OH
H H
Ribose
diester bond
H
H
To the 3' end
1. Flavin adenine dinucleotide
OH OH
(FAD)
2. RNA (section)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
82
Biomolecules
Messenger RNAs (mRNAs) transfer genetic
RNA
information from the cell nucleus to the cyto-
Ribonucleic acids (RNAs) are polymers con-
plasm. The primary transcripts are substan-
sisting of nucleoside phosphate components
tially modified while still in the nucleus
that are linked by phosphoric acid diester
(mRNA maturation; see p. 246). Since mRNAs
bonds (see p. 80). The bases the contain are
have to be read codon by codon in the ribo-
mainly uracil, cytosine, adenine, and guanine,
some, they must not form a stable tertiary
but many unusual and modified bases are also
structure. This is ensured in part by the at-
found in RNAs (B).
tachment of RNA-binding proteins, which pre-
vent base pairing. Due to the varying amounts
of information that they carry, the lengths of
A. Ribonucleic acids (RNAs)
mRNAs also vary widely. Their lifespan is usu-
RNAs are involved in all the individual steps of
ally short, as they are quickly broken down
gene expression and protein biosynthesis (see
after translation.
pp. 242-253). The properties of the most im-
Small nuclear RNAs (snRNAs) are involved
portant forms of RNA are summarized in the
in the splicing of mRNA precursors
(see
table. The schematic diagram also gives an
p. 246). They associate with numerous pro-
idea of the secondary structure of these mol-
teins to form “spliceosomes.”
ecules.
In contrast to DNA, RNAs do not form ex-
B. Transfer RNA (tRNAPhe)
tended double helices. In RNAs, the base pairs
(see p. 84) usually only extend over a few
The transfer RNAs (tRNAs) function during
residues. For this reason, substructures often
translation (see p. 250) as links between the
arise that have a finger shape or clover-leaf
nucleic acids and proteins. They are small
shape in two-dimensional representations. In
RNA molecules consisting of 70-90 nucleoti-
these, the paired stem regions are linked by
des, which “recognize” specific mRNA codons
loops. Large RNAs such as ribosomal 16S-
with their anticodons through base pairing. At
rRNA (center) contain numerous “stem and
the same time, at their 3
end (sequence
loop” regions of this type. These sections are
.. CCA-3 ) they carry the amino acid that is
again folded three-dimensionally—i. e., like
assigned to the relevant mRNA codon accord-
proteins, RNAs have a tertiary structure (see
ing to the genetic code (see p. 248).
p. 86). However, tertiary structures are only
The base sequence and the tertiary struc-
known of small RNAs, mainly tRNAs. The dia-
ture of the yeast tRNA specific for phenylala-
grams in Fig. B and on p. 86 show that the
nine (tRNAPhe) is typical of all tRNAs. The
“clover-leaf” structure is not recognizable in
molecule (see also p. 86) contains a high pro-
a three-dimensional representation.
portion of unusual and modified components
Cellular RNAs vary widely in their size,
(shaded in dark green in Fig. 1). These include
structure, and lifespan. The great majority of
pseudouridine (Ψ), dihydrouridine (D), thymi-
them are ribosomal RNA (rRNA), which in
dine (T), which otherwise only occurs in DNA,
several forms is a structural and functional
and many methylated nucleotides such as 7-
component of ribosomes (see p. 250). Riboso-
methylguanidine
(m7G) and—in the anti-
mal RNA is produced from DNA by transcrip-
codon—2 -O-methylguanidine (m2G). Numer-
tion in the nucleolus, and it is processed there
ous base pairs, sometimes deviating from the
and assembled with proteins to form ribo-
usual pattern, stabilize the molecule’s confor-
some subunits (see pp. 208, 242). The bacte-
mation (2).
rial 16S-rRNA shown in Fig. A, with 1542 nu-
cleotides (nt), is a component of the small
ribosomae subunit, while the much smaller
5S-rRNA (118 nt) is located in the large sub-
unit.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nucleic Acids
83
A. Ribonucleic acids (RNAs)
5'
3'
tRNA
mRNA U1-snRNA
5'
5'
3'
5'
3'
5'
3'
3'
16S-RNA
5S-rRNA
tRNA
rRNA
Type
mRNA
snRNA
>50
4
Species per cell
> 1000
~ 10
74 - 95
120 - 5000
Length (b)
400 - 6000
100 - 300
10-20%
80%
Proportion
5%
< 1%
Long
Long
Lifespan
Short
Long
Translation
Translation
Function
Translation
Splicing
B. Transfer RNA (tRNAPhe)
O
O
5'
Phe
TΨ loop
C
C
H2C
NH
HN
NH
3'
H2C
C
HC
C
N
5'
C
O
O
D loop
O
CH2
O
CH2
O
O
H H
H H
H
H
H
H
Variable
loop
O
OH
3'
O OH
A
Phe
C
C
mRNA
Dihydrouridine (D)
5'
Pseudouridine (Ψ)
A
G
C
Anticodon
C
G
Codon
G
C
G
U
TΨ loop
Normal base
A
U
pairing
U
A
C U
*
2. Conformation
U
A
A
Unusual base
G A C AC
AU
G
pairing
DD
GA
C U G U
G
CUCG
C
C
G
T
U
Y
G AG C
*
G
G
G
A
G
G
*
C
GA
*
D loop
C
G
2'-O-methylguanidine (m
2G)
7-methylguanidine (m7G)
A
U
G
C
Variable
O
O
*
CH3
A
Y
loop
C
A
C
C
*
N
N
U
Y
C
NH
C
NH
G
HC
HC
A A
C
C
C
C
Anticodon
N
N
O
CH2
N
NH2
O
CH2
N
NH
*
O
O
1'
H H
H H
* Methylated base
H
H
H
H
3'
O OCH3
O OH
1. Structure
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
84
Biomolecules
sugar and phosphate residues in the back-
DNA
bone. Along the whole length of the DNA
molecule, there are two depressions—re-
A. DNA: structure
ferred to as the “minor groove” and the “ma-
Like RNAs (see p. 82), deoxyribonucleic acids
jor groove”—that lie between the strands.
(DNAs) are polymeric molecules consisting of
nucleotide building blocks. Instead of ribose,
B. Coding of genetic information
however, DNA contains 2 -deoxyribose, and
the uracil base in RNA is replaced by thymine.
In all living cells, DNA serves to store genetic
The spatial structure of the two molecules
information. Specific segments of DNA
also differs (see p. 86).
(“genes”) are transcribed as needed into
The first evidence of the special structure
RNAs, which either carry out structural or
of DNA was the observation that the amounts
catalytic tasks themselves or provide the basis
of adenine and thymine are almost equal in
for synthesizing proteins (see p. 82). In the
every type of DNA. The same applies to gua-
latter case, the DNA codes for the primary
nine and cytosine. The model of DNA struc-
structure of proteins. The “language” used in
ture formulated in 1953 explains these con-
this process has four letters (A, G, C, and T). All
stant base ratios: intact DNA consists of two
of the words (“codons”) contain three letters
polydeoxynucleotide molecules
(“strands”).
(“triplets”), and each triplet stands for one of
Each base in one strand is linked to a comple-
the 20 proteinogenic amino acids.
mentary base in the other strand by H-bonds.
The two strands of DNA are not function-
Adenine is complementary to thymine, and
ally equivalent. The template strand (the (-)
guanine is complementary to cytosine. One
strand or “codogenic strand,” shown in light
purine base and one pyrimidine base are
gray in Fig. 1) is the one that is read during the
thus involved in each base pair.
synthesis of RNA (transcription; see p. 242).
The complementarity of A with T and of G
Its sequence is complementary to the RNA
with C can be understood by considering the
formed. The sense strand (the (+) strand or
H bonds that are possible between the differ-
“coding strand,” shown in color in Figs. 1 and
ent bases. Potential donors
(see p. 6) are
2) has the same sequence as the RNA, except
amino groups (Ade, Cyt, Gua) and ring NH
that T is exchanged for U. By convention, it is
groups. Possible acceptors are carbonyl oxy-
agreed that gene sequences are expressed by
gen atoms (Thy, Cyt, Gua) and ring nitrogen
reading the sequence of the sense strand in
atoms. Two linear and therefore highly stable
the 5
3
direction. Using the genetic code
bonds can thus be formed in A-T pairs, and
(see p. 248), in this case the protein sequence
three in G-C pairs.
(3) is obtained directly in the reading direc-
Base pairings of this type are only possible,
tion usual for proteins—i. e., from the N termi-
however, when the polarity of the two strands
nus to the C terminus.
differs—i. e., when they run in opposite direc-
tions (see p. 80). In addition, the two strands
have to be intertwined to form a double helix.
Due to steric hindrance by the 2 -OH groups
of the ribose residues, RNA is unable to form a
double helix. The structure of RNA is therefore
less regular than that of DNA (see p. 82).
The conformation of DNA that predomi-
nates within the cell (known as B-DNA) is
shown schematically in Fig. A2 and as a van
der Waals model in Fig. B1. In the schematic
diagram
(A2), the deoxyribose-phosphate
“backbone” is shown as a ribbon. The bases
(indicated by lines) are located on the inside
of the double helix. This area of DNA is there-
fore apolar. By contrast, the molecule’s surface
is polar and negatively charged, due to the
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nucleic Acids
85
A. DNA: structure
5'
3'
H
H
N
C
H
C
O
O P O
H
O
C
N
C
O
N
C
H
C
C
H
N
C
N
G
CH2
C
N
O
C
G
O
C
H
N
O
A
T
N
H2C
H
O
H
O
O
P
O
CH3
N H
O
O
P
O
H
N
O
C
C C
C C
O
T
C H
N
A
N
H
N
C
CH2
C
N
O
N
C
O
O
H2 C
H
Minor
Major
O
OH
CH
OH
groove
groove
2
O
O
P
O
2'-deoxy-
D-ribose
5'
3'
2
OH
1. Formula
2. Double strand
B. Coding of genetic information
C
3'
5'
3'
3'
C
Codon
T
Phe
(triplet)
T
C
Sense strand
(+) strand,
A
Tyr
“coding strand”
T
T
C
Ala
G
T
C
Thr
A
A
Template
A
Glu
strand
G
(-) strand,
A
“codogenic
A
Glu
strand”
G
G
T
Met
A
5'
N
1.
2.
3.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
86
Biomolecules
A-DNA arises when B-DNA is dehydrated. It
Molecular models: DNA and RNA
probably does not occur in the cell.
The illustration opposite shows selected nuc-
In the Z-conformation (3), which can occur
leic acid molecules. Fig. A shows various con-
within GC-rich regions of B-DNA, the organ-
formations of DNA, and Fig. B shows the spa-
ization of the nucleotides is completely differ-
tial structures of two small RNA molecules. In
ent. In this case, the helix is left-handed, and
both, the van der Waals models (see p. 6) are
the backbone adopts a characteristic zig-zag
accompanied by ribbon diagrams that make
conformation (hence “Z-DNA”). The Z double
the course of the chains clear. In all of the
helix has a smaller pitch than B-DNA. DNA
models, the polynucleotide
“backbone” of
segments in the Z conformation probably
the molecule is shown in a darker color, while
have physiological significance, but details
the bases are lighter.
are not yet known.
A. DNA: conformation
B. RNA
Investigations of synthetic DNA molecules
RNA molecules are unable to form extended
have shown that DNA can adopt several dif-
double helices, and are therefore less highly
ferent conformations. All of the DNA seg-
ordered than DNA molecules. Nevertheless,
ments shown consist of 21 base pairs (bp)
they have defined secondary and tertiary
and have the same sequence.
structures, and a large proportion of the nu-
By far the most common form is B-DNA (2).
cleotide components enter into base pairings
As discussed on p. 84, this consists of two
with other nucleotides. The examples shown
antiparallel polydeoxynucleotide strands in-
here are 5S-rRNA (see p. 242), which occurs as
tertwined with one another to form a right-
a structural component in ribosomes, and a
handed double helix. The “backbone” of these
tRNA molecule from yeast (see p. 82) that is
strands is formed by deoxyribose and phos-
specific for phenylalanine.
phate residues linked by phosphoric acid di-
Both molecules are folded in such a way
ester bonds.
that the 3
end and the 5
end are close to-
In the B conformation, the aromatic rings of
gether. As in DNA, most of the bases are lo-
the nucleobases are stacked at a distance of
cated in the inside of the structures, while the
0.34 nm almost at right angles to the axis of
much more polar “backbone” is turned out-
the helix. Each base is rotated relative to the
wards. An exception to this is seen in the
preceding one by an angle of 35°. A complete
three bases of the anticodon of the tRNA
turn of the double helix (360°) therefore con-
(pink), which have to interact with mRNA
tains around 10 base pairs (abbreviation: bp),
and therefore lie on the surface of the mole-
i. e., the pitch of the helix is 3.4 nm. Between
cule. The bases of the conserved CCA triplet at
the backbones of the two individual strands
the 3
end (red) also jut outward. During
there are two grooves with different widths.
amino acid activation (see p. 248), they are
The major groove is visible at the top and
recognized and bound by the ligases.
bottom, while the narrower minor groove is
seen in the middle. DNA-binding proteins and
transcription factors (see pp. 118, 244) usually
enter into interactions in the area of the major
groove, with its more easily accessible bases.
In certain conditions, DNA can adopt the A
conformation (1). In this arrangement, the
double helix is still right-handed, but the
bases are no longer arranged at right angles
to the axis of the helix, as in the B form. As can
be seen, the A conformation is more compact
than the other two conformations. The minor
groove almost completely disappears, and the
major groove is narrower than in the B form.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nucleic Acids
87
A. DNA: conformation
3'
5'
3'
5'
3'
5'
5'
3'
5'
Backbone
5'
3'
3'
1. A - DNA
Bases
2. B - DNA
3. Z - DNA
B. RNA
3'
5'
5'
3'
1. 5S-rRNA
2. Phe-tRNAPhe
(118 nucleotides)
(77 nucleotides)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
88
Metabolism
Enzymes: basics
C. Enzyme classes
Enzymes are biological catalysts—i. e., sub-
More than 2000 different enzymes are cur-
stances of biological origin that accelerate
rently known. A system of classification has
chemical reactions (see p. 24). The orderly
been developed that takes into account both
course of metabolic processes is only possible
their reaction specificity and their substrate
because each cell is equipped with its own
specificity. Each enzyme is entered in the En-
genetically determined set of enzymes. It is
zyme Catalogue with a four-digit Enzyme
only this that allows coordinated sequences
Commission number (EC number). The first
of reactions (metabolic pathways; see p. 112).
digit indicates membership of one of the six
Enzymes are also involved in many regulatory
major classes. The next two indicate sub-
mechanisms that allow the metabolism to
classes and subsubclasses. The last digit indi-
adapt to changing conditions (see p. 114). Al-
cates where the enzyme belongs in the sub-
most all enzymes are proteins. However,
subclass. For example, lactate dehydrogenase
there are also catalytically active ribonucleic
(see pp. 98-101) has the EC number 1.1.1.27
acids, the “ribozymes” (see pp. 246, 252).
(class 1, oxidoreductases; subclass 1.1, CH-OH
group as electron donor; sub-subclass 1.1.1,
NAD(P)+ as electron acceptor).
A. Enzymatic activity
Enzymes with similar reaction specificities
The catalytic action of an enzyme, its activity,
are grouped into each of the six major classes:
is measured by determining the increase in
The oxidoreductases (class 1) catalyze the
the reaction rate under precisely defined con-
transfer of reducing equivalents from one re-
ditions—i. e., the difference between the turn-
dox system to another.
over (violet) of the catalyzed reaction (or-
The transferases
(class
2) catalyze the
ange) and uncatalyzed reaction (yellow) in a
transfer of other groups from one molecule
specific time interval. Normally, reaction rates
to another. Oxidoreductases and transferases
are expressed as the change in concentration
generally require coenzymes (see pp. 104ff.).
per unit of time (mol
1-1 s-1; see p. 22).
The hydrolases (class 3) are also involved in
Since the catalytic activity of an enzyme is
group transfer, but the acceptor is always a
independent of the volume, the unit used
water molecule.
for enzymes is usually turnover per unit time,
Lyases (class 4, often also referred to as
expressed in katal (kat, mol s-1). However,
“synthases”) catalyze reactions involving ei-
the international unit U is still more com-
ther the cleavage or formation of chemical
monly used (µmol turnover min-1; 1 U =
bonds, with double bonds either arising or
16.7 nkat).
disappearing.
The isomerases
(class
5) move groups
within a molecule, without changing the
B. Reaction and substrate specificity
gross composition of the substrate.
The action of enzymes is usually very specific.
The ligation reactions catalyzed by ligases
This applies not only to the type of reaction
(“synthetases,” class 6) are energy-dependent
being catalyzed (reaction specificity), but also
and are therefore always coupled to the hy-
to the nature of the reactants (“substrates”)
drolysis of nucleoside triphosphates.
that are involved (substrate specificity; see
p. 94). In Fig. B, this is illustrated schemati-
In addition to the enzyme name, we also
cally using a bond-breaking enzyme as an
usually give its EC number. The annotated
example. Highly specific enzymes (type A,
enzyme list (pp. 420ff.) includes all of the en-
top) catalyze the cleavage of only one type
zymes mentioned in this book, classified ac-
of bond, and only when the structure of the
cording to the Enzyme Catalog system.
substrate is the correct one. Other enzymes
(type B, middle) have narrow reaction specif-
icity, but broad substrate specificity. Type C
enzymes (with low reaction specificity and
low substrate specificity, bottom) are very
rare.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
89
A. Enzymatic activity
B. Reaction and substrate specificity
Group
Bond
A
Turnover (mol product. s-1)
without enzyme
Group
B
B
B
B
Turnover (mol product. s-1) with enzyme
Enzyme activity (mol. s-1 = kat)
C
C
C
C
C
C
C
C
Reaction specificity
Substrate specificity
1 Katal (kat): Amount of enzyme
A
High
High
which increases
-1
B
High
Low
turnover by 1 mol . s
C
Low
Low
C. The enzyme classes
Class
Reaction type
Important subclasses
= Reduction equivalent
Dehydrogenases
Oxidases, peroxidases
1 Oxidoreductases
+
Reductases
+
Monooxygenases
Dioxygenases
Ared
Box
Aox
Bred
C1-Transferases
Glycosyltransferases
2 Transferases
+
+
Aminotransferases
Phosphotransferases
A-B
C
A
B-C
Esterases
Glycosidases
3 Hydrolases
+
Peptidases
Amidases
A-B
H2O
A-H
B-OH
C-C-Lyases
C-O-Lyases
4 Lyases
+
+
C-N-Lyases
(“synthases”)
C-S-Lyases
A
B
A-B
Epimerases
cis trans Isomerases
5 Isomerases
Intramolecular
transferases
A
Iso-A
B
X=A,G,U,C
XDP
C-C-Ligases
X
X
C-O-Ligases
6 Ligases
+
P P P
+
P P
C-N-Ligases
(“synthetases”)
C-S-Ligases
P
A
XTP
A-B
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
90
Metabolism
that productive A-B complexes will arise. In
Enzyme catalysis
addition, binding of the substrates results in
Enzymes are extremely effective catalysts.
removal of their hydration shells. As a result
They can increase the rate of a catalyzed re-
of the exclusion of water, very different con-
action by a factor of 1012 or more. To grasp the
ditions apply in the active center of the en-
mechanisms involved in enzyme catalysis, we
zyme during catalysis than in solution (3-5).
can start by looking at the course of an un-
A third important factor is the stabilization of
catalyzed reaction more closely.
the transition state as a result of interactions
between the amino acid residues of the pro-
tein and the substrate (4). This further re-
A. Uncatalyzed reaction
duces the activation energy needed to create
The reaction A + B
C + D is used as an
the transition state. Many enzymes also take
example. In solution, reactants A and B are
up groups from the substrates or transfer
surrounded by a shell of water molecules
them to the substrates during catalysis.
(the hydration shell), and they move in ran-
Proton transfers are particularly common.
dom directions due to thermal agitation. They
This acid-base catalysis by enzymes is much
can only react with each other if they collide
more effective than the exchange of protons
in a favorable orientation. This is not very
between acids and bases in solution. In many
probable, and therefore only occurs rarely.
cases, chemical groups are temporarily bound
Before conversion into the products C + D,
covalently to the amino acid residues of the
the collision complex A-B has to pass through
enzyme or to coenzymes during the catalytic
a transition state, the formation of which usu-
cycle. This effect is referred to as covalent
ally requires a large amount of activation
catalysis (see the transaminases, for example;
energy, Ea (see p. 22). Since only a few A-B
p. 178). The principles of enzyme catalysis
complexes can produce this amount of en-
sketched out here are discussed in greater
ergy, a productive transition state arises
detail on p. 100 using the example of lactate
even less often than a collision complex. In
dehydrogenase.
solution, a large proportion of the activation
energy is required for the removal of the hy-
C. Principles of enzyme catalysis
dration shells between A and B. However,
charge displacements and other chemical
Although it is dif cult to provide quantitative
processes within the reactants also play a
estimates of the contributions made by indi-
role. As a result of these limitations, conver-
vidual catalytic effects, it is now thought that
sion only happens occasionally in the absence
the enzyme’s stabilization of the transition
of a catalyst, and the reaction rate v is low,
state is the most important factor. It is not
even when the reaction is thermodynamically
tight binding of the substrate that is impor-
possible—i. e., when ∆G < 0 (see p. 18).
tant, therefore—this would increase the acti-
vation energy required by the reaction, rather
than reducing it—but rather the binding of the
B. Enzyme-catalyzed reaction
transition state. This conclusion is supported
Shown here is a sequential mechanism in
by the very high af nity of many enzymes for
which substrates A and B are bound and prod-
analogues of the transition state (see p. 96). A
ucts C and D are released, in that order. An-
simple mechanical analogy may help clarify
other possible reaction sequence, known as
this (right). To transfer the metal balls (the
the “ping-pong mechanism,” is discussed on
reactants) from location EA (the substrate
p. 94.
state) via the higher-energy transition state
Enzymes are able to bind the reactants
to EP (the product state), the magnet (the
(their substrates) specifically at the active cen-
catalyst) has to be orientated in such a way
ter. In the process, the substrates are oriented
that its attractive force acts on the transition
in relation to each other in such a way that
state (bottom) rather than on EA (top).
they take on the optimal orientation for the
formation of the transition state (1-3). The
proximity and orientation of the substrates
therefore strongly increase the likelihood
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
91
A. Uncatalyzed reaction
Reactants
Collision
Transition
Collision
Products
complex 1
state
complex 2
C
+
+
D
A
B
B. Enzyme-catalyzed reaction
A
B
X
Y
X
Y
Z
Z
2. E.A-complex
3. E.A.B-complex
X
Y
X
Y
Z
Active site
Z
1. Free enzyme E
4.Transition state E
X
Y
X
Y
Z
Z
D
C
6. E.D-complex
5. E.C.D-complex
C. Principles of enzyme catalysis
Approximation and orientation
a
of the substrates
Transition
state
b
Exclusion of water
c
Stabilization of the
transition state
EA
EP
Ea
d
Group transfer
Stabilization
G
c
a
c
of the EA complex
X
Y
b
EA
EP
d
Z
Stabilization
of the transition state
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
92
Metabolism
enzymes whose substrate saturation curves
Enzyme kinetics I
are shown in diagram 1, enzyme 2 has the
The kinetics of enzyme-catalyzed reactions
higher af nity for A
[Km = 1 mmol l-1);
(i. e., the dependence of the reaction rate on
Vmax, by contrast, is much lower than with
the reaction conditions) is mainly determined
enzyme 1.
by the properties of the catalyst. It is therefore
Since v approaches V asymptotically with
more complex than the kinetics of an uncata-
increasing values of [A], it is dif cult to obtain
lyzed reaction (see p. 22). Here we discuss
reliable values for Vmax—and thus for Km as
these issues using the example of a simple
well—from diagrams plotting v against [A]. To
first-order reaction (see p. 22)
get around this, the Michaelis-Menten equa-
tion can be arranged in such a way that the
measured points lie on a straight line. In the
A. Michaelis-Menten kinetics
Lineweaver-Burk plot
(2),
1/v is plotted
In the absence of an enzyme, the reaction rate v
against 1/[A]. The intersections of the line of
is proportional to the concentration of sub-
best fit with the axes then produce 1/Vmax
stance A (top). The constant k is the rate con-
and—1/Km. This type of diagram is very clear,
stant of the uncatalyzed reaction. Like all cat-
but for practical purposes it is less suitable for
alysts, the enzyme E (total concentration [E]t)
determining Vmax and Km. Calculation meth-
creates a new reaction pathway. Initially, A is
ods using personal computers are faster and
bound to E (partial reaction 1, left). If this
more objective.
reaction is in chemical equilibrium, then
with the help of the law of mass action—and
B. Isosteric and allosteric enzymes
taking into account the fact that [E]t = [E] +
[EA]—one can express the concentration [EA]
Many enzymes can occur in various conforma-
of the enzyme-substrate complex as a func-
tions (see p. 72), which have different catalytic
tion of [A] (left). The Michaelis constant Km
properties and whose proportion of the total
thus describes the state of equilibrium of the
number of enzyme molecules is influenced by
reaction. In addition, we know that kcat > k—in
substrates and other ligands (see pp. 116 and
other words, enzyme-bound substrate reacts
280, for example). Allosteric enzymes of this
to B much faster than A alone (partial reaction
type, which are usually present in oligomeric
2, right). kcat, the enzyme’s turnover number,
form, can be recognized by their S-shaped
corresponds to the number of substrate mol-
(sigmoidal) saturation curves, which cannot
ecules converted by one enzyme molecule per
be described using the Michaelis model. In
second. Like the conversion A B, the forma-
the case of isosteric enzymes (with only one
tion of B from EA is a first-order reaction—i. e.,
enzyme conformation, 1), the ef ciency of
v = k
[EA] applies. When this equation is
substrate binding
(dashed curve) declines
combined with the expression already de-
constantly with increasing [A], because the
rived for EA, the result is the Michaelis-
number of free binding sites is constantly
Menten equation.
decreasing. In most allosteric enzymes (2),
In addition to the variables v and [A], the
the binding ef ciency initially rises with in-
equation also contains two parameters that do
creasing
[A], because the free enzyme is
not depend on the substrate concentration
present
in a low-af nity conformation
[A], but describe properties of the enzyme
(square symbols), which is gradually con-
itself: the product kcat
[E]g is the limiting
verted into a higher-af nity form (round sym-
value for the reaction rate at a very high [A],
bols) as a result of binding with A. It is only at
the maximum velocity Vmax of the reaction
high [A] values that a lack of free binding sites
(recommended abbreviation: V). The Michae-
becomes noticeable and the binding strength
lis constant Km characterizes the af nity of the
decreases again. In other words, the af nity of
enzyme for a substrate. It corresponds to the
allosteric enzymes is not constant, but de-
substrate concentration at which v reaches
pends on the type and concentration of the
half of Vmax (if v = Vmax/2, then [A]/(Km +
ligand.
[A]) = 1/2, i. e. [A] is then = Km). A high af nity
of the enzyme for a substrate therefore leads
to a low Km value, and vice versa. Of the two
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
93
A. Michaelis Menten kinetics
Uncatalyzed reaction
Enzyme-catalyzed reaction
A
B
Enzyme-catalyzed reaction
v = k · [A ]
Partial reaction 1:
Partial reaction 2:
formation and decay
formation of the
of enzyme-substrate
product of EA
E
complex EA
Km
kcat
Km
kcat
EA
E + A
EA
[EA]
E + B
[A]
A
[EA] = [E]
·
g
v = k
K
cat · [EA]
m + [A]
E
Michaelis constant
Maximum velocity
Km (mol · l-1)
·
[A]
Vmax = kcat · [E]g (mol · l-1· s-1)
kcat [E]g
v =
+
[A]
Km
10
1.0
Enzyme 1
8
0.8
6
0.6
-1/K
m
4
0.4
-1/K
m
Enzyme 2
2
0.2
1/V
max
1/V
Vmax
Vmax
max
0
0.0
Km
Km
0
2
4
6
8
10
90
95
100
-1.0
-0.5
0.0
0.5
1.0
1.5
2.0
Concentration [A] (mM)
Reciprocal concentration 1/[A] (mM-1)
1. Hyperbolic plot
2. Lineweaver-Burk plot
B. Isosteric and allosteric enzymes
Isosteric
Allosteric
(n=1)
(n=3)
Vmax
Vmax
Bonding
strenght
Bonding
strenght
[A]
K
O.5
m
0
5
10
15
20
0
5
10
15
20
[A] (mmol · l-1)
[A] (mmol · l-1)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
94
Metabolism
B. Substrate specificity
Enzyme kinetics II
Enzymes “recognize” their substrates in a
The catalytic properties of enzymes, and con-
highly specific way (see p. 88). It is only the
sequently their activity (see p. 90), are influ-
marked substrate specificity of the enzymes
enced by numerous factors, which all have to
that makes a regulated metabolism possible.
be optimized and controlled if activity meas-
This principle can be illustrated using the ex-
urements are to be carried out in a useful and
ample of the two closely related proteinases
reproducible fashion. These factors include
trypsin and chymotrypsin. Both belong to the
physical quantities (temperature, pressure),
group of serine proteinases and contain the
the chemical properties of the solution (pH
same “triad” of catalytically active residues
value, ionic strength), and the concentrations
(Asp-His-Ser, shown here in green; see
of the relevant substrates, cofactors, and in-
p. 176). Trypsin selectively cleaves peptide
hibitors.
bonds on the C-terminal side of basic amino
acids (lysine and arginine), while chymotryp-
A. pH and temperature dependency of
sin is specific for hydrophobic residues. The
enzyme activity
substrate binding “pockets” of both enzymes
have a similar structure, but their amino acid
The effect of enzymes is strongly dependent
sequences differ slightly. In trypsin, a nega-
on the pH value (see p. 30). When the activity
tively charged aspartate residue
(Asp-189,
is plotted against pH, a bell-shaped curve is
red) is arranged in such a way that it can
usually obtained (1). With animal enzymes,
bind and fix the basic group in the side chain
the pH optimum—i. e., the pH value at which
of the substrate. In chymotrypsin, the “bind-
enzyme activity is at its maximum—is often
ing pocket” is slightly narrower, and it is lined
close to the pH value of the cells (i. e., pH 7).
with neutral and hydrophobic residues that
However, there are also exceptions to this. For
stabilize the side chains of apolar substrate
example, the proteinase pepsin (see p. 270),
amino acids through hydrophobic interac-
which is active in the acidic gastric lumen,
tions (see p. 28).
has a pH optimum of 2, while other enzymes
(at least in the test tube) are at their most
active at pH values higher than 9. The bell
C. Bisubstrate kinetics
shape of the activity-pH profile results from
Almost all enzymes—in contrast to the sim-
the fact that amino acid residues with ioniz-
plified description given on p. 92—have more
able groups in the side chain are essential for
than one substrate or product. On the other
catalysis. In example (1), these are a basic
hand, it is rare for more than two substrates to
group B (pKa = 8), which has to be protonated
be bound simultaneously. In bisubstrate reac-
in order to become active, and a second acidic
tions of the type A + B C + D, a number of
amino acid AH (pKa = 6), which is only active
reaction sequences are possible. In addition to
in a dissociated state. At the optimum pH of 7,
the sequential mechanisms
(see p. 90), in
around 90% of both groups are present in the
which all substrates are bound in a specific
active form; at higher and lower values, one
sequence before the product is released, there
or the other of the groups increasingly passes
are also mechanisms in which the first sub-
into the inactive state.
strate A is bound and immediately cleaved. A
The temperature dependency of enzymatic
part of this substrate remains bound to the
activity is usually asymmetric. With increas-
enzyme, and is then transferred to the second
ing temperature, the increased thermal
substrate B after the first product C has been
movement of the molecules initially leads to
released. This is known as the ping-pong
a rate acceleration (see p.22). At a certain
mechanism, and it is used by transaminases,
temperature, the enzyme then becomes un-
for example (see p. 178). In the Lineweaver—
stable, and its activity is lost within a narrow
Burk plot (right; see p. 92), it can be recog-
temperature difference as a result of denatu-
nized in the parallel shifting of the lines when
ration (see p. 74). The optimal temperatures of
[B] is varied.
the enzymes in higher organisms rarely ex-
ceed 50 °C, while enzymes from thermophilic
bacteria found in hot springs, for instance,
may still be active at 100 °C.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
95
A. pH and temperature dependency of enzyme activity
+
-
Proportion BH
Proportion A
1.0
1.0
BH+
A-
Denaturation
Enzyme
0.8
0.8
active
Increased
0.6
0.6
activity due
BH+
AH
BH+
A-
B
A-
to temperature
increase
0.4
0.4
BH+
0.2
0.2
A-
Enzyme
0.0
0.0
denatured
4
5
6
7
8
9
10
0
10
20
30
40
50
pH
Temperature (°C)
B. Substrate specificity
Asp-189
Substrate (-X-Arg-Y-)
Ser-189
Substrate (-X-Phe-Y-)
Ser-195
Ser-195
His-57
His-57
Asp-102
Asp-102
1
Trypsin (3.4.21.4)
2
Chymotrypsin (3.4.21.1)
-X-Y-Arg (Lys)-Z-
-X-Y-Tyr (Trp, Phe, Leu)-Z-
C. Bisubstrate kinetics
3. Covalent
A
intermediate
2. EA complex
product E'
X
Y
X
Y
{B}
Z
Z
X
Y
P1
Active center
B
Z
1. Free enzyme E
X
Y
X
Y
Z
Z
P2
0
2
4
5. EP2 complex
4. E'B complex
1/{A}(mM-1)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
96
Metabolism
“Suicide substrates” (5) are substrate ana-
Inhibitors
logs that also contain a reactive group. Ini-
Many substances can affect metabolic pro-
tially, they bind reversibly, and then they
cesses by influencing the activity of
form a covalent bond with the active center
enzymes. Enzyme inhibitors are particularly
of the enzyme. Their effect is therefore also
important here. A large proportion of
non-competitive. A well-known example of
medicines act as enzyme inhibitors. Enzyme-
this is the antibiotic penicillin (see p. 254).
kinetic experiments are therefore an impor-
Allosteric inhibitors bind to a separate
tant aspect of drug development and testing
binding site outside the active center
(6).
procedures. Natural metabolites are also in-
This results in a conformational change in
volved in regulatory processes as inhibitors
the enzyme protein that indirectly reduces
(see p. 114).
its activity (see p. 116). Allosteric effects prac-
tically only occur in oligomeric enzymes. The
kinetics of this type of system can no longer
A. Types of inhibitor
be described using the simple Micha-
Most enzyme inhibitors act reversibly—i. e.,
elis-Menten model.
they do not cause any permanent changes in
the enzyme. However, there are also irrever-
B. Inhibition kinetics
sible inhibitors that permanently modify the
target enzyme. The mechanism of action of an
In addition to the Lineweaver-Burk plot (see
inhibitor—its inhibition type—can be deter-
p. 92), the Eadie-Hofstee plot is also com-
mined by comparing the kinetics (see p. 92)
monly used. In this case, the velocity v is
of the inhibited and uninhibited reactions (B).
plotted against v /[A]. In this type of plot,
This makes it possible to distinguish compet-
Vmax corresponds to the intersection of the
itive inhibitors
(left) from noncompetitive
approximation lines with the v axis, while
inhibitors
(right), for example. Allosteric
Km is derived from the gradient of the lines.
inhibition is particularly important for meta-
Competitive and non-competitive inhibitors
bolic regulation (see below).
are also easily distinguishable in the Eadie—
Substrate analogs (2) have properties sim-
Hofstee plot. As mentioned earlier, competi-
ilar to those of one of the substrates of the
tive inhibitors only influence Km, and not
target enzyme. They are bound by the en-
Vmax. The lines obtained in the absence and
zyme, but cannot be converted further and
presence of an inhibitor therefore intersect on
therefore reversibly block some of the enzyme
the ordinate. Non-competitive inhibitors pro-
molecules present. A higher substrate concen-
duce lines that have the same slope (Km un-
tration is therefore needed to achieve a half-
changed) but intersect with the ordinate at a
maximum rate; the Michaelis constant Km
lower level. Another type of inhibitor, not
increases (B). High concentrations of the sub-
shown here, in which Vmax and Km are re-
strate displace the inhibitor again. The max-
duced by the same factor, is referred to as
imum rate Vmax is therefore not influenced by
uncompetitive. Inhibitors with purely uncom-
this type of inhibition. Because the substrate
petitive effects are rare. A possible explana-
and the inhibitor compete with one another
tion for this type of inhibition is selective
for the same binding site on the enzyme, this
binding of the inhibitor to the EA complex.
type of inhibition is referred to as compe-
Allosteric enzymes shift the target en-
titive. Analogs of the transition state (3) usu-
zyme’s saturation curve to the left
(see
ally also act competitively.
p. 92). In Eadie-Hofstee and Lineweaver-Burk
When an inhibitor interacts with a group
plots (see p. 92), allosteric enzymes are recog-
that is important for enzyme activity, but does
nizable because they produce curved lines
not affect binding of the substrate, the inhi-
(not shown).
bition is non-competitive (right). In this case,
Km remains unchanged, but the concentration
of functional enzyme [E]t, and thus Vmax, de-
crease. Non-competitive inhibitors generally
act irreversibly, by modifying functional
groups of the target enzyme (4).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
97
A. Types of inhibitor
Competitive
1.
Uninhibited
Noncompetitive
I
C
I
C
C
C
C
C
C
C
C
a
4. Modifying reagent
Allosteric
C
C
C
C
I
I
C
C
b
a
2. Substrate analogs
I
v
C
I
C
C
C
I
C
C
b
[A]
3. Transition state
6.
5. “Suicide substrate”
analog
B. Kinetics of inhibition
Competitive
Maximal
inhibition:
velocity V
10
V unchanged
10
8
8
Slope = Km
6
6
Non-
competitive
4
inhibition:
4
V reduced
2
2
Km
increased
Km
unchanged
0
0
0
10
20
80 90100
0.0
0.5
1.0
1.5
2.0
Substrate concentration [A] (mM)
v/[A] (nkat · mM -o)
1. Hyperbolic plot
2. Eadie Hofstee plot
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
98
Metabolism
The differences in sequence between the M
Lactate dehydrogenase: structure
and H subunits are mainly conservative—i. e.,
Lactate dehydrogenase (LDH, EC 1.1.1.27) is
both residues are of the same type, e. g. gly-
discussed in some detail here and on the
cine (G) and alanine (A), or arginine (R) and
next page as an example of the structure
lysine (K). Non-conservative exchanges are
and function of an enzyme.
less frequent—e. g., lysine (K) for glutamine
(Q), or threonine (T) for glutamic acid (E).
Overall, the H subunit contains more acidic
A. Lactate dehydrogenase: structure
and fewer basic residues than the M form, and
The active form of lactate dehydrogenase
it therefore has a more strongly negative
(mass 144 kDa) is a tetramer consisting of
charge. This fact is exploited to separate the
four subunits (1). Each monomer is formed
isoenzymes using electrophoresis
(2; see
by a peptide chain of
334
amino acids
pp. 78, 276). The isoenzyme LDH-1, consisting
(36 kDa). In the tetramer, the subunits
of four H subunits, migrates fastest, and the
occupy equivalent positions (1); each mono-
M4 isoenzyme is slowest.
mer has an active center. Depending on met-
abolic conditions, LDH catalyzes NADH-de-
The separation and analysis of isoenzymes
pendent reduction of pyruvate to lactate, or
in blood samples is important in the diagnosis
NAD+-dependent oxidation of lactate to pyru-
of certain diseases. Normally, only small
vate (see p. 18).
amounts of enzyme activity are found in se-
The active center of an LDH subunit is
rum. When an organ is damaged, intracellular
shown schematically in Fig. 2. The peptide
enzymes enter the blood and can be demon-
backbone is shown as a light blue tube. Also
strated in it (serum enzyme diagnosis). The
shown are the substrate lactate (red), the
total activity of an enzyme reflects the se-
coenzyme NAD+ (yellow), and three amino
verity of the damage, while the type of iso-
acid side chains (Arg-109, Arg-171, and His-
enzyme found in the blood provides evidence
195; green), which are directly involved in the
of the site of cellular injury, since each of the
catalysis. A peptide loop
(pink) formed by
genes is expressed in the various organs at
amino acid residues 98-111 is also shown. In
different levels. For example, the liver and
the absence of substrate and coenzyme, this
skeletal muscles mainly produce M subunits
partial structure is open and allows access to
of lactate dehydrogenase
(M for muscle),
the substrate binding site (not shown). In the
while the brain and cardiac muscle mainly
enzyme lactate NAD+ complex shown, the
express H subunits (H for heart). In conse-
peptide loop closes the active center. The cat-
quence, each organ has a characteristic isoen-
alytic cycle of lactate dehydrogenase is dis-
zyme pattern (3). Following cardiac infarction,
cussed on the next page.
for example, there is a strong increase in the
amount of LDH-1 in the blood, while the con-
centration of LDH-5 hardly changes. The iso-
B. Isoenzymes
enzymes of creatine kinase (see p. 336) are
There are two different LDH subunits in the
also of diagnostic importance.
organism—M and H—which have a slightly
different amino acid sequence and conse-
quently different catalytic properties. As these
two subunits can associate to form tetramers
randomly, a total of five different isoenzymes
of LDH are found in the body.
Fig. 1 shows sections from the amino acid
sequences of the two subunits, using the sin-
gle-letter notation (see p. 60). A common pre-
cursor gene was probably duplicated at some
point in evolution. The two genes then con-
tinued to develop further independently of
each other through mutation and selection.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
99
A. Lactate dehydrogenase: structure
Lactate
Pyruvate
Lactate
dehydrogenase
1.1.1.27
NADH + H
NAD
N
A
N
A
1. Tetramer 144 kDa
2. Active center
Substrate
Mobile
Coenzyme
Essential
(lactate)
loop
(NAD
)
amino
Enzyme
acids
protein
Arg-109
His-195
Arg-171
B. Isoenzymes
Lactate dehydrogenase M
RYLMGERLGVHPLSCHGWVLGEHGDSSVPVWSGMNVAGCSL
KTLHPDLGTD..
1. Gene
RYLMAEKLG I HPSSCHGW
I
LGEHGDSSVAVWSG
V
NVAG
VSLQELNPEMGTD..
Lactate dehydrogenase H
LDH1
(H4)
Skeletal muscle
LDH2
(M1H3)
Liver
LDH3
(M2H2)
Brain
LDH4
(M3H1)
Cardiac muscle
LDH5
(M4)
LDH5
LDH4
LDH3
LDH2
LDH1
2. Forms
3. Separation by gel electrophoresis
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
100
Metabolism
and coenzyme, so that water molecules are
Lactate dehydrogenase:
largely excluded during the electron transfer.
mechanism
We can now look at the partial reactions
involved in LDH-catalyzed pyruvate reduc-
The principles of enzyme catalysis discussed
tion.
on p. 90 can be illustrated using the reaction
mechanism of lactate dehydrogenase (LDH)
In the free enzyme, His195 is protonated
as an example.
(1). This form of the enzyme is therefore de-
scribed as E H+. The coenzyme NADH is
A. Lactate dehydrogenase: catalytic cycle
bound first (2), followed by pyruvate (3). It
is important that the carbonyl group of the
LDH catalyzes the transfer of hydride ions (see
pyruvate in the enzyme and the active site in
p. 32) from lactate to NAD+ or from NADH to
the nicotinamide ring of the coenzyme should
pyruvate.
have a fairly optimal position in relation to
each other, and that this orientation should
L-lactate + NAD+ ↔pyruvate + NADH + H+
become fixed (proximity and orientation of the
substrates). The 98-111 loop now closes over
The equilibrium of the reaction strongly fa-
the active center. This produces a marked
vors lactate formation. At high concentrations
decrease in polarity, which makes it easier
of lactate and NAD+, however, oxidation of
to achieve the transition state (4; water ex-
lactate to pyruvate is also possible
(see
clusion). In the transition state, a hydride ion,
p. 18). LDH catalyzes the reaction in both di-
H- (see p. 32), is transferred from the coen-
rections, but—like all enzymes—it has no ef-
zyme to the carbonyl carbon (group transfer).
fect on chemical equilibrium.
The transient—and energetically unfavora-
As the reaction is reversible, the catalytic
ble—negative charge on the oxygen that oc-
process can be represented as a closed loop.
curs here is stabilized by electrostatic inter-
The catalytic cycle of LDH is reduced to six
action with Arg-109 (stabilization of the tran-
“snapshots” here. Intermediate steps in catal-
sition state). At the same time, a proton from
ysis such as those shown here are extremely
His-195 is transferred to this oxygen atom
short-lived and therefore dif cult to detect.
(group transfer), giving rise to the enzyme-
Their existence was deduced indirectly from a
bound products lactate and NAD+ (5). After
large number of experimental findings—e. g.,
the loop opens, lactate dissociates from the
kinetic and binding measurements.
enzyme, and the temporarily uncharged imi-
Many amino acid residues play a role in the
dazole group in His-195 again binds a proton
active center of LDH. They can mediate the
from the surrounding water (6). Finally, the
binding of the substrate and coenzyme, or
oxidized coenzyme NAD+ is released, and the
take part in one of the steps in the catalytic
initial state (1) is restored. As the diagram
cycle directly. Only the side chains of three
shows, the proton that appears in the reaction
particularly important residues are shown
equation (NADH + H+) is not bound together
here. The positively charged guanidinium
with NADH, but after release of the lacta-
group of arginine-171 binds the carboxylate
te—i. e., between steps
(5) and
(6) of the
group of the substrate by electrostatic inter-
previous cycle.
action. The imidazole group of histidine-195 is
Exactly the same steps occur during the
involved in acid-base catalysis, and the side
oxidation of lactate to pyruvate, but in the
chain of arginine-109 is important for the sta-
opposite direction. As mentioned earlier, the
bilization of the transition state. In contrast to
direction which the reaction takes depends
His-195, which changes its charge during cat-
not on the enzyme, but on the equilibrium
alysis, the two essential arginine residues are
state—i. e., on the concentrations of all the
constantly protonated. In addition to these
reactants and the pH value (see p. 18).
three residues, the peptide loop 98-111 men-
tioned on p. 98 is also shown here schemati-
cally (red). Its function consists of closing the
active center after binding of the substrate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
101
A. Lactate dehydrogenase: catalytic cycle
E. H
E . H . NADH
Arg-109
HN NH2
HN NH2
NH2
H
C
C
C
H2N
H2N
O
N
H
Loop
H
H
98-111
H
H
H
C
N
C
N
NADH
HC
CH
HC
CH
N
N
H
H
His-195
H2N
NH2
H2N
NH2
C
Arg-171
C
HN
HN
1. Free enzyme
2. NADH bound
NH2
H
HN NH2
NH2
HN NH
C
H
2
C
C
O
N
C
H2N
O
N
H2N
H
H
H
H
H
H
H3C
H
H
H
O
C
C
N
NAD
C
N
Pyruvat
HC
CH
HC
CH
O
O
N
N
H
H
H2N
NH2
H2N
NH2
C
C
HN
HN
6. NAD bound
3. Pyruvate bound
NH2
NH2
H
H
HN
NH2
C
HN NH2
C
C
O
N
C
N
O
H2N
H2N
H
H
H
H3C
H
H
H
O
C
H
H
3C
C
N H
O O
O
C
H
HC
CH
C
N
Lactate
N
H
H
H2N
NH2
HC
CH
O O
C
N
Proton
Hydride
H
transfer
HN
transfer
H2N
NH2
C
HN
5. Lactate bound
4. Redox reaction
E . NAD . Lactate
Transition state
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
102
Metabolism
absorption is recorded at a constant
Enzymatic analysis
wavelength of 340 nm. The uncatalyzed LDH
Enzymes play an important role in biochem-
reaction is very slow. It is only after addition
ical analysis. In biological material—e. g., in
of the enzyme that measurable quantities of
body fluids—even tiny quantities of an en-
NADH are formed and absorption increases.
zyme can be detected by measuring its cata-
Since according to the Beer-Lambert law the
lytic activity. However, enzymes are also used
rate of the increase in absorption ∆A/∆t is
as reagents to determine the concentrations
proportional to the reaction rate ∆c/∆t. The
of metabolites—e. g., the blood glucose level
absorption coef cient ε at 340 nm or compar-
(C). Most enzymatic analysis procedures use
ison with a standard solution can be used to
the method of spectrophotometry (A).
calculate LDH activity.
A. Principle of spectrophotometry
C. Enzymatic determination of glucose
Many substances absorb light in the visible or
Most biomolecules do not show any absorp-
ultraviolet region of the spectrum. This prop-
tion in the visible or ultraviolet spectrum. In
erty can be used to determine the concentra-
addition, they are usually present in the form
tion of such a substance. The extent of light
of mixtures with other—similar—compounds
absorption depends on the type and concen-
that would also react to a chemical test pro-
tration of the substance and on the wave-
cedure. These two problems can be avoided
length of the light used. Monochromatic
by using an appropriate enzyme to produce a
light—i. e., light with a defined wavelength
colored dye selectively from the metabolite
isolated from white light using a monochrom-
that is being analyzed. The absorption of the
ator—is therefore used. Monochromatic light
dye can then be measured.
with an intensity of I0 is passed through a
A procedure (1) that is often used to mea-
rectangular vessel made of glass or quartz (a
sure glucose when monitoring blood glucose
cuvet), which contains a solution of the ab-
levels (see p. 160) involves two successive re-
sorbing substance. The absorption A of the
actions. The glucose-specific enzyme glucose
solution (often also referred to as its extinc-
oxidase (obtained from fungi) first produces
tion) is defined as the negative decadic loga-
hydrogen peroxide, H2O2, which in the second
rithm of the quotient I/I0. The Beer-Lambert
step—catalyzed by a peroxidase—oxidizes a
law states that A is proportional to the con-
colorless precursor into a green dye
(2).
centration c of the absorbing substance and
When all of the glucose in the sample has
the thickness d of the solution it passes
been used up, the amount of dye formed—
through. As mentioned earlier, the absorption
which can be measured on the basis of its
coef•cient ε depends on the type of substance
light absorption—is equivalent to the quantity
and the wavelength.
of glucose originally present.
B. Measurement of lactate dehydrogenase
activity
Measurement of lactate dehydrogenase (LDH)
activity takes advantage of the fact that while
the reduced coenzyme NADH + H+ absorbs
light at 340 nm, oxidized NAD+ does not. Ab-
sorption spectra (i. e., plots of A against the
wavelength) for the substrates and the coen-
zymes of the LDH reaction are shown in Fig. 1.
Differences in absorption behavior between
NAD+ and NADH between 300 and 400 nm
result from changes in the nicotinamide ring
during oxidation or reduction (see p. 32). To
measure the activity, a solution containing
lactate and NAD+ is placed in a cuvet, and
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
103
A. Principle of spectrophotometry
Monochroma-
Monochroma-
White light
tic light,
tic light,
intensity
0
intensity
d
Absorption
Light source
Monochromator
Detector
Instrument
Light
Sample solution,
absorption
concentration c
Beer Lambert
Absorption
A = -log I
= ε . c. d
I
0
law
B. Assay of lactate dehydrogenase activity
∆A
∆c
∆c
v
2.0
∆A =
=
=
v ≈ Activity
ε
;
∆t · ε
∆t
;
Lactate
Pyruvate
0.1 mM
NAD
each
1.5
NADH
Addition
3 nkat
1.0
of LDH
2 nkat
NADH
1 nkat
Lactate; NAD
0.5
Lactate
∆A
NAD
Pyruvate
0
0
240
280
320
360
400
∆t
Time
1.
2.
Wavelength (nm)
C. Enzymatic determination of glucose
O2
Glucose-containing
sample solution
Glucose
Glucono-
Glucose
oxidase
lactone
1.1.3.4 [FAD]
Enzymes
2 H2O
Colorless
H2O2
precursor
∆A
∆A
—
[Green dye]
∞ =
ε
Peroxidase
Green
Colorless
[Green dye]
∞ = [Glucose]0
1.11.1.7
dye
precursor
[Heme]
0
10
20
Time (min)
1. Reaction
2. Procedure
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
104
Metabolism
metabolism. In contrast, reduced NADP+ is the
Coenzymes 1
most important reductant involved in biosyn-
thesis (see p. 112).
A. Coenzymes: definitions
The flavin coenzymes FMN and FAD (2, 3)
In many enzyme-catalyzed reactions, elec-
contain flavin (isoalloxazine) as a redox-active
trons or groups of atoms are transferred
group. This is a three-membered, N-contain-
from one substrate to another. This type of
ing ring system that can accept a maximum of
reaction always also involves additional mol-
two electrons and two protons during reduc-
ecules, which temporarily accept the group
tion. FMN carries the phosphorylated sugar
being transferred. Helper molecules of this
alcohol ribitol at the flavin ring. FAD arises
type are called coenzymes. As they are not
from FMN through bonding with AMP. The
catalytically active themselves, the less fre-
two coenzymes are functionally similar.
quently used term “cosubstrate” would be
They are found in dehydrogenases, oxidases,
more appropriate. In contrast to substrates
and monooxygenases. In contrast to the pyri-
for which a given enzyme is usually specific
dine nucleotides, flavin reactions give rise to
(see p. 88), coenzymes cooperate with many
radical intermediates (see p. 32). To prevent
enzymes of varying substrate specificity. We
damage to cell components, the flavins al-
have rather arbitrarily divided the coenzymes
ways remain bound as prosthetic groups in
here into group-transferring and redox coen-
the enzyme protein.
zymes. Strictly speaking, redox coenzymes
The role of ubiquinone (coenzyme Q, 4) in
also transfer groups—namely, reducing equiv-
transferring reducing equivalents in the res-
alents (see p. 32).
piratory chain is discussed on p. 140. During
Depending on the type of interaction with
reduction, the quinone is converted into the
the enzyme, a distinction is made between
hydroquinone (ubiquinol). The isoprenoid side
soluble coenzymes and prosthetic groups.
chain of ubiquinone can have various lengths.
Soluble coenzymes
(1) are bound like
It holds the molecule in the membrane, where
substrates during a reaction, undergo a chem-
it is freely mobile. Similar coenzymes are also
ical change, and are then released again. The
found in photosynthesis (plastoquinone; see
original form of the coenzyme is regenerated
p. 132). Vitamins E and K (see p. 52) also be-
by a second, independent reaction. Prosthetic
long to the quinone/hydroquinone systems.
groups (2), on the other hand, are coenzymes
L-Ascorbic acid (vitamin C, 5) is a powerful
that are tightly bound to the enzyme and re-
reducing agent. As an antioxidant, it provides
main associated with it during the reaction.
nonspecific protection against oxidative dam-
The part of the substrate bound by the coen-
age (see p. 284), but it is also an essential
zyme is later transferred to another substrate
cofactor for various monooxygenases and di-
or coenzyme of the same enzyme (not shown
oxygenases. Ascorbic acid is involved in the
in Fig. 2).
hydroxylation of proline and lysine residues
during the biosynthesis of collagen
(see
p. 344), in the synthesis of catecholamines
B. Redox coenzymes 1
(see p. 352) and bile acids (see p. 314), as
All oxidoreductases (see p. 88) require coen-
well as in the breakdown of tyrosine (see
zymes. The most important of these redox
p. 415). The reduced form of the coenzyme
coenzymes are shown here. They can act in
is a relatively strong acid and forms salts,
soluble form (S) or prosthetically (P). Their
the ascorbates. The oxidized form is known
normal potentials E0
are shown in addition
as dehydroascorbic acid. The stimulation of
to the type of reducing equivalent that they
the immune system caused by ascorbic acid
transfer (see p. 18).
has not yet been fully explained.
The pyridine nucleotides NAD+ and NADP+
(1) are widely distributed as coenzymes of
dehydrogenases. They transport hydride ions
(2e- and 1 H+; see p. 32) and always act in
soluble form. NAD+ transfers reducing equiv-
alents from catabolic pathways to the respi-
ratory chain and thus contributes to energy
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
105
A. Coenzymes: definitions
Substrate 1
Coenzyme
Substrate 2
Coenzyme
Group
1.
(form 1)
transfer
(form 2)
a
b
c
d
Prosthetic
Prosthetic
group (form1)
Substrate 1
Substrate 2
group (form 2)
2.
B. Redox coenzymes
Trans-
Eol
Coenzyme
Oxidized form
Reduced form
Type
ferred
(V)
1. NAD(P)
O
L
H
-0.32
H
H
H
H
N
H
C
NH2
C
N
N
A
N
A
C
N
O
O H
C
H
N
H
H
O
C
C
CH2O P O
P O CH2
N
N
H
C
O
O
N
A
N
A
NH2
O
O
H H
H H
P
P
H
H
H
H
H
N
H
ox.
red.
OH OH
OH OH
2. Flavin
O
H
H
O
P
2[H]
-0.3
mononucleotide
to
H3C
N
C
H3C
C
N
C
H
(FMN)
NH
C
C
N
+0.2
C
C
C
C
H3C
N
N
O
H3C
C
N
N
O
H
CH2
H
F
F
H
C
OH
H
C
OH
Ribitol (Rit)
ox.
red.
H
C
OH
O
H2C O P
O
O
3. Flavin
H
O
P
2[H]
-0.3
H
H
adenine
to
H3C
N
C
N
dinucleotide
NH
+0.2
C
(FAD)
C
N
C
N
H3C
N N
O
H
C
C
C
H
N
O
O
N
F
A
F
A
O
O P O P O CH2
H3C
N
C
Ribitol
NH
O
O
O
ox.
red.
H H
C
H
3C
N
N
O
H
H
OH OH
4. Ubiquinone
O
OH
L
2[H]
-0
(coenzym Q)
to
H3CO
CH3
H3CO
CH3
OH
+0.2
H3CO
H3CO
H
O
OH
CH3
6 -10
OH
5. Ascorbic
O
O
HO
O
L
2[H]
+0.1
acid
O
O
O
HO
HO
C
H
HO
C H
CH2OH
CH2OH
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
106
Metabolism
of energetic coupling (see p. 124) also allow
Coenzymes 2
endergonic processes to proceed. Metabolites
are often made more reactive (“activated”) as
A. Redox coenzymes 2
a result of the transfer of phosphate residues
In lipoic acid (6), an intramolecular disulfide
(phosphorylation). Bonding with nucleoside
bond functions as a redox-active structure. As
diphosphate residues (mainly UDP and CDP)
a result of reduction, it is converted into the
provides activated precursors for polysac-
corresponding dithiol. As a prosthetic group,
charides and lipids (see p. 110). Endergonic
lipoic acid is usually covalently bound to a
formation of bonds by ligases (enzyme class
lysine residue (R) of the enzyme, and it is
6) also depends on nucleoside triphosphates.
then referred to as lipoamide. Lipoamide is
Acyl residues are usually activated by
mainly involved in oxidative decarboxylation
transfer to coenzyme A (2). In coenzyme A
of
2-oxo acids
(see p. 134). The peptide
(see p. 12), pantetheine is linked to 3 -phos-
coenzyme glutathione is a similar disulfide/
pho-ADP by a phosphoric acid anhydride
dithiol system (not shown; see p. 284).
bond. Pantetheine consists of three compo-
Iron-sulfur clusters (7) occur as prosthetic
nents connected by amide bonds—pantoic
groups in oxidoreductases, but they are also
acid, E-alanine, and cysteamine. The latter
found in lyases—e. g., aconitase (see p. 136)
two components are biogenic amines formed
and other enzymes. Iron-sulfur clusters con-
by the decarboxylation of aspartate and
sist of 2-4 iron ions that are coordinated with
cysteine, respectively. The compound formed
cysteine residues of the protein (-SR) and
from pantoic acid and β−alanine (pantothenic
with anorganic sulfide ions (S). Structures of
acid) has vitamin-like characteristics for hu-
this type are only stable in the interior of
mans (see p. 368). Reactions between the
proteins. Depending on the number of iron
thiol group of the cysteamine residue and
and sulfide ions, distinctions are made be-
carboxylic acids give rise to thioesters, such
tween [Fe2S2], [Fe3S4], and [Fe4S4] clusters.
as acetyl CoA. This reaction is strongly ender-
These structures are particularly numerous
gonic, and it is therefore coupled to exergonic
in the respiratory chain
(see p. 140), and
processes. Thioesters represent the activated
they are found in all complexes except com-
form of carboxylic acids, because acyl residues
plex IV.
of this type have a high chemical potential
Heme coenzymes (8) with redox functions
and are easily transferred to other molecules.
exist in the respiratory chain (see p. 140), in
This property is often exploited in metabo-
photosynthesis (see p. 128), and in monooxy-
lism.
genases and peroxidases (see p. 24). Heme-
Thiamine diphosphate (TPP, 3), in coopera-
containing proteins with redox functions are
tion with enzymes, is able to activate alde-
also referred to as cytochromes. In cyto-
hydes or ketones as hydroxyalkyl groups and
chromes, in contrast to hemoglobin and myo-
then to pass them on to other molecules. This
globin, the iron changes its valence (usually
type of transfer is important in the transketo-
between +2 and +3). There are several classes
lase reaction, for example (see p. 152). Hy-
of heme (a, b, and c), which have different
droxyalkyl residues also arise in the decar-
types of substituent - R1 to - R3. Hemoglobin,
boxylation of oxo acids. In this case, they are
myoglobin, and the heme enzymes contain
released as aldehydes or transferred to lipo-
heme b. Two types of heme a are found in
amide residues of 2-oxoacid dehydrogenases
cytochrome c oxidase
(see p. 132), while
(see p. 134). The functional component of TPP
heme c mainly occurs in cytochrome c, where
is the sulfur- and nitrogen-containing thiazole
it is covalently bound with cysteine residues
ring.
of the protein part via thioester bonds.
B. Group-transferring coenzymes 1
The nucleoside phosphates (1) are not only
precursors for nucleic acid biosynthesis; many
of them also have coenzyme functions. They
serve for energy conservation, and as a result
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
107
A. Redox coenzymes 2
Coenzyme
Oxidized form
Reduced form
Type
Trans-
Eol
ferred
6. Lipoamide
P
2[H]
–0.29
S S
H
H
HC
CH
S S
H
S
S
HS SH
C
H
HC
CH
N
2
C
H2
ox.
red.
O
n+
m+
-0.6
7. Iron-
[Fe2 S2]
[Fe4 S4]
P
1e
to
sulfur
S Fe
RS
S
SR
RS
Fe S
+0.5
cluster
Fe
Fe
Fe
S
RS
S
SR
RS
S Fe
SR
1e
0
8. Heme
P
R2
CH3
R2
CH3
to
H3C
R3
H3C
R3
+0.5
A
B
A
B
N
N
N
N
Fe3+
Fe2+
N
N
N
N
3+
2+
D
C
D
C
R1
CH3
R1
CH3
ox.
red.
COO
COO
COO
COO
B. Group-transferring coenzymes 1
Coenzyme
Free form
Charged form
Group(s)
Important
(symbol)
trans-
enzymes
ferred
1. Nucleoside
P
Phospho-
1. phosphates
transferases
O
O
O
Base
Nucleotidyl-
B-Rib
transferases
O
P
O
P
O
P
O
CH2
O
B-Rib-
P
(2.7.n.n)
O
O
O
Ligases
N
H
P P P
(6.n.n.n)
B-Rib- P
P
H
H
OH OH
H
H CH3
O
2. Coenzyme A
Acyl
Acyltrans-
CH2
N C C C CH2
O P O
residues
ferases
CH2
O OH CH3
O
NH2
(2.3.n.n)
C
O
C O
O
P O
N
N
H
N
O
HC
CoA trans-
H
N
CH
2
CH2
N
N
ferases
S
O
CH2
CH2
(2.8.3.n)
A
H H
S
H
H
CH2
C O
O OH
CH3
SH
O
P O
O
3. Thiamine
Hydroxy-
Decarboxy-
R
3. diphosphate
H
alkyl
lases (4.1.1.n)
H3C
N
NH2
H
C
OH
residues
Oxoacid de-
N S
hydrogenases
N
CH2 N S
(1.2.4. n)
TPP
Transketolase
H3C
CH2
(2.2.1.1)
H3C
CH2
CH2O
P P
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
108
Metabolism
THF derivatives in the biosynthesis of DNA
Coenzymes 3
precursors, the enzymes involved in THF me-
tabolism are primary targets for cytostatic
A. Group-transferring coenzymes 2
drugs (see p. 402).
Pyridoxal phosphate (4) is the most important
coenzyme in amino acid metabolism. Its role
The cobalamins (7) are the chemically most
in transamination reactions is discussed in
complex form of coenzyme. They also repre-
detail on p. 178. Pyridoxal phosphate is also
sent the only natural substances that contain
involved in other reactions involving amino
the transition metal cobalt (Co) as an essential
acids, such as decarboxylations and dehydra-
component. Higher organisms are unable to
tions. The aldehyde form of pyridoxal phos-
synthesize cobalamins themselves, and are
phate shown here (left) is not generally found
therefore dependent on a supply of vitamin
in free form. In the absence of substrates, the
B12 synthesized by bacteria (see p. 368).
aldehyde group is covalently bound to the ε-
The central component of the cobalamins
amino group of a lysine residue as aldimine
is the corrin ring, a member of the tetrapyr-
(“Schiff’s base”). Pyridoxamine phosphate
roles, at the center of which the cobalt ion is
(right) is an intermediate of transamination
located. The end of one of the side chains of
reactions. It reverts to the aldehyde form by
the ring carries a nucleotide with the unusual
reacting with 2-oxoacids (see p. 178).
base dimethylbenzimidazole. The ligands for
the metal ion are the four N atoms of the
Biotin (5) is the coenzyme of the carboxy-
pyrrole ring, a nitrogen from dimethylbenzi-
lases. Like pyridoxal phosphate, it has an
midazole, and a group X, which is organo-
amide-type bond via the carboxyl group
metallically bound—i. e., mainly covalently.
with a lysine residue of the carboxylase. This
bond is catalyzed by a specific enzyme. Using
In methylcobalamin, X is a methyl group.
ATP, biotin reacts with hydrogen carbonate
This compound functions as a coenzyme for
(HCO3-) to form N-carboxybiotin . From this
several methyltransferases, and among other
activated form, carbon dioxide (CO2) is then
things is involved in the synthesis of methio-
transferred to other molecules, into which a
nine from homocysteine (see p. 418). How-
carboxyl group is introduced in this way. Ex-
ever, in human metabolism, in which methio-
amples of biotindependent reactions of this
nine is an essential amino acid, this reaction
type include the formation of oxaloacetic acid
does not occur.
from pyruvate (see p. 154) and the synthesis
of malonyl-CoA from acetyl-CoA (see p. 162).
Adenosylcobalamin (coenzyme B12) carries
a covalently bound adenosyl residue at the
Tetrahydrofolate (THF, 6) is a coenzyme
metal atom. This is a coenzyme of various
that can transfer C1 residues in different oxi-
isomerases, which catalyze rearrangements
dation states. THF arises from the vitamin folic
following a radical mechanism. The radical
acid (see p. 366) by double hydrogenation of
arises here through homolytic cleavage of the
the heterocyclic pterin ring. The C1
units
bond between the metal and the adenosyl
being transferred are bound to N-5, N-10, or
group. The most important reaction of this
both nitrogen atoms. The most important de-
type in animal metabolism is the rearrange-
rivatives are:
ment of methylmalonyl-CoA to form succinyl-
a) N5-formyl-THF and N10-formyl-THF, in
CoA, which completes the breakdown of odd-
which the formyl residue has the oxidation
numbered fatty acids and of the branched
state of a carboxylic acid;
amino acids valine and isoleucine
(see
b) N5-methylene-THF, with a C1 residue in
pp. 166 and 414).
the oxidation state of an aldehyde; and
c) N5-methyl-THF, in which the methyl
group has the oxidation state of an alcohol.
C1 units transferred by THF play a role in
the synthesis of methionine (see p. 412), pu-
rine nucleotides (see p. 188), and dTMP (see
p. 190), for example. Due to the central role of
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
109
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
110
Metabolism
3. Phosphoadenosine phosphosulfate (PAPS)
Activated metabolites
Sulfate residues occur as strongly polar
Many coenzymes (see pp. 104ff.) serve to ac-
groups in various biomolecules—e. g., in gly-
tivate molecules or groups that are poorly
cosaminoglycans (see p. 346) and conjugates
reactive. Activation consists of the formation
of steroid hormones and xenobiotics
(see
of reactive intermediate compounds in which
p. 316). In the synthesis of the “activated sul-
the group concerned is located at a higher
fate” PAPS, ATP first reacts with anorganic
chemical potential and can therefore be
sulfate to form adenosine phosphosulfate
transferred to other molecules in an exer-
(APS, a). This intermediate already contains
gonic reaction (see p. 124). Acetyl-CoA is an
the “energy-rich” mixed anhydride bond be-
example of this type of compound (see p. 12).
tween phosphoric acid and sulfuric acid. In
ATP and the other nucleoside triphosphate
the second step, the 3 -OH group of APS is
coenzymes not only transfer phosphate resi-
phosphorylated, with ATP being used again.
dues, but also provide the nucleotide compo-
After transfer of the sulfate residue to OH
nents for this type of activation reaction. On
groups (c), adenosine-3 ,5 -bisphosphate re-
this page, we discuss metabolites or groups
mains.
that are activated in the metabolism by bond-
ing with nucleosides or nucleotides. Inter-
mediates of this type are mainly found in
4. S-adenosyl methionine (SAM)
the metabolism of complex carbohydrates
The coenzyme tetrahydrofolate (THF) is the
and lipids.
main agent by which C1 fragments are trans-
ferred in the metabolism. THF can bind this
A. Activated metabolites
type of group in various oxidation states and
pass it on (see p. 108). In addition, there is
1. Uridine diphosphate glucose (UDPglucose)
“activated methyl,” in the form of S-adenosyl
The inclusion of glucose residues into poly-
methionine (SAM). SAM is involved in many
mers such as glycogen or starches is an ender-
methylation reactions—e. g., in creatine syn-
gonic process. The activation of the glucose
thesis (see p. 336), the conversion of norepi-
building blocks that is required for this takes
nephrine into epinephrine (see p. 352), the
places in several steps, in which two ATPs are
inactivation of norepinephrine by methyla-
used per glucose. After the phosphorylation of
tion of a phenolic OH group (see p. 316), and
free glucose, glucose 6-phosphate is isomer-
in the formation of the active form of the
ized to glucose 1-phosphate (a), reaction with
cytostatic drug
6-mercaptopurine
(see
UTP (b) then gives rise to UDPglucose, in
p. 402).
which the anomeric OH group at C-1 of the
SAM is derived from degradation of the
sugar is bound with phosphate. This “energy-
proteinogenic amino acid methionine, to
rich” compound (an acetal phosphate) allows
which the adenosyl residue of an ATP mole-
exergonic transfer of glucose residues to gly-
cule is transferred. After release of the acti-
cogen (c; see pp. 156, 408) or other acceptors.
vated methyl group, S-adenosyl homocys-
teine (SAH) is left over. This can be converted
back into methionine in two further steps.
2. Cytidine diphosphate choline (CDPcholine)
Firstly, cleavage of the adenosine residue
The amino alcohol choline is activated for in-
gives rise to the non-proteinogenic amino
clusion in phospholipids following a similar
acid homocysteine, to which a methyl group
principle (see p. 170). Choline is first phos-
is transferred once again with the help of N5-
phorylated by ATP to form choline phosphate
methyl-THF (see p. 418). Alternatively, homo-
(a), which by reaction with CTP and cleavage
cysteine can also be broken down into pro-
of diphosphate, then becomes CDPcholine. In
pionyl-CoA.
contrast to (1), it is not choline that is trans-
ferred from CDPcholine, but rather choline
phosphate, which with diacylglycerol yields
phosphatidylcholine (lecithin).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Enzymes
111
A. Activated metabolites
ADP
U
P
A
P
P
P
Glc
P
P
P
P
a
O
Glc
HOCH2
Glucose
C
Glc
H
O
H
1-phosphate
HC
NH
b
O
O
OH
H H
HC
C
Glycogen
Extended glycogen
HO
O P O
P
O
CH2
N
O
P
P
H
OH
O
Glc
O
O
H H
H
H
U
U
c
OH OH
Glc
P
P
P
P
UDP-glucose
1. Uridine diphosphate glucose (UDP-glucose)
ADP
C
A
NH2
P
P
P
Choline P
P
P
P
a
C
HC
NH
Choline
CH3
O
O
HC
C
b
N
O
Diacylglycerol
Phosphatidylcholine
H3C
N
(CH2)2
O P O
P O
CH2
O
CH
3
O
O
H H
P
P
H
H
P
Choline
OH OH
CDP-choline
C
C
c
Choline
P
P
P
2. Cytidine diphosphate choline (CDPcholine)
P P
A
A
A
H
H
P
P
P
P
P
P
P
N
O3S
a
C
N
C
N
b
SO42
O
O
HC
A
C
CH
N
P
P
O
S
O
P
O
CH2
N
OH
O
SO3
O
O
O
H H
Sulfated
H
H
substrate
A
A
P
c
P
O OH
O3S
O
P O
P
P
O
PAPS
3. Phosphoadenosine phosphosulfate (PAPS)
A
a
P
P
P
Methionine
Homocysteine
CH3
A
N5-Methyl-
THF
THF
Adenosine
b
THF
COO
H
H
N
d
H3N
C
H
3
P
CH2
C
N
C
N
CH3
CH
H
C
2
C
C
R
N
Methylated
H3C
S
CH2
N
H
R
substrate
O
H3C
S
c
A
S
H H
A
H
H
4. S-adenosyl methionine (SAM)
S-adenosyl-
OH OH
homocysteine
SAM
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
112
Metabolism
arrows). Of the numerous metabolites in the
Intermediary metabolism
pool, only three particularly important repre-
Hundreds of chemical reactions are con-
sentatives—pyruvate, acetyl-CoA, and gly-
stantly taking place in every cell, and taken
cerol—are shown here. These molecules
together these are referred to as the metabo-
represent connecting links between the
lism. The chemical compounds involved in
metabolism of proteins, carbohydrates, and
this are known as metabolites. Outside of
lipids. The metabolite pool also includes the
the cell, almost all of the chemical changes
intermediates of the tricarboxylic acid cycle
in metabolites would only take place very
(6). This cyclic pathway has both catabolic and
slowly and without any specific direction. By
anabolic functions—i. e., it is amphibolic (vio-
contrast, organized sequences of chemical re-
let; see p. 138).
actions with a high rate of throughput, known
Waste products from the degradation of
as metabolic pathways, become possible
organic substances in animal metabolism
through the existence of specific enzymes
include carbon dioxide (CO2), water (H2O),
(see p. 88).
and ammonia (NH3). In mammals, the toxic
substance ammonia is incorporated into urea
and excreted in this form (see p. 182).
A. Intermediary metabolism: overview
The most important form of storage for
A number of central metabolic pathways are
chemical energy in all cells is adenosine
common to most cells and organisms. These
triphosphate (ATP, see p. 122). ATP synthesis
pathways, which serve for synthesis, degra-
requires energy—i. e., the reaction is ender-
dation, and interconversion of important me-
gonic. Conversely, cleavage of ATP into ADP
tabolites, and also for energy conservation,
and phosphate releases energy. Exergonic hy-
are referred to as the intermediary metabo-
drolysis of ATP, as a result of energetic
lism.
coupling (see p. 16), makes energy-depend-
In order to survive, all cells constantly re-
ent (endergonic) processes possible. For ex-
quire organic and inorganic nutrients, as well
ample, most anabolic pathways, as well as
as chemical energy, which is mainly derived
movement and transport processes, are en-
from ATP (see below). Depending on the way
ergy-dependent.
in which these needs are satisfied, organisms
The most important pathway for the syn-
can be classified into autotrophic and hetero-
thesis of ATP is oxidative phosphorylation
trophic groups. The autotrophs, which in-
(see p. 122). In this process, catabolic path-
clude plants and many microorganisms, can
ways first form reduced cofactors (NADH+H+,
synthesize organic molecules from inorganic
QH2, ETFH2). Electrons are then transferred
precursors (CO2). An autotrophic lifestyle is
from these compounds to oxygen. This
possible through photosynthesis, for exam-
strongly exergonic process is catalyzed by
ple (see p. 128). The heterotrophs—e. g., ani-
the respiratory chain and used indirectly for
mals and fungi—depend on organic substan-
the ATP synthesis (see p. 140). In anaerobic
ces supplied in their diet. The schema shown
conditions—i. e., in the absence of oxygen—
on this page provides an overview of animal
most organisms can fall back on ATP that
metabolism.
arises in glycolysis
(3). This less ef cient
The polymeric substances contained in the
type of ATP synthesis is referred to as fermen-
diet
(proteins, carbohydrates, and nucleic
tation (see p. 146).
acids—top) cannot be used by the organism
While NADH exclusively supplies oxidative
directly. Digestive processes first have to de-
phosphorylation, NADPH+H+—a very similar
grade them to monomers (amino acids, sug-
coenzyme—is the reducing agent for anabolic
ars, nucleotides). These are then mostly bro-
pathways. NADPH + H+ is mainly formed in
ken down by catabolic pathways (pink ar-
the pentose phosphate pathway (PPP, 1; see
rows) into smaller fragments. The metabolites
p. 152).
produced in this way (generally referred to as
the “metabolite pool”) are then either used to
obtain energy through further catabolic con-
version, or are built up again into more com-
plex molecules by anabolic pathways (blue
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic Regulation
113
A. Intermediary metabolism: overview
Food
Poly-
Nucleic
Iso-
Proteins
Fats
saccharides
acids
prenoids
Bases
Lipids
P
Amino
Fatty
Glucose
1
Pentoses
acids
acids
Glycerol
ATP
NADPH
2
3
Other
NADH
biosyn-
4
5
theses
NADH
Pyruvate
ETF
Acetyl
CoA
Catabolic pathway
Anabolic pathway
6
Amphibolic pathway
Storage compound
NADH
QH2
1
Pentose phosphate pathway
2
Gluconeogenesis
NH
3
3
Glycolysis
4
β-Oxidation
O2
Oxidative
5
Fatty acid biosynthesis
7
phospho-
6
Tricarboxylic acid cycle
ATP
rylation
7
Urea cycle
Urea
H2O
CO2
Waste products
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
114
Metabolism
Interconversion processes in most cases
Regulatory mechanisms
involve ATP-dependent phosphorylation of
the enzyme protein by a protein kinase or
A. Fundamental mechanisms of metabolic
dephosphorylation of it by a protein phospha-
regulation
tase (see p. 120). The phosphorylated form of
The activities of all metabolic pathways are
the key enzyme is usually the more active
subject to precise regulation in order to adjust
one, but the reverse may also occur.
the synthesis and degradation of metabolites
Modulation by ligands. An important vari-
to physiological requirements. An overview of
able that regulates flow through a metabolic
the regulatory mechanisms is presented here.
pathway is precursor availability (metabolite
Further details are shown on pp. 116ff.
A in the case shown here). The availability of
Metabolite flow along a metabolic pathway
precursor A increases along with the activity
is mainly determined by the activities of the
of the metabolic pathways that form A (3) and
enzymes involved (see p. 88). To regulate the
it decreases with increasing activity of other
pathway, it is suf cient to change the activity
pathways that also consume A (4). Transport
of the enzyme that catalyzes the slowest step
from one cell compartment to another can
in the reaction chain. Most metabolic path-
also restrict the availability of A.
ways have key enzymes of this type on which
Coenzyme availability can also often have a
the regulatory mechanisms operate. The ac-
limiting effect (5). If the coenzyme is regen-
tivity of key enzymes is regulated at three
erated by a second, independent metabolic
independent levels:
pathway, the speed of the second pathway
Transcriptional control. Here, Biosynthesis
can limit that of the first one. For example,
of the enzyme protein is influenced at the
glycolysis and the tricarboxylic acid cycle are
genetic level
(1). Interventions in enzyme
mainly regulated by the availability of NAD+
synthesis mainly affect synthesis of the cor-
(see p. 146). Since NAD+ is regenerated by the
responding mRNA—i. e., transcription of the
respiratory chain, the latter indirectly con-
gene coding for the enzyme. The term “tran-
trols the breakdown of glucose and fatty acids
scriptional control” is therefore used
(see
(respiratory control, see p. 144).
pp. 118, 244). This mechanism is mediated by
Finally, the activity of key enzymes can be
regulatory proteins (transcription factors) that
regulated by ligands
(substrates, products,
act directly on DNA. The genes have a special
coenzymes, or other effectors), which as allo-
regulatory segment for this purpose, known
steric effectors do not bind at the active center
as the promoter region, which contains bind-
itself, but at another site in the enzyme,
ing sites
(control elements) for regulatory
thereby modulating enzyme activity (6; see
proteins. The activity of these proteins is, in
p. 116). Key enzymes are often inhibited by
turn, affected by metabolites or hormones.
immediate reaction products, by end prod-
When synthesis of a protein is increased by
ucts of the reaction chain concerned (“feed-
transcriptional control, the process is referred
back” inhibition), or by metabolites from com-
to as induction; when it is reduced or sup-
pletely different metabolic pathways. The
pressed, it is referred to as repression. Induc-
precursors for a reaction chain can stimulate
tion and repression processes take some time
their own utilization through enzyme activa-
and are therefore not immediately effective.
tion.
Interconversion of key enzymes (2) takes
effect considerably faster than transcriptional
control. In this case, the enzyme is already
present at its site of effect, but it is initially
still inactive. It is only when needed that it is
converted into the catalytically active form,
after signaling and mediation from second
messengers (see p. 120) through an activating
enzyme (E1). If the metabolic pathway is no
longer required, an inactivating enzyme (E2)
returns the key enzyme to its inactive resting
state.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic Regulation
115
A. Fundamental mechanisms of metabolic regulation
Transcriptional
Induction
Repression
control
R
Gene
H
RP
RP
H
Transcription
1
RP
RP
Regulator
mRNA
protein
A
Translation
Z
Interconversion
Hormone
H
Key enzyme
(inactive)
A
P
P P P
2
Second
Inactivated
E1
E2
messenger
enzyme
A
Activated
P P
enzyme
Modulation
by ligands
End product
Key enzyme
A
A
B
C
Z
(active)
P
6
Meta-
bolic
pathway
Feedback
Coenzyme
Coenzyme
inhibition
3
Availability pre-
Competition
Regeneration
cursors, com-
for substrates
of the coenzyme
partmentation
or coenzymes
5
4
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
116
Metabolism
causes a left shift; it reduces both [A]0.5 and
Allosteric regulation
h (curve III). This type of allosteric effect was
The regulation of aspartate carbamoyltrans-
first observed in hemoglobin
(see p. 280),
ferase (ACTase), a key enzyme of pyrimidine
which can be regarded as an “honorary” en-
biosynthesis (see p. 188) is discussed here as
zyme.
an example of allosteric regulation of enzyme
activity. Allosteric effects are mediated by the
C. R and T states
substrate itself or by inhibitors and activators
(allosteric effectors, see p. 114). The latter bind
Allosteric enzymes are almost always oligo-
at special sites outside the active center, pro-
mers with 2-12 subunits. ACTase consists of
ducing a conformational change in the en-
six catalytic subunits (blue) and six regulatory
zyme protein and thus indirectly lead to an
subunits (yellow). The latter bind the allo-
alteration in its activity.
steric effectors CTP and ATP. Like hemoglobin,
ACTase can also be present in two conforma-
tions—the less active T state (for “tense”) and
A. Aspartate carbamoyltransferase:
the more active R state (for “relaxed”). Sub-
reaction
strates and effectors influence the equili-
ACTase catalyzes the transfer of a carbamoyl
brium between the two states, and thereby
residue from carbamoyl phosphate to the
give rise to sigmoidal saturation behavior.
amino group of L-aspartate. The N-carbamoyl
With increasing aspartate concentration, the
L-aspartate formed in this way already con-
equilibrium is shifted more and more toward
tains all of the atoms of the later pyrimidine
the R form. ATP also stabilizes the R confor-
ring (see p. 188). The ACTase of the bacterium
mation by binding to the regulatory subunits.
Escherichia coli is inhibited by cytidine tri-
By contrast, binding of CTP to the same sites
phosphate (CTP), an end product of the ana-
promotes a transition to the T state. In the
bolic metabolism of pyrimidines, and is ac-
case of ACTase, the structural differences be-
tivated by the precursor ATP.
tween the R and T conformations are partic-
ularly dramatic. In T R conversion, the cat-
alytic subunits separate from one another by
B. Kinetics
1.2 nm, and the subunits also rotate around
In contrast to the kinetics of isosteric (normal)
the axis of symmetry. The conformations of
enzymes, allosteric enzymes such as ACTase
the subunits themselves change only slightly,
have sigmoidal (S-shaped) substrate satura-
however.
tion curves (see p. 92). In allosteric systems,
the enzyme’s af nity to the substrate is not
D. Structure of a dimer
constant, but depends on the substrate con-
centration [A]. Instead of the Michaelis con-
The subunits of ACTase each consist of two
stant Km (see p. 92), the substrate concentra-
domains—i. e., independently folded partial
tion at half-maximal rate ([A]0.5) is given. The
structures. The N-terminal domain of the reg-
sigmoidal character of the curve is described
ulatory subunit (right) mediates interaction
by the Hill coef•cient h. In isosteric systems,
with CTP or ATP (green). A second, Zn2+-con-
h = 1, and h increases with increasing sig-
taining domain (Zn2+ shown in light blue)
moidicity.
establishes contact with the neighboring cat-
Depending on the enzyme, allosteric effec-
alytic subunit. Between the two domains of
tors can influence the maximum rate Vmax,
the catalytic subunit lies the active center,
the semi-saturation concentration [A]0.5, and
which is occupied here by two substrate ana-
the Hill coef cient h. If it is mainly Vmax that is
logs (red).
changed, the term “V system” is used. Much
more common are “K systems”, in which al-
losteric effects only influence [A]0.5 and h.
The K type also includes ACTase. The inhib-
itor CTP in this case leads to right-shifting of
the curve, with an increase in [A]0.5 and h
(curve II). By contrast, the activator ATP
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic Regulation
117
A. Aspartate carbamoyltransferase: reaction
C
COO
P P P
L-Aspartate
CH2
CTP
CTP
C
H3N H COO
COO
Aspartate
H2N
CH2
carbamoyl-
UMP
CO2
NH2
transferase
C
C
R-NH2
O
N
H
COO
C
2.1.3.2 [ Zn2
]
H
ATP
O
P
P
N-Carbamoyl
N-Carbamoyl-
P
L-aspartate
A
P
P
P
ATP
B. Kinetics
C. R and T conformation
7
Active
C1
C
2
C3
center
6
R1
2 mM ATP
R2
h = 1.4
III
5
R1
R2
Effector-
C4
C6
4
binding
C5
Without
I
site
[A]0.5
effector
h = 2.0
T conformation
3
(less active)
0.5 mM CTP
2
II
h = 2.3
1
C2
C1
C3
0
R1
R2
0.0
5.0
10.0
15.0
R3
[Aspartate] (mM)
R1
R2
C4
C6
D. Structure of a dimer
C5
R conformation
(more active)
Regulatory
Substrate
subunit
ATP (CTP)
Catalytic
subunit
Zn2 domain
ATP/CTP-binding domain
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
118
Metabolism
B. Lactose operon
Transcription control
The well-investigated lactose operon of the
A. Functioning of regulatory proteins
bacterium Escherichial coli can be used here
as an example of transcriptional control. The
Regulatory proteins (transcription factors) are
lac operon is a DNA sequence that is simul-
involved in controlling gene expression in all
taneously subject to negative and positive
cells. These regulatory proteins bind to spe-
control. The operon contains the structural
cific DNA sequences and thereby activate or
genes for three proteins that are required for
inhibit the transcription of genes
(Tran-
the utilization of lactose (one transporter and
scription control). The effects of transcription
two enzymes), as well as control elements that
factors are usually reversible and are often
serve to regulate the operon.
controlled by ligands or by interconversion.
Since lactose is converted to glucose in the
The nomenclature for transcription factors
cell, there is no point in expressing the genes
is confusing. Depending on their mode of ac-
if glucose is already available. And indeed, the
tion, various terms are in use both for the
genes are in fact only transcribed when glu-
proteins themselves and for the DNA sequen-
cose is absent and lactose is present (3). This is
ces to which they bind. If a factor blocks tran-
achieved by interaction between two regula-
scription, it is referred to as a repressor; oth-
tory proteins. In the absence of lactose, the lac
erwise, it is called an inducer. DNA sequences
repressor blocks the promoter region
(2).
to which regulatory proteins bind are referred
When lactose is available, it is converted
to as control elements. In prokaryotes, control
into allolactose, which binds to the repressor
elements that serve as binding sites for RNA
and thereby detaches it from the operator (3).
polymerases are called promoters, whereas
However, this is still not suf cient for the
repressor-binding sequences are usually
transcription of the structural genes. For bind-
called operators. Control elements that bind
ing of the RNA polymerase to take place, an
activating factors are termed enhancers,
inducer—the catabolite activator protein
while elements that bind inhibiting factors
(CAP)—is required, which only binds to the
are known as silencers.
DNA when it is present as a complex with
The numerous regulatory proteins that are
3,5 -cyclo-AMP (cAMP; see p. 386). cAMP, a
known can be classified into four different
signal for nutrient deficiency, is only formed
groups (1-4), based on their mechanisms of
by E. coli in the absence of glucose.
action. Negative gene regulation—i. e.,
The interaction between the CAP-cAMP
switching off of the gene concerned—is car-
complex and DNA is shown in Fig. 4. Each
ried out by repressors. Some repressors only
subunit of the dimeric inducer (yellow or or-
bind to DNA (1a) in the absence of specific
ange) binds one molecule of cAMP (red). Con-
ligands (L). In this case, the complex between
tact with the DNA (blue) is mediated by two
the repressor and the ligand loses its ability to
“recognition helices” that interact with the
bind to the DNA, and the promoter region
major groove of the DNA. The bending of the
becomes accesible for binding of RNA poly-
DNA strand caused by CAP has functional sig-
merase (1b). It is often the free repressor that
nificance.
does not bind to the DNA, so that transcrip-
Transcription control is much more com-
tion is only blocked in the presence of the
plex in eukaryotes (see p. 244). The number
ligand (2a, 2b). A distinction between two
of transcription factors involved is larger, and
different types of positive gene regulation
in addition the gene activity is influenced by
can be made in the same way. If it is only
the state of the chromatin (see p. 238).
the free inducer that binds, then transcription
is inhibited by the appropriate ligand
(3).
Conversely, many inducers only become ac-
tive when they have bound a ligand (4). This
group includes the receptors for steroid hor-
mones, for example (see p. 378).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic Regulation
119
A. Functions of regulatory proteins
Negative regulation
Positive regulation
Without ligand
With ligand
Without ligand
With ligand
RNA
mRNA
poly-
RNA
merase
poly-
merase
DNA
Gen
Inducer
Promoter
L
Repressor
Ligand
L
1a
1b
3a
3b
L
L
2a
2b
4a
4b
B. Lactose operon
Genes for
5'
Catabolite
Adenin
permease
CH
2
activator
galactosidase
O
protein
Promoter
transacetylase
O
(CAP)
CAP
H H
O
P
H
H
3'
O
O OH
CAP-binding
Operator
1.
site
RNA polymerase
Recognition
cAMP
(only binds in the
helix
presence of
CAP · cAMP)
Lactose
lac Repressor
2.
cAMP
Lactose catabolism
Glucose
mRNA
CAP · cAMP
Transcription
Repressor
Lactose
allo-
lactose
Allolactose
complex
3.
4. CAP · cAMP bound to DNA
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
120
Metabolism
glycogen breakdown, releasing glucose, and
Hormonal control
at the same time inhibits glycogen synthesis.
In higher organisms, metabolic and other
Glucagon binds to receptors in the plasma
processes (growth, differentiation, control of
membrane (bottom left) and, with mediation
the internal environment) are controlled by
by a G-protein (see p. 386), activates the
hormones (see pp. 370 ff.)
enzyme adenylate cyclase, which forms the
second messenger
3,5 -cyclo-AMP
(cAMP)
from ATP. cAMP binds to another enzyme,
A. Principles of hormone action
protein kinase A
(PK-A), and activates it.
Depending on the type of hormone, hormone
PK-A has several points of attack. Through
signals are transmitted to the target cells in
phosphorylation, it converts the active form
different ways. Apolar (lipophilic) hormones
of glycogen synthase into the inactive form,
penetrate the cell and act in the cell nucleus,
thereby terminating the synthesis of glyco-
while polar (hydrophilic) hormones act on the
gen. Secondly, it activates another protein
external cell membrane.
kinase (not shown), which ultimately con-
Lipophilic hormones, which include the
verts the inactive form of glycogen phosphor-
steroid hormones, thyroxine, and retinoic
ylase into the active form through phosphor-
acid, bind to a specific receptor protein inside
ylation. The active phosphorylase releases glu-
their target cells. The complex formed by the
cose 1-phosphate from glycogen, which after
hormone and the receptor then influences
conversion into glucose 6-phosphate supplies
transcription of specific genes in the cell nu-
free glucose. In addition, via an inhibitor (I) of
cleus (see pp. 118, 244). The group of hydro-
protein phosphatase (PP), active PK-A inhibits
philic hormones (see p. 380) consists of hor-
inactivation of glycogen phosphorylase. When
mones derived from amino acids, as well as
the cAMP level falls again, phosphoprotein
peptide hormones and proteohormones.
phosphatases become active, which dephos-
Their receptors are located in the plasma
phorylate the various phosphoproteins in the
membrane. Binding of the hormone to this
cascade described, and thereby arrest glyco-
type of receptor triggers a signal that is trans-
gen breakdown and re-start glycogen synthe-
mitted to the interior of the cell, where it
sis. Activation and inactivation of proteins
controls the processes that allow the hor-
through phosphorylation or dephosphoryla-
mone signal to take effect (signal transduc-
tion is referred to as interconversion.
tion; see pp. 384ff.)
In contrast to glucagon, the peptide
hormone insulin (see p. 76) increases glyco-
gen synthesis and inhibits glycogen break-
B. Hormonal regulation of glucose
down. Via several intermediates, it inhibits
metabolism in the liver
protein kinase GSK-3 (bottom right; for de-
The liver plays a major role in glucose homeo-
tails, see p. 388) and thereby prevents inacti-
stasis in the organism (see p. 310). If glucose
vation of glycogen synthase. In addition, in-
deficiency arises, the liver releases glucose
sulin reduces the cAMP level by activating
into the blood, and when blood sugar levels
cAMP phosphodiesterase (PDE).
are high, it takes glucose up from the blood
Regulation by transcriptional control (top).
and converts it into different metabolites.
If the liver’s glycogen reserves have been ex-
Several hormones from both groups are in-
hausted, the steroid hormone cortisol main-
volved in controlling these processes. A very
tains glucose release by initiating the conver-
simplified version of the way in which they
sion of amino acids into glucose (gluconeo-
work is presented here. Glycogen is the form
genesis; see p. 154). In the cell nucleus, the
in which glucose is stored in the liver and
complex of cortisol and its receptor
(see
muscles. The rate of glycogen synthesis is
p. 378) binds to the promoter regions of var-
determined by glycogen synthase
(bottom
ious key enzymes of gluconeogenesis and
right), while its breakdown is catalyzed by
leads to their transcription. The active en-
glycogen phosphorylase (bottom left).
zymes are produced through translation of
Regulation by interconversion (bottom). If
the mRNA formed. Control of the transcrip-
the blood glucose level falls, the peptide
tion of the gluconeogenesis enzyme PEP car-
hormone glucagon is released. This activates
boxykinase is discussed on p. 244.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic Regulation
121
A. Principles of hormone action
H
H
Lipophilic
Hydrophilic
hormones
hormones
H
Receptor
Receptor
H
DNA
H
H
Tran-
Second messenger
scription
mRNA
Polysome
Translation
Cell response
Cell response
Protein
B. Hormonal regulation of glucose metabolism in the liver
Gene 1
Gene 2
Gene 3
Hormone
Transcription
receptor
complex
mRNA 1
mRNA 2
mRNA 3
Translation
Cortisol
Gluco-
corticoid
Enzyme 1
Enzyme 2
Enzyme 3
receptor
Glu-
Amino
Glucose
Glc-6-
P
cose
acids
Gluconeogenesis
ATP
cAMP
AMP
Adenylate
cyclase
PDE
Glc-1-
P
GSK-3
PKA
PI3K
P
P
Gα
I
IRS
P
Glycogen
Glucagon
PP
PP
receptor
Glycogen
Glycogen
Insulin
phosphorylase
synthase
receptor
Glucagon
Insulin
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
122
Metabolism
In standard conditions, the change in free
ATP
enthalpy ∆G0
(see p. 18) that occurs in the
The nucleotide coenzyme adenosine
hydrolysis of phosphoric acid anhydride
triphosphate
(ATP) is the most important
bonds amounts to -30 to -35 kJ mol-1 at
form of chemical energy in all cells. Cleavage
pH 7. The particular anhydride bond of ATP
of ATP is strongly exergonic. The energy this
that is cleaved only has a minor influence on
provides (∆G; see p. 16) is used to drive ender-
∆G0
(1-2). Even the hydrolysis of diphos-
gonic processes (such as biosynthesis and
phate (also known as pyrophosphate; 4) still
movement and transport processes) through
yields more than -30 kJ mol-1. By contrast,
energetic coupling (see p. 124). The other nu-
cleavage of the ester bond between ribose and
cleoside triphosphate coenzymes (GTP, CTP, and
phosphate only provides -9 kJ mol-1 (3).
UTP) have similar chemical properties to ATP,
In the cell, the ∆G of ATP hydrolysis is sub-
but they are used for different tasks in metab-
stantially larger, because the concentrations
olism (see p. 110).
of ATP, ADP and Pi are much lower than in
standard conditions and there is an excess of
ATP over ADP (see p. 18). The pH value and
A. ATP: structure
Mg2+ concentration also affect the value of ∆G.
In ATP, a chain of three phosphate residues is
The physiological energy yield of ATP hydrol-
linked to the 5 -OH group of the nucleoside
ysis to ADP and anorganic phosphate (Pi) is
adenosine (see p. 80). These phosphate resi-
probably around -50 kJ mol-1.
dues are termed α, β, and γ. The α phosphate
is bound to ribose by a phosphoric acid ester
C. Types of ATP formation
bond. The linkages between the three phos-
phate residues, on the other hand, involve
Only a few compounds contain phosphate
much more unstable phosphoric acid anhy-
residues with a group transfer potential (see
dride bonds. The active coenzyme is in fact
p. 18) that is high enough to transfer them to
generally a complex of ATP with an Mg2+
ADP and thus allow ATP synthesis. Processes
ion, which is coordinatively bound to the α
that raise anorganic phosphate to this type of
and β phosphates (Mg2+ ATP4-). However,
high potential are called substrate level phos-
the term “ATP” is usually used for the sake
phorylations (see p. 124). Reactions of this
of simplicity.
type take place in glycolysis (see p. 150) and
in the tricarboxylic acid cycle (see p. 136).
Another “energy-rich” phosphate compound
B. Hydrolysis energies
is creatine phosphate, which is formed from
The formula for phosphate residues shown in
ATP in muscle and can regenerate ATP as
Fig. A, with single and double bonds, is not an
needed (see p. 336).
accurate representation of the actual charge
Most cellular ATP does not arise in the way
distribution. In ATP, the oxygen atoms of all
described above (i. e., by transfer of phosphate
three phosphate residues have similarly
residues from organic molecules to ADP), but
strong negative charges (orange), while the
rather by oxidative phosphorylation. This
phosphorus atoms represent centers of posi-
process takes place in mitochondria (or as
tive charge. One of the reasons for the insta-
light-driven phosphorylation in chloroplasts)
bility of phosphoric anhydride bonds is the
and is energetically coupled to a proton gra-
repulsion between these negatively charged
dient over a membrane. These H+ gradients
oxygen atoms, which is partly relieved by
are established by electron transport chains
cleavage of a phosphate residue. In addition,
and are used by the enzyme ATP synthase as a
the free phosphate anion formed by hydroly-
source of energy for direct linking of anor-
sis of ATP is better hydrated and more strongly
ganic phosphate to ADP. In contrast to sub-
resonance-stabilized than the corresponding
strate level phosphorylation, oxidative phos-
residue in ATP. This also contributes to the
phorylation requires the presence of oxygen
strongly exergonic character of ATP hydroly-
(i. e., aerobic conditions).
sis.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
123
A. ATP: structure
Phosphoric acid
Phosphoric acid
N-glycosidic
NH
anhydride bonds
ester bond
2
bond
C
N
C
N
O
O
O
HC
C
CH
N
O
P
O
P
O
P
O
CH2
N
γ
β
α
O
O
O
O
H H
1. Formula
H
H
O
O
OH OH
O
P
O
P
α
O
CH2
β
O
O
O
H
γ
β
α
Ribose
Mg
H
Phosphate residue
OH
Adenosine
2. Mg2 -Complex
B. Hydrolysis energies
1
2
3
4
∆GOI
P
P
P
P
1
2
3
kJ · mol-1
+
P
P
+
+
P
P
-10
+
P
Positive
-20
P
Neutral
-30
Negative
1. Hydrolysis energies
2. ATP: charge density
C. Types of ATP formation
Electrons
Phosphorylated
substrate
Protons
ATP
A
synthase
P
P
P
P
A
H
ATP
P
P
P
P
Ared
ATP
Enzyme
e
Substrate
chain phos-
phorylation
A
H
A
A
P
P P
P P
Substrate
ADP
ADP
P
1. Phosphate transfer
2. Oxidative phosphorylation
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
124
Metabolism
phorylations,” as they represent individual
Energetic coupling
steps within metabolic pathways.
The cell stores chemical energy in the form of
In the glyceraldehyde 3-phosphate dehy-
“energy-rich” metabolites. The most impor-
drogenation reaction, a step involved in gly-
tant metabolite of this type is adenosine tri-
colysis (1; see also C), the aldehyde group in
phosphate (ATP), which drives a large number
glyceraldehyde 3-phosphate is oxidized into a
of energy-dependent reactions via energetic
carboxyl group. During the reaction, an anor-
coupling (see p. 16).
ganic phosphate is also introduced into the
product, producing a mixed acid anhy-
dride—1,3-bisphosphoglycerate. Phosphopyr-
A. Energetic coupling
uvate hydratase (“enolase”, 2) catalyzes the
The change in free enthalpy ∆G0 during hy-
elimination of water from 2-phosphoglycer-
drolysis (see p. 18) has been arbitrarily se-
ate. In the enol phosphate formed (phosphoe-
lected as a measure of the group transfer
nol pyruvate), the phosphate residue—in con-
potential of “energy-rich” compounds. How-
trast to
2-phosphoglycerate—is at an ex-
ever, this does not mean that ATP is in fact
tremely high potential (∆G0
of hydrolysis:
hydrolyzed in energetically coupled reactions.
-62 kJ mol-1). A third reaction of this type
If ATP hydrolysis and an endergonic process
is the formation of succinyl phosphate, which
were simply allowed to run alongside each
occurs in the tricarboxylic acid cycle as an
other, the hydrolysis would only produce
individual step in the succinyl CoA ligase re-
heat, without influencing the endergonic
action. Here again, anorganic phosphate is
process. For coupling, the two reactions have
introduced into a mixed acid anhydride
to be linked in such a way that a common
bond to be transferred from there to GDP.
intermediate arises. This connection is illus-
Succinyl phosphate is only an intermediate
trated here using the example of the gluta-
here, and is not released by the enzyme.
mine synthetase reaction.
In the literature, the term “substrate level
Direct transfer of NH3 to glutamate is en-
phosphorylation” is used inconsistently.
dergonic
(∆G0 = +14 kJ mol-1; see p. 18),
Some authors use it to refer to reactions in
and can therefore not take place. In the cell,
which anorganic phosphate is raised to a high
the reaction is divided into two exergonic
potential, while others use it for the subse-
steps. First, the γ-phophate residue is trans-
quent reactions, in which ATP or GTP is
ferred from ATP to glutamate. This gives rise
formed from the energy-rich intermediates.
to an “energy-rich” mixed acid anhydride. In
the second step, the phosphate residue from
C. Glyceraldehyde-3-phosphate
the intermediate is substituted by NH3, and
dehydrogenase
glutamine and free phosphate are produced.
The energy balance of the reaction as a whole
The reaction catalyzed during glycolysis by
(∆G0 = -17 kJ mol-1) is the sum of the
glyceraldehyde-3-phosphate dehydrogenase
changes in free enthalpy of direct glutamine
(GADPH) is shown here in detail. Initially,
synthesis (∆G0 = 14 kJ mol-1) plus ATP hy-
the SH group of a cysteine residue of the
drolysis (∆G0
= -31 kJ mol-1), although ATP
enzyme is added to the carbonyl group of
has not been hydrolyzed at all.
glyceraldehyde 3-phosphate (a). This inter-
mediate is oxidized by NAD+ into an “en-
ergy-rich” thioester (b). In the third step (c),
B. Substrate-level phosphorylation
anorganic phosphate displaces the thiol, and
As mentioned earlier (see p. 122), there are a
the mixed anhydride 1,3-bisphosphoglycerate
few metabolites that transfer phosphate to
arises. In this bond, the phosphate residue is
ADP in an exergonic reaction and can there-
at a high enough potential for it to be trans-
fore form ATP. In ATP synthesis, anorganic
ferred to ADP in the next step (not shown; see
phosphate or phosphate bound in an ester-
p. 150).
like fashion is transferred to bonds with a
high phosphate transfer potential. Reactions
of this type are termed “substrate-level phos-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
125
A. Energetic coupling
COO
Glu-γ-
Gln
phosphate
COO
H
3
N
C H
P
NH3
H3N
C H
(CH
)
2 2
H2O
(CH )
C
2 2
O
NH2
C
∆GOI =
∆GOI = -17 kJ . mol-1
O
O
+14 kJ .
COO
HO
P
O
mol-1
NH4
H
3
N
C H
O
(CH )
2 2
A
A
P P
P
ATP
P P
ADP
C
O
O
–
Glu
1. Glutamine synthetase reaction
Reaction 1: Glutamate + NH3
+14 kJ . mol-1
Glutamine
+ H2O
Reaction 2: ATP
+ H2O
- 31 kJ . mol-1
ADP
+
P
Total:
Glutamate + NH3+ ATP
- 17 kJ . mol-1
Glutamine +
ADP
+
P
2. Energy balance
B. Substrate level phosphorylation
C. Glyceraldehyde 3-phosphate
dehydrogenase
H
Chemical
Phospho-
H
potential
enolpyruvate
S
Glyceraldehyde
70
C
P
3-phosphate
R
O
Glycer-
60
1,3-bisphospho-
a
aldehyde-
glycerate
3-phosphate
Succinyl-
dehydro-
P
phosphate
genase
H
50
P
Inter-
R
C
S
mediate 1
40
OH
N
A
P
A
b
P P P
30
ATP (GTP)
N
A
Inter-
N
A
mediate 2
R
C
S
2
CoA
20
1
3
O
Inorganic
Thioester
N
A
P
Succinyl-
P
phosphate
CoA
10
c
H
Glycer-
2-Phospho-
S
aldehyde
glycerate
3-phos-
1,3-Bisphos-
R
C
O P
phate
phoglycerate
Inorganic
P
P
O
phosphate
Mixed anhydride
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
126
Metabolism
constantly maintained by the enzyme Na+/
Energy conservation at membranes
K+-ATPase, which consumes ATP.
Metabolic energy can be stored not only in the
form of “energy-rich” bonds (see p. 122), but
B. Proton motive force
also by separating electric charges from each
other using an insulating layer to prevent
Hydronium ions (“H+ ions”) can also develop
them from redistributing. In the field of tech-
electrochemical gradients. Such a proton gra-
nology, this type of system would be called a
dient plays a decisive part in cellular ATP syn-
condenser. Using the same principle, energy is
thesis (see p. 142). As usual, the energy con-
also stored (“conserved”) at cell membranes.
tent of the gradient depends on the concen-
The membrane functions as an insulator;
tration gradients—i. e., on the pH difference
electrically charged atoms and molecules
∋pH between the two sides of the membrane.
(ions) function as charges.
In addition, the membrane potential ∆ψ also
makes a contribution. Together, these two
values give the proton motive force ∆p, a
A. Electrochemical gradient
measure for the work that the H+ gradient
Although artificial lipid membranes are al-
can do. The proton gradient across the inner
most impermeable to ions, biological mem-
mitochondrial membrane thus delivers ap-
branes contain ion channels that selectively
proximately 24 kJ per mol H+.
allow individual ion types to pass through
(see p. 222). Whether an ion can cross this
C. Energy conservation in proton gradients
type of membrane, and if so in which direc-
tion, depends on the electrochemical gra-
Proton gradients can be built up in various
dient—i. e., on the concentrations of the ion
ways. A very unusual type is represented by
on each side of the membrane (the concen-
bacteriorhodopsin (1), a light-driven proton
tration gradient) and on the difference in the
pump that various bacteria use to produce
electrical potential between the interior and
energy. As with rhodopsin in the eye, the
exterior, the membrane potential.
light-sensitive component used here is cova-
The membrane potential of resting cells
lently bound retinal (see p. 358). In photosyn-
(resting potential; see p. 350) is
-0.05 to
thesis
(see p. 130), reduced plastoquinone
-0.09 V—i. e., there is an excess negative
(QH2) transports protons, as well as electrons,
charge on the inner side of the plasma mem-
through the membrane (Q cycle, 2). The for-
brane. The main contributors to the resting
mation of the proton gradient by the respira-
potential are the two cations Na+ and K+, as
tory chain is also coupled to redox processes
well as Cl- and organic anions (1). Data on the
(see p. 140). In complex III, a Q cycle is respon-
concentrations of these ions outside and in-
sible for proton translocation (not shown). In
side animal cells, and permeability coef -
cytochrome c oxidase (complex IV, 3), H+ trans-
cients, are shown in the table (2).
port is coupled to electron flow from
The behavior of an ion type is described
cytochrome c to O2.
quantitatively by the Nernst equation
(3).
In each of these cases, the H+ gradient is
∆ψG is the membrane potential (in volts, V)
utilized by an ATP synthase (4) to form ATP.
at which there is no net transport of the ion
ATP synthases consist of two components—a
concerned across the membrane (equilibrium
proton channel (F0) and an inwardly directed
potential). The factor R T/F n has a value of
protein complex (F1), which conserves the
0.026 V for monovalent ions at 25 °C. Thus,
energy of back-flowing protons through ATP
for K+, the table (2) gives an equilibrium po-
synthesis (see p. 142).
tential of ca. -0.09 V—i. e., a value more or less
the same as that of the resting potential. By
contrast, for Na+ ions, ∆ψG is much higher than
the resting potential, at
+0.07 V. Na+ ions
therefore immediately flow into the cell
when Na+ channels open (see p. 350). The
disequilibrium between Na+ and K+ ions is
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
127
C. Energy conservation
A. Electrochemical gradient
in proton gradients
Inside
Outside
2H
2H
1. Light-driven
proton pump
Retinal
Na
K
Cl
Organic anions
(bacterio-
1. Cause
rhodopsin)
Ion
Concentrations
Permeability
coefficient
2e
PS II
Cyto-
Extracelluar
plasm space
Q
(mM)
(mM)
(cm · s-1 · 109)
2H
K
100
5
500
2. Q cycle
QH2
Na
15
150
5
(plasto-
quinone
Ca2
0.0002
2
cycle
in plants)
Cl
13
150
10
Q
Organic
138
34
0
anions
QH2
2H
2. Concentrations
b/f-
complex
2e
Cyto-
C
outside
R · T
chrome C
∆ΨG =
·
ln
F · n
C
inside
Fe
O2
CuA
R = gas constant
n = Ion charge
8H
T = temperature (K)
F = Faraday constant
Fe
4H
2 H2O
3. Nernst equation
CuB
3. Electron-
Heme/a3
B. Proton motive force
driven
proton
pump
(cyto-
chrome C
oxidase)
Membrane
pH gradient
F
F
1
0
potential
∆pH = pHa - pH
i
∆ψ = ψa - ψ
i
(in pH units)
nH
P
A
Protein
P P
Proton motive force
≈ 0.06 V
A
P P P
ATP
2.3 · R · T
∆p = ∆ψ -
·
∆pH
F
4. Proton-
∆G
= -F · ∆p
driven ATP
Proton flow
Ha H i
synthesis
Electron flow
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
128
Metabolism
koids or flattened membrane sacs are stacked
Photosynthesis: light reactions
on top of each other to form grana. The inside
Sunlight is the most important source of en-
of the thylakoid is referred to as the lumen.
ergy for nearly all living organisms. With the
The light reactions are catalyzed by enzymes
help of photosynthesis, light energy is used to
located in the thylakoid membrane, whereas
produce organic substances from CO2 and
the dark reactions take place in the stroma.
water. This property of phototrophic organ-
As in the respiratory chain (see p. 140), the
isms (plants, algae, and some bacteria) is ex-
light reactions cause electrons to pass from
ploited by heterotrophic organisms (e. g., ani-
one redox system to the next in an electron
mals), which are dependent on a supply of
transport chain. However, the direction of
organic substances in their diet (see p. 112).
transport is opposite to that found in the res-
The atmospheric oxygen that is vital to higher
piratory chain. In the respiratory chain, elec-
organisms is also derived from photosynthe-
trons flow from NADH+H+ to O2, with the
sis.
production of water and energy.
In photosynthesis, electrons are taken up
from water and transferred to NADP+, with an
A. Photosynthesis: overview
expenditure of energy. Photosynthetic electron
The chemical balance of photosynthesis is
transport is therefore energetically
“uphill
simple. Six molecules of CO2
are used to
work.” To make this possible, the transport
form one hexose molecule (right). The hydro-
is stimulated at two points by the absorption
gen required for this reduction process is
of light energy. This occurs through two pho-
taken from water, and molecular oxygen is
tosystems—protein complexes that contain
formed as a by-product (left). Light energy is
large numbers of chlorophyll molecules and
required, since water is a very poor reducing
other pigments (see p. 132). Another compo-
agent and is therefore not capable of reducing
nent of the transport chain is the cytochrome
CO2.
b/f complex, an aggregate of integral mem-
In the light-dependent part of photosyn-
brane proteins that includes two cytochromes
thesis—the “light reactions ”—H2O molecules
(b563
and f). Plastoquinone, which is com-
are split into protons, electrons, and oxygen
parable to ubiquinone, and two soluble pro-
atoms. The electrons undergo excitation by
teins, the coppercontaining plastocyanin and
light energy and are raised to an energy level
ferredoxin, function as mobile electron car-
that is high enough to reduce NADP+. The
riers. At the end of the chain, there is an
NADPH+H+ formed in this way, in contrast to
enzyme that transfers the electrons to NADP+.
H2O, is capable of “fixing” CO2
reductive-
Because photosystem II and the cyto-
ly—i. e., of incorporating it into organic bonds.
chrome b/f complex release protons from re-
Another product of the light reactions is ATP,
duced plastoquinone into the lumen (via a Q
which is also required for CO2 fixation. If
cycle), photosynthetic electron transport es-
NADPH+H+, ATP, and the appropriate en-
tablishes an electrochemical gradient across
zymes are available, CO2 fixation can also
the thylakoid membrane (see p. 126), which is
take place in darkness. This process is there-
used for ATP synthesis by an ATP synthase.
fore known as the “dark reaction.”
ATP and NADPH+H+, which are both needed
The excitation of electrons to form NADPH
for the dark reactions, are formed in the
is a complex photochemical process that
stroma.
involves chlorophyll, a tetrapyrrole dye con-
taining Mg2+ that bears an extra phytol resi-
due (see p. 132).
B. Light reactions
In green algae and higher plants, photosyn-
thesis occurs in chloroplasts. These are organ-
elles, which—like mitochondria—are sur-
rounded by two membranes and contain their
own DNA. In their interior, the stroma, thyla-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
129
A. Photosynthesis: overview
Phytol residue
Hexose
N
A
P
H
2
CH
12 NADPH + H
H3C
CH
H
N
CH3
O
H3C
N
Mg
N
A
C
P P P
O
H
N
“Dark
CH
2
CH3
H
2+
18 ATP
reactions”
Mg
H
O
C
(Calvin
CH3
O
cycle)
O
A
H3C
P P
P
Chlorophyll a
3 O2
12 H
18 ADP + Pi
N
A
Electron flow
P
6 H2O
Proton flow
12 NADP
6 CO2
Cytoplasm
Nucleus
Outer
Thylakoid
Lumen
membrane
Inner
membrane
Plasma
membrane
Stroma
Granum
Chloroplast
Vacuole
B. Light reactions
A
nH
A
P P
P P P
N
A
P
ATP
P
Stroma
2
NADPH + H
Thylakoid membrane
N
A
1
P
Lumen
NADP
Ferredoxin (Fd)
2 H
2 H
H2O
[O]
Mg2+
Light
Photosystem I
Plasto-
quinone
Ferredoxin-
1
Plasto-
NADP
-
Mg
2+
Q
cyanin
reductase
1.18.1.2
QH2
(PC)
ATP synthase
2
3.6.1.34
Photosystem II
2 H
Cytochrome b/f complex
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
130
Metabolism
quinone pool takes place as described on the
Photosynthesis: dark reactions
preceding page and shown in B.
The “light reactions” in photosynthesis bring
about two strongly endergonic reactions—the
B. Redox series
reduction of NADP+ to NADPH+H+ and ATP
synthesis (see p. 122). The chemical energy
It can be seen from the normal potentials E0
needed for this is produced from radiant en-
(see p. 18) of the most important redox sys-
ergy by two photosystems.
tems involved in the light reactions why two
excitation processes are needed in order to
transfer electrons from H2O to NADP+. After
A. Photosystem II
excitation in PS II, E0 rises from around -1 V
The photosynthetic electron transport chain
back to positive values in plastocyanin
in plants starts in photosystem II (PS II; see
(PC)—i. e., the energy of the electrons has to
p. 128). PS II consists of numerous protein
be increased again in PS I. If there is no NADP+
subunits
(brown) that contain bound pig-
available, photosynthetic electron transport
ments—i. e., dye molecules that are involved
can still be used for ATP synthesis. During
in the absorption and transfer of light energy.
cyclic photophosphorylation, electrons return
The schematic overview of PS II presented
from ferredoxin (Fd) via the plastoquinone
here (1) only shows the important pigments.
pool to the b/f complex. This type of electron
These include a special chlorophyll molecule,
transport does not produce any NADPH, but
the reaction center P680; a neighboring Mg2+
does lead to the formation of an H+ gradient
free chlorophyll (pheophytin); and two bound
and thus to ATP synthesis.
plastoquinones (QA and QB). A third quinone
(QP) is not linked to PS II, but belongs to the
C. Calvin cycle
plastoquinone pool. The white arrows indi-
cate the direction of electron flow from water
The synthesis of hexoses from CO2 is only
to QP. Only about 1 % of the chlorophyll mol-
shown in a very simplified form here; a com-
ecules in PS II are directly involved in photo-
plete reaction scheme is given on p. 407. The
chemical excitation (see p. 128). Most of them
actual CO2 fixation—i. e., the incorporation of
are found, along with other pigments, in what
CO2 into an organic compound—is catalyzed
are known as light-harvesting or antenna
by ribulose bisphosphate carboxylase/oxygen-
complexes (green). The energy of light quanta
ase (“rubisco”). Rubisco, the most abundant
striking these can be passed on to the reaction
enzyme on Earth, converts ribulose 1,5-bis-
center, where it can be utilized.
phosphate, CO2 and water into two mole-
In Fig. 2, photosynthetic electron transport
cules of 3-phosphoglycerate. These are then
in PS II is separated into the individual steps
converted, via 1,3-bisphosphoglycerate and
involved. Light energy from the light-harvest-
3-phosphoglycerate, into glyceraldehyde
ing complexes (a) raises an electron of the
3-phosphate (glyceral 3-phosphate). In this
chlorophyll in the reaction center to an
way, 12 glyceraldehyde 3-phosphates are syn-
excited “singlet state.” The excited electron is
thesized from six CO2. Two molecules of this
immediately passed on to the neighboring
intermediate are used by gluconeogenesis re-
pheophytin. This leaves behind an “electron
actions to synthesize glucose
6-phosphate
gap” in the reaction center—i. e., a positively
(bottom right). From the remaining 10 mole-
charged P680 radical (b). This gap is now filled
cules, six molecules of ribulose 1,5-bisphos-
by an electron removed from an H2O mole-
phate are regenerated, and the cycle then
cule by the water-splitting enzyme (b). The
starts over again. In the Calvin cycle, ATP is
excited electron passes on from the pheophy-
required for phosphorylation of 3-phospho-
tin via QA to QB, converting the latter into a
glycerate
and ribulose
5-phosphate.
semiquinone radical (c). QB is then reduced to
NADPH+H+, the second product of the light
hydroquinone by a second excited electron,
reaction, is consumed in the reduction of 1,3-
and is then exchanged for an oxidized qui-
bisphosphoglycerate to glyceraldehyde
3-
none (QP) from the plastoquinone pool. Fur-
phosphate.
ther transport of electrons from the plasto-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
131
A. Photosystem II
B. Redox series
EOl (V)
Photo-
Bound
Exchangeable Plasto-
chemical
-1.2
quinone
quinone
quinone
excitation
Q
-1.0
Fe
-0.8
Fe/S
Ph
Mg2+
Mg2+
Mg2+
QA
Q
Q
-0.6
B
P
QA
Fd
Mg2+
Mg2+
Mg2+
-0.4
QB
H
N
A
QP
-0.2
P
b/f
Light-
Reaction
NADP
0
harvesting
Mg2+
center
PC
complex
(P680)
+0.2
PS I
Pheophytin
+0.4
(P
PS II
700)
(P
680)
+0.6
Cyclic
Water-
+0.8
photophos-
splitting
4 Mn2+
H2O
phorylation
enzyme
H2O
4 ions
2 H
1/2 O2
1.
Plasto-
a
b
c
d
quinone
OH
(ox.)
OH
Mg2+
Mg2+
Mg2+
Mg2+
Exchange
Q
QP(ox)
B (red)
2.
H2O
C. Calvin cycle
A
A
6 ADP
P
P
P
P
P
6 ATP
H
H O O
P
H2C C
C C CH2
4
P
O OH OH
6 CO2
H2O
O C O
CO2, Mg2
1
CO2
Light
1
6 H2O
4 P
OH
O
P
Light
O
O
H2C C
C
C
C
CH2
Light
P O OHO
O
Ribulose-bisphosphate
12
P
1
carboxylase 4.1.1.39
2
A
3
Phosphoglycerate kinase
P
P
P
Glucose
2
6-phosphate
2.7.2.3
12 ATP
A
P P
Glyceraldehyde 3-phosphate
3
N
A
dehydrogenase (NADP
) 1.2.1.13
12 ADP
Gluco-
N
A
P
neogenesis
P
4
Phosphoribulokinase 2.7.1.19
12 NADPH
12 NADP
P
Glyceraldehyde
2
3-phosphate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
132
Metabolism
are taken up by two channels (D and K, not
Molecular models:
shown). The mechanism that links proton
membrane proteins
transport to electron transfer is still being
investigated.
The plates show, in simplified form, the struc-
tures of cytochrome c oxidase (A; complex IV
of the respiratory chain) and of photosystem I
B. Reaction center of Synechococcus
of a cyanobacterium (B). These two molecules
elongatus
are among the few integral membrane pro-
Photosystem I (PS I) in the cyanobacterium
teins for which the structure is known in de-
Synechococcus elongatus is the first system of
tail. Both structures were determined by X-
this type for which the structure has been
ray crystallography.
solved in atomic detail. Although the bacterial
photosystem differs slightly from the systems
in higher plants, the structure provides val-
A. Cytochrome c oxidase
uable hints about the course of the light re-
The enzyme cytochrome c oxidase (“COX,” EC
actions in photosynthesis (see p. 128). The
1.9.3.1) catalyzes the final step of the respira-
functioning of the photosystem is discussed
tory chain. It receives electrons from the small
in greater detail on p. 130.
heme protein cytochrome c and transfers
The functional form of PS I in S. elongatus
them to molecular oxygen, which is thereby
consists of a trimer with a mass of more than
reduced to water (see p. 140). At the same
106 Da that is integrated into the membrane.
time, 2-4 protons per water molecule formed
Only one of the three subunits is shown here.
are pumped from the matrix into the inter-
This consists of
12 different polypeptides
membrane space.
(gray-blue), 96 chlorophyll molecules (green),
Mammalian COX (the illustration shows
22 carotenoids (orange), several phylloqui-
the enzyme from bovine heart) is a dimer
nones (yellow), and other components. Most
that has two identical subunits with masses
of the chlorophyll molecules are so-called an-
of 204 kDa each. Only one subunit is shown in
tenna pigments. These collect light energy
detail here; the other is indicated by gray
and conduct it to the reaction center, which
lines. Each subunit consists of 13 different
is located in the center of the structure and
polypeptides, which all span the inner mito-
therefore not visible. In the reaction center, an
chondrial membrane. Only polypeptides I
electron is excited and transferred via various
(light blue) and II (dark blue) and the linked
intermediate steps to a ferredoxin molecule
cofactors are involved in electron transport.
(see p. 128). The chlorophylls (see formula)
The other chains, which are differently ex-
are heme-like pigments with a highly modi-
pressed in the different organs, probably
fied tetrapyrrole ring, a central Mg2+ ion, and
have regulatory functions. The two heme
an apolar phytol side chain. Shown here is
groups, heme a (orange) and heme a1 (red)
chlorophyll a, which is also found in the re-
are bound in polypeptide 1. The copper center
action center of the S. elongatus photosystem.
CuA consists of two copper ions
(green),
The yellow and orange-colored carot-
which are coordinated by amino acid residues
enoids—e. g., E-carotene
(see formula)—are
in polypeptide II. The second copper (CuB) is
auxiliary pigments that serve to protect the
located in polypeptide I near heme a3.
chloroplasts from oxidative damage. Danger-
To reduce an O2 molecule to two molecules
ous radicals can be produced during the light
of H2O, a total of four electrons are needed,
reaction—particularly singlet oxygen. Caroten-
which are supplied by cytochrome c (pink, top
oids prevent compounds of this type from
left) and initially given off to CuA. From there,
arising, or render them inactive. Carotenoids
they are passed on via heme a and heme a3 to
are also responsible for the coloring of leaves
the enzyme’s reaction center, which is located
seen during fall. They are left behind when
between heme a3 and CuB. The reduction of
plants break down chlorophyll in order to
the oxygen takes place in several steps, with-
recover the nitrogen it contains.
out any intermediate being released. The four
protons needed to produce water and the H+
ions pumped into the intermembrane space
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
133
A. Cytochrome C oxidase
Outer
mitochondrial
membrane
Intermembrane space
Cytochrome C
Cu A
Cu B
Heme a3
Inner
Matrix
Heme a
mitochondrial
membrane
B. Photosystem I
O
H3C
H3C
O
O
H
CH3
H3C
C
CH
Plastoquinone
O
H
N
N
CH2
H
H3CO
Mg2+
C
N
N
OO
CH3
CH3
CH3
Chlorophyll
β-carotene
Thylakoid membrane
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
134
Metabolism
An important aspect of PDH catalysis is the
Oxoacid dehydrogenases
spatial relationship between the components
The intermediary metabolism has multien-
of the complex. The covalently bound lipo-
zyme complexes which, in a complex reaction,
amide coenzyme is part of a mobile domain
catalyze the oxidative decarboxylation of 2-
of E2, and is therefore highly mobile. This
oxoacids and the transfer to coenzyme A of
structure is known as the lipoamide arm, and
the acyl residue produced. NAD+ acts as the
swings back and forth between E1 and E3
electron acceptor. In addition, thiamine di-
during catalysis. In this way, lipoamide can
phosphate, lipoamide, and FAD are also in-
interact with the TPP bound at E1, with solute
volved in the reaction. The oxoacid
coenzyme A, and also with the FAD that
dehydrogenases include a) the pyruvate dehy-
serves as the electron acceptor in E3.
drogenase complex (PDH, pyruvate
acetyl
CoA), b) the
2-oxoglutarate dehydrogenase
B. PDH complex of Escherichia coli
complex of the tricarboxylic acid cycle (ODH,
2-oxoglutarate
succinyl CoA), and c) the
The PDH complex of the bacterium Escheri-
branched chain dehydrogenase complex, which
chia coli has been particularly well studied. It
is involved in the catabolism of valine, leu-
has a molecular mass of 5.3
106, and with a
cine, and isoleucine (see p. 414).
diameter of more than 30 nm it is larger than
a ribosome. The complex consists of a total of
60 polypeptides (1, 2): 24 molecules of E2
A. Pyruvate dehydrogenase: reactions
(eight trimers) form the almost cube-shaped
The pyruvate dehydrogenase reaction takes
core of the complex. Each of the six surfaces of
place in the mitochondrial matrix
(see
the cube is occupied by a dimer of E3 compo-
p. 210). Three different enzymes
[E1-E3]
nents, while each of the twelve edges of the
form the PDH multienzyme complex (see B).
cube is occupied by dimers of E1 molecules.
[1] Initially, pyruvate dehydrogenase [E1]
Animal oxoacid dehydrogenases have similar
catalyzes the decarboxylation of pyruvate
structures, but differ in the numbers of sub-
and the transfer of the resulting hydroxyethyl
units and their molecular masses.
residue to thiamine diphosphate (TPP, 1a). The
same enzyme then catalyzes oxidation of the
Further information
TPP-bound hydroxyethyl group to yield an
acetyl residue. This residue and the reducing
The PDH reaction, which is practically irrever-
equivalents obtained are then transferred to
sible, occupies a strategic position at the inter-
lipoamide (1b).
face between carbohydrate and fatty acid me-
[2] The second enzyme, dihydrolipoamide
tabolism, and also supplies acetyl residues to
acetyltransferase [E2], shifts the acetyl residue
the tricarboxylic acid cycle. PDH activity is
from lipoamide to coenzyme A (2), with dihy-
therefore strictly regulated (see p. 144). Inter-
drolipoamide being left over.
conversion is particularly important in animal
[3] The third enzyme, dihydrolipoamide de-
cells (see p. 120). Several PDH-specific protein
hydrogenase
[E3], reoxidizes dihydrolipo-
kinases inactivate the E1 components through
amide, with NADH+H+ being formed. The
phosphorylation, while equally specific pro-
electrons are first taken over by enzyme-
tein phosphatases reactivate it again. The
bound FAD (3a) and then transferred via a
binding of the kinases and phosphatases to
catalytically active disulfide bond in the E3
the complex is in turn regulated by metabo-
subunit (not shown) to soluble NAD+ (3b).
lites. For example, high concentrations of ace-
The five different coenzymes involved are
tyl CoA promote binding of kinases and
associated with the enzyme components in
thereby inhibit the reaction, while Ca2+ in-
different ways. Thiamine diphosphate is
creases the activity of the phosphatase. Insu-
non-covalently bound to E1, whereas lipo-
lin activates PDH via inhibition of phosphor-
amide is covalently bound to a lysine residue
ylation.
of E2 and FAD is bound as a prosthetic group to
E3. NAD+ and coenzyme A, being soluble
coenzymes, are only temporarily associated
with the complex.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
135
A. Pyruvate dehydrogenase: reactions
E1
CO2
R
OH
N
Pyruvate
H3C
Hydroxyalkyl thiamine
C
C
CH3
O
diphosphate
S
H
OOC
C CH3
R
1
E2
2
O
S
Acetyl
R1
S
S C CH3
lipoamide
N
S
H3C
H
H
S
S
A
R2
Thiamine
3
CoA
diphosphate
TPP
O
S
S
C CH3
A
S
S
H
S
Acetyl CoA
H
E3
4a
Dihydrolipoamide
F
A
F
A
4b
Pyruvate dehydro-
E1
genase 1.2.4.1
Dihydrolipoamide
E2
acetyltransferase 2.3.1.12
N
A
N
A
E3
Dihydrolipoamide
dehydrogenase 1.8.1.4
NAD
NADH + H
B. PDH complex of Escherichia coli
10nm
E1
12 · 2 = 24
E1
E3
E2
8 · 3 = 24
E2
E3
6 · 2 =12
E1+ E2 + E3 = 60
1.
2.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
136
Metabolism
p. 134). NADH+H+ is once again formed in
Tricarboxylic acid cycle: reactions
this reaction.
The tricarboxylic acid cycle (TCA cycle, also
[5] The subsequent cleavage of the thio-
known as the citric acid cycle or Krebs cycle)
ester succinylCoA into succinate and coen-
is a cyclic metabolic pathway in the mitochon-
zyme A by succinic acid-CoA ligase (succinyl
drial matrix (see p. 210). In eight steps, it oxi-
CoA synthetase, succinic thiokinase) is
dizes acetyl residues (CH3-CO-) to carbon di-
strongly exergonic and is used to synthesize
oxide (CO2). The reducing equivalents ob-
a phosphoric acid anhydride bond (“substrate
tained in this process are transferred to
level phosphorylation ”, see p. 124). However, it
NAD+ or ubiquinone, and from there to the
is not ATP that is produced here as is other-
respiratory chain (see p. 140). Additional met-
wise usually the case, but instead guanosine
abolic functions of the cycle are discussed on
triphosphate (GTP). However, GTP can be con-
p. 138.
verted into ATP by a nucleoside diphosphate
kinase (not shown).
[6] Via the reactions described so far, the
A. Tricarboxylic acid cycle
acetyl residue has been completely oxidized
The acetyl-CoA that supplies the cycle with
to CO2. At the same time, however, the carrier
acetyl residues is mainly derived from E-
molecule oxaloacetate has been reduced to
oxidation of fatty acids (see p. 164) and from
succinate. Three further reactions in the cycle
the pyruvate dehydrogenase reaction. Both of
now regenerate oxaloacetate from succinate.
these processes take place in the mitochon-
Initially, succinate dehydrogenase oxidizes
drial matrix.
succinate to fumarate. In contrast to the other
[1] In the first step of the cycle, citrate
enzymes in the cycle, succinate dehydrogenase
synthase catalyzes the transfer of an acetyl
is an integral protein of the inner mitochondrial
residue from acetyl CoA to a carrier molecule,
membrane. It is therefore also assigned to the
oxaloacetic acid. The product of this reaction,
respiratory chain as complex II. Although suc-
tricarboxylic acid, gives the cycle its name.
cinate dehydrogenase contains FAD as a pros-
[2] In the next step, tricarboxylic acid
thetic group, ubiquinone is the real electron
undergoes isomerization to yield isocitrate.
acceptor of the reaction.
In the process, only the hydroxyl group is
[7] Water is now added to the double bond
shifted within the molecule. The correspond-
of fumarate by fumarate hydratase (“fuma-
ing enzyme is called aconitate hydratase
rase”), and chiral (2S)-malate is produced.
(“aconitase”), because unsaturated aconitate
[8] In the last step of the cycle, malate is
arises as an enzyme-bound intermediate dur-
again oxidized by malate dehydrogenase into
ing the reaction (not shown; see p. 8). Due to
oxaloacetate, with NADH+H+ again being pro-
the properties of aconitase, the isomerization
duced. With this reaction, the cycle is com-
is absolutely stereospecific. Although citrate is
plete and can start again from the beginning.
not chiral, isocitrate has two chiral centers, so
As the equilibrium of the reaction lies well on
that it could potentially appear in four iso-
the side of malate, the formation of oxaloace-
meric forms. However, in the tricarboxylic
tic acid by reaction
[8] depends on the
acid cycle, only one of these stereoisomers,
strongly exergonic reaction [1], which imme-
(2R,3S)-isocitrate, is produced.
diately removes it from the equilibrium.
[3] The first oxidative step now follows.
The net outcome is that each rotation of
Isocitrate dehydrogenase oxidizes the hy-
the tricarboxylic acid cycle converts one ace-
droxyl group of isocitrate into an oxo group.
tyl residue and two molecules of H2O into two
At the same time, a carboxyl group is released
molecules of CO2. At the same time, one GTP,
as CO2, and 2-oxoglutarate (also known as α-
three NADH+H+ and one reduced ubiquinone
ketoglutarate) and NADH+H+ are formed.
(QH2) are produced. By oxidative phosphory-
[4] The next step, the formation of succinyl
lation (see p. 122), the cell obtains around
CoA, also involves one oxidation and one de-
nine molecules of ATP from these reduced
carboxylation. It is catalyzed by 2-oxogluta-
coenzymes (see p. 146). Together with the
rate dehydrogenase, a multienzyme complex
directly formed GTP, this yields a total of 10
closely resembling the PDH complex
(see
ATP per acetyl group.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
137
A. Tricarboxylic acid cycle
Center of chirality
COO
Acetyl CoA
H
C
OH
H
C H
7
8
+28.1
S
C CH3
H2O
-3.8
COO
A
O
∆G0I
(2S) - Malate
COO
kJ · mol-1
COO
H
C
C
O
N
A
N
A
C H
CH2
COO
COO
Fumarate
Oxaloacetate
OH
Ubiquinol
QH2
H2O
6
±0
1
-38.2
OH
S
O
A
Respiratory
OOC
COO
COO
chain
H2C
C
OH
CH
O
2
CH2
CH2
COO
COO
Citrate
Succinate
G
P P
P
GTP
N
A
N
A
5
-8.8
2
+6.7
G
N
A
N
A
P P
OOC
OOC
COO
P
H2C
CH2
H2C
C
H
S
C O
H
C
OH
A
COO
Succinyl CoA
(2R,3S)-Isocitrate
S
OOC
A
-37.0
-7.1
H2C
CH2
4
3
C
O
O C O
CO2
O C O
CO2
COO
2-Oxoglutarate
Citrate synthase
2-Oxoglutarate DH complex
Succinate DH 1.3.5.1
1
4
6
4.1.3.7
1.2.4.2, 1.8.1.4, 2.3.1.61
[FAD, Fe2S2, Fe4S4]
Aconitase
Succinate-CoA ligase
Fumarate hydratase
2
5
7
4.2.1.3 [Fe4S4]
6.2.1.4
4.2.1.2
3
Isocitrate DH 1.1.1.41
DH = dehydrogenase
8
Malate DH 1.1.1.37
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
138
Metabolism
carboxylase [1]. It allows pyruvate yielding
Tricarboxylic acid cycle: functions
amino acids and lactate to be used for gluco-
neogenesis.
A. Tricarboxylic acid cycle: functions
By contrast, acetyl CoA does not have ana-
The tricarboxylic acid cycle (see p. 136) is
plerotic effects in animal metabolism. Its car-
often described as the “hub of intermediary
bon skeleton is completely oxidized to CO2
metabolism.” It has both catabolic and ana-
and is therefore no longer available for bio-
bolic functions—it is amphibolic.
synthesis. Since fatty acid degradation only
As a catabolic pathway, it initiates the “ter-
supplies acetyl CoA, animals are unable to
minal oxidation” of energy substrates. Many
convert fatty acids into glucose. During peri-
catabolic pathways lead to intermediates of
ods of hunger, it is therefore not the fat re-
the tricarboxylic acid cycle, or supply metab-
serves that are initially drawn on, but pro-
olites such as pyruvate and acetyl-CoA that
teins. In contrast to fatty acids, the amino
can enter the cycle, where their C atoms are
acids released are able to maintain the blood
oxidized to CO2. The reducing equivalents
glucose level (see p. 308).
(see p. 14) obtained in this way are then
used for oxidative phosphorylation—i. e., to
The tricarboxylic acid cycle not only takes
aerobically synthesize ATP (see p. 122).
up acetyl CoA from fatty acid degradation, but
also supplies the material for the biosynthesis
The tricarboxylic acid cycle also supplies
of fatty acids and isoprenoids. Acetyl CoA,
important precursors for anabolic pathways.
which is formed in the matrix space of mito-
Intermediates in the cycle are converted into:
chondria by pyruvate dehydrogenase
(see
p. 134), is not capable of passing through the
• Glucose (gluconeogenesis; precursors: oxa-
inner mitochondrial membrane. The acetyl
loacetate and malate—see p. 154)
residue is therefore condensed with oxalo-
• Porphyrins
(precursor: succinyl-CoA—see
acetate by mitochondrial citrate synthase to
p. 192)
form citrate. This then leaves the mitochon-
• Amino acids (precursors: 2-oxoglutarate,
dria by antiport with malate
(right; see
oxaloacetate—see p. 184)
p. 212). In the cytoplasm, it is cleaved again
• Fatty acids and isoprenoids (precursor: cit-
by ATP-dependent citrate lyase [4] into acetyl-
rate—see below)
CoA and oxaloacetate. The oxaloacetate
The intermediates of the tricarboxylic acid
formed is reduced by a cytoplasmic malate
cycle are present in the mitochondria only in
dehydrogenase to malate [2], which then re-
very small quantities. After the oxidation of
turns to the mitochondrion via the antiport
acetyl-CoA to CO2, they are constantly regen-
already mentioned. Alternatively, the malate
erated, and their concentrations therefore re-
can be oxidized by “malic enzyme” [5], with
main constant, averaged over time. Anabolic
decarboxylation, to pyruvate. The NADPH+H+
pathways, which remove intermediates of the
formed in this process is also used for fatty
cycle (e. g., gluconeogenesis) would quickly
acid biosynthesis.
use up the small quantities present in the
mitochondria if metabolites did not reenter
Additional information
the cycle at other sites to replace the com-
pounds consumed. Processes that replenish
Using the so-called glyoxylic acid cycle, plants
the cycle in this way are called anaplerotic
and bacteria are able to convert acetyl-CoA
reactions.
into succinate, which then enters the tricar-
The degradation of most amino acids is
boxylic acid cycle. For these organisms, fat
anaplerotic, because it produces either inter-
degradation therefore functions as an ana-
mediates of the cycle or pyruvate (glucogenic
plerotic process. In plants, this pathway is
amino acids; see p. 180). Gluconeogenesis is in
located in special organelles, the glyoxysomes.
fact largely sustained by the degradation of
amino acids. A particularly important ana-
plerotic step in animal metabolism leads
from pyruvate to oxaloacetic acid. This ATP-
dependent reaction is catalyzed by pyruvate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
139
A. Tricarboxylic acid cycle: functions
* Essential
Asp, Asn,
Glucose
* amino acid
Arg, Met*,
GTP
Thr*, Ile*
Fat
Oxaloacetate
3
PEP
Phe*, Tyr, Trp*
2
Irreversible
Ala, Ser, Thr*,
Cys, Gly
Acyl-
Ala, Leu*,
Carbo-
Malate
Pyruvate
CoA
Val*
hydrates
CO2
Cytoplasm
N
A
P
Carnitine
5
shuttle
N
A
CO2
Pyruvate
P
Irreversible
ATP
CO
Oxalo-
2
Malate
2
acetate
Acyl
Acetyl
CoA
CoA
β-Oxidation
1
Mitochondrial
Malate
2
ATP
ADP+P
i
matrix
Oxalo-
acetate
Citrate
4
Fumarate
Asp
Asp
Phe*
Asn
Citrate
S
Tyr
A
His
CoA
Acetyl
Ile*
Pro
CoA
Val*
Arg
Succinate
Met*
Gln
Trp*
Glu
Isocitrate
Fatty
GTP
acids
Succinyl
2-Oxo-
CoA
CO2
glutarate
Fat
CO2
Glu, Gln,
Porphyrins
Pro, Arg
Pyruvate carboxylase
1
6.4.1.1
OH
Malate dehydrogenase
2
1.1.1.37
N
A
NADH+H
2[H]
O2
2[H]
Ubiquinol
PEP carboxykinase
(QH2)
3
4.1.1.32
OH
4
Citrate lyase 4.1.3.8
A
Oxidative
A
P P
+
P
phosphorylation
P P P
5
Malic enzyme 1.1.1.40
ADP
P
ATP
Catabolic pathway
i
H2O
Anabolic pathway
Anaplerotic reaction
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
140
Metabolism
zyme-bound FADH2 and the electron-trans-
Respiratory chain
porting flavoprotein (ETF; see p. 164). Ubiq-
The respiratory chain is one of the pathways
uinol passes electrons on to complex III, which
involved in oxidative phosphorylation
(see
transfers them via two b-type heme groups,
p. 122). It catalyzes the steps by which elec-
one Fe/S cluster, and heme c1 to the small
trons are transported from NADH+H+ or re-
heme protein cytochrome c. Cytochrome c
duced ubiquinone (QH2) to molecular oxygen.
then transports the electrons to complex
Due to the wide difference between the redox
IV—cytochrome c oxidase. Cytochrome c oxi-
potentials of the donor (NADH+H+ or QH2)
dase contains redox-active components in the
and the acceptor (O2), this reaction is strongly
form of two copper centers (CuA and CuB) and
exergonic (see p. 18). Most of the energy re-
hemes a and a3, through which the electrons
leased is used to establish a proton gradient
finally reach oxygen (see p. 132). As the result
across the inner mitochondrial membrane
of the two-electron reduction of O2, the
(see p. 126), which is then ultimately used to
strongly basic O2- anion is produced (at least
synthesize ATP with the help of ATP synthase.
formally), and this is converted into water by
binding of two protons. The electron transfer
is coupled to the formation of a proton gradi-
A. Components of the respiratory chain
ent by complexes I, III, and IV (see p. 126).
The electron transport chain consists of three
protein complexes (complexes I, III, and IV),
B. Organization
which are integrated into the inner mitochon-
drial membrane, and two mobile carrier mol-
Proton transport via complexes I, III, and IV
ecules—ubiquinone (coenzyme Q) and cyto-
takes place vectorially from the matrix into
chrome c. Succinate dehydrogenase, which ac-
the intermembrane space. When electrons
tually belongs to the tricarboxylic acid cycle,
are being transported through the respiratory
is also assigned to the respiratory chain as
chain, the H+ concentration in this space in-
complex II. ATP synthase (see p. 142) is some-
creases—i. e., the pH value there is reduced by
times referred to as complex V, although it is
about one pH unit. For each H2O molecule
not involved in electron transport. With the
formed, around 10 H+ ions are pumped into
exception of complex I, detailed structural in-
the intermembrane space. If the inner mem-
formation is now available for every complex
brane is intact, then generally only ATP syn-
of the respiratory chain.
thase (see p. 142) can allow protons to flow
All of the complexes in the respiratory
back into the matrix. This is the basis for the
chain are made up of numerous polypeptides
coupling of electron transport to ATP synthe-
and contain a series of different protein
sis, which is important for regulation pur-
bound redox coenzymes (see pp. 104, 106).
poses (see p. 144).
These include flavins (FMN or FAD in com-
As mentioned, although complexes I
plexes I and II), iron-sulfur clusters (in I, II,
through V are all integrated into the inner
and III), and heme groups (in II, III, and IV).
membrane of the mitochondrion, they are not
Of the more than 80 polypeptides in the res-
usually in contact with one another, since the
piratory chain, only 13 are coded by the mi-
electrons are transferred by ubiquinone and
tochondrial genome (see p. 210). The remain-
cytochrome c. With its long apolar side chain,
der are encoded by nuclear genes, and have to
ubiquinone is freely mobile within the mem-
be imported into the mitochondria after
brane. Cytochrome c is water-soluble and is
being synthesized in the cytoplasm
(see
located on the outside of the inner membrane.
p. 228).
NADH oxidation via complex I takes place
Electrons enter the respiratory chain in var-
on the inside of the membrane—i. e., in the
ious different ways. In the oxidation of
matrix space, where the tricarboxylic acid
NADH+H+ by complex I, electrons pass via
cycle and β-oxidation (the most important
FMN and Fe/S clusters to ubiquinone (Q). Elec-
sources of NADH) are also located. O2 reduc-
trons arising during the oxidation of succinate,
tion and ATP formation also take place in the
acyl CoA, and other substrates are passed to
matrix.
ubiquinone by succinate dehydrogenase or
other mitochondrial dehydrogenases via en-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
141
A. Components of the respiratory chain
Liponamide-H
Pyruvate
2
Complex I
EOI
2-Oxoglutarate
HS SH
(V
H
NADH dehydrogenase (ubiquinone)
1.6.5.3
Mr 700 - 800 kDa, 25 - 30 subunits
I
-0.3
1 FMN, 2 Fe2S2, 4 - 5 Fe4S
4
Succinate
Complex II
3-Hydroxybutyrate
3-Hydroxyacyl CoA
N
A
Succinate dehydrogenase 1.3.5.1
Malate
Isocitrate
Mr 125 kDa, 4 - 6 subunits
NADH
Q
II
+0.1
1 FAD, 1 Fe2S2, 1 Fe4S4, 1Fe3S
4
2 ubiquinone, 1 heme b
Complex III
H
Fumarate
Ubiquinol-cytochrome C
Acyl CoA
reductase 1.10.2.2
III
α-Glycerophosphate
ETF
Mr ≈ 400 kDa, 11 subunits
Dihydroorotat
2 Fe2S2, 2 heme b, 1 heme c
Cholin
1
Complex IV
Cytochrome C
Mr 12 kDa
Cytochrome Coxidase 1.9.3.1
nH
1 heme
+0.3
Mr ≈ 200 kDa, 8 - 13 subunits
2 Cu, 1 Zn, 1 heme a, 1 heme a3
V
H
Complex V
CN
H -transporting ATP synthase
IV
CO
A
A
3.6.1.34
P P
P
nH
P P P
Mr >400 kDa, >20 subunits
2
O
2 H
1/2 O
Electron flow
2
+0.8
Proton flow
H2O
B. Organization
Outer
mitochon-
drial
membrane
Inter-
membrane
space
4 H
2 H
4 H
II
Inner
I
Cyto-
mitochon-
?
Q
∆p
chrome C
drial
III
IV
V
membrane
4 H
2-4 H
2e
1/2 O
Matrix
4 H
2
O2
3 P
space
2 H
N
A
N
A
A
A
3
3
P P
P P P
ATP
Tricarboxylic
10 H
NAD
H2O
acid cycle
NADH+H
β-Oxidation
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
142
Metabolism
form complexes with inorganic sulfide and
ATP synthesis
the SH groups of cysteine residues
(see
In the respiratory chain (see p. 140), electrons
p. 286). Ubiquinone
(coenzyme Q; see
are transferred from NADH or ubiquinol (QH2)
p. 104) is a mobile carrier that takes up elec-
to O2. The energy obtained in this process is
trons from complexes I and II and from re-
used to establish a proton gradient across the
duced ETF and passes them on to complex III.
inner mitochondrial membrane. ATP synthe-
Heme groups are also involved in electron
sis is ultimately coupled to the return of pro-
transport in a variety of ways. Type b hemes
tons from the intermembrane space into the
correspond to that found in hemoglobin (see
matrix.
p. 280). Heme c in cytochrome c is covalently
bound to the protein, while the tetrapyrrole
ring of heme a is isoprenylated and carries a
A. Redox systems of the respiratory chain
formyl group. In complex IV, a copper ion (CuB)
The electrons provided by NADH do not reach
and heme a3 react directly with oxygen.
oxygen directly, but instead are transferred to
it in various steps. They pass through at least
B. ATP synthase
10 intermediate redox systems, most of which
are bound as prosthetic groups in complexes
The ATP synthase (EC 3.6.1.34, complex V) that
I, III, and IV. The large number of coenzymes
transports H+ is a complex molecular ma-
involved in electron transport may initially
chine. The enzyme consists of two parts—a
appear surprising. However, as discussed on
proton channel (Fo, for “oligomycin-sensitive”)
p. 18, in redox reactions, the change in free
that is integrated into the membrane; and a
enthalpy ∆G—i. e., the chemical work that is
catalytic unit (F1) that protrudes into the ma-
done—depends only on the difference in re-
trix. The Fo part consists of 12 membrane-
dox potentials ∆E between the donor and the
spanning c-peptides and one a-subunit. The
acceptor. Introducing additional redox sys-
“head” of the F1 part is composed of three α
tems does not alter the reaction’s overall en-
and three β subunits, between which there
ergy yield. In the case of the respiratory chain,
are three active centers. The “stem” between
the difference between the normal potential
Fo and F1 consists of one γ and one ε subunit.
of the donor (NAD+/NADH+H+, E0 = -0.32 V)
Two more polypeptides, b and δ, form a kind
and that of the acceptor
(O2/H2O, E0 =
of “stator,” fixing the α and β subunits relative
+0.82 V) corresponds to an energy difference
to the Fo part.
∆G0
of more than 200 kJ mol-1. This large
The catalytic cycle can be divided into
amount is divided into smaller, more
three phases, through each of which the three
manageable “packages,” the size of which is
active sites pass in sequence. First, ADP and Pi
determined by the difference in redox poten-
are bound (1), then the anhydride bond forms
tials between the respective intermediates. It
(2), and finally the product is released (3).
is assumed that this division is responsible for
Each time protons pass through the Fo chan-
the astonishingly high energy yield (about
nel protein into the matrix, all three active
60%) achieved by the respiratory chain.
sites change from their current state to the
The illustration shows the important redox
next. It has been shown that the energy for
systems involved in mitochondrial electron
proton transport is initially converted into a
transport and their approximate redox poten-
rotation of the γ subunit, which in turn cycli-
tials. These potentials determine the path fol-
cally alters the conformation of the α and β
lowed by the electrons, as the members of a
subunits, which are stationary relative to the
redox series have to be arranged in order of
Fo part, and thereby drives ATP synthesis.
increasing redox potential if transport is to
occur spontaneously (see p. 32).
In complex 1, the electrons are passed from
NADH+H+ first to FMN (see p. 104) and then
on to several iron-sulfur (Fe/S) clusters. These
redox systems are only stable in the interior
of proteins. Depending on the type, Fe/S clus-
ters may contain two to six iron ions, which
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
143
A. Redox systems of the respiratory chain
∆G'
E'
FMN/FMNH2
(kJ · mol-1)
(V)
F
F
NAD
-0.4
N
A
I
CoQ
Hemes b
(ubi-
0
quinone)
3+
2+
N
A
–0.2
O
NADH
+H
III
0
O
Cyto-
chrome C
Heme a
OH
Cu2
3+
2+
+0.2
3+
Cu
Fe/S centers
IV
OH
2+
+0.4
Fe/S center
Cu2
Cu
3+
2+
+0.6
Heme c1
3+
2+
∆G = -n · F · ∆E
O
-220
+0.8
Heme a
O2
3
B. ATP synthase
Inside:
A
P P
P
matrix
2
Formation of ATP
A
H2O
P P P
A
Outside:
H
P P P
2O
Inter-
membrane
H
H
space
outside
outside
α β
α
δ
β
β
α
F1
A
H
P P
P
inside
H
b2
ε
A
A
P P
P
P P P
γ
A
P P P
a
F0
A
A
H2O
P
P P
H2O
P
P P
A
A
P P
P
P P P
H2O
H2O
H2O
1
Binding of ADP and Pi
3
Release of ATP
H
H
outside
1. Structure and location
2. Catalytic cycle
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
144
Metabolism
to the inner membrane (1) or by lipid-soluble
Regulation
substances that can transport protons
The amount of nutrient degradation and ATP
through the membrane, such as
2,4-
synthesis have to be continually adjusted to
dinitrophenol (DNP, 2). Thermogenin (uncou-
the body’s changing energy requirements.
pling protein-1, UCP-1,
3)—an ion channel
The need to coordinate the production and
(see p. 222) in mitochondria of brown fat tis-
consumption of ATP is already evident from
sue—is a naturally occurring uncoupler.
the fact that the total amounts of coenzymes
Brown fat is found, for example, in newborns
in the organism are low. The human body
and in hibernating animals, and serves exclu-
forms about 65 kg ATP per day, but only con-
sively to generate heat. In cold periods, nor-
tains 3-4 g of adenine nucleotides (AMP, ADP,
epinephrine activates the hormone-sensitive
and ATP). Each ADP molecule therefore has to
lipase (see p. 162). Increased lipolysis leads
be phosphorylated to ATP and dephosphory-
to the production of large quantities of free
lated again many thousand times a day.
fatty acids. Like DNP, these bind H+ ions in the
intermembrane space, pass the UCP in this
form, and then release the protons in the
A. Respiratory control
matrix again. This makes fatty acid degrada-
The simple regulatory mechanism which en-
tion independent of ADP availability—i. e., it
sures that ATP synthesis is “automatically”
takes place at maximum velocity and only
coordinated with ATP consumption is known
produces heat (A). It is becoming increasingly
as respiratory control. It is based on the fact
clear that there are also UCPs in other cells,
that the different parts of the oxidative phos-
which are controlled by hormones such as
phorylation process are coupled via shared
thyroxine
(see p. 374). This regulates the
coenzymes and other factors (left).
ATP yield and what is known as the basal
If a cell is not using any ATP, hardly any ADP
metabolic rate.
will be available in the mitochondria. Without
ADP, ATP synthase (3) is unable to break down
C. Regulation of the tricarboxylic acid cycle
the proton gradient across the inner mito-
chondrial membrane. This in turn inhibits
The most important factor in the regulation of
electron transport in the respiratory chain
the cycle is the NADH/NAD+ ratio. In addition
(2), which means that NADH+H+ can no lon-
to pyruvate dehydrogenase (PDH) and oxoglu-
ger be reoxidized to NAD+. Finally, the result-
tarate dehydrogenase (ODH; see p. 134), cit-
ing high NADH/NAD+ ratio inhibits the tricar-
rate synthase and isocitrate dehydrogenase are
boxylic acid cycle (C), and thus slows down
also inhibited by NAD+ deficiency or an excess
the degradation of the substrate SH2 (1). Con-
of NADH+H+. With the exception of isocitrate
versely, high rates of ATP utilization stimulate
dehydrogenase, these enzymes are also sub-
nutrient degradation and the respiratory
ject to product inhibition by acetyl-CoA, suc-
chain via the same mechanism.
cinyl-CoA, or citrate.
If the formation of a proton gradient is
Interconversion processes (see p. 120) also
prevented
(right), substrate oxidation
(1)
play an important role. They are shown here
and electron transport
(2) proceed much
in detail using the example of the PDH com-
more rapidly. However, instead of ATP, only
plex
(see p. 134). The inactivating protein
heat is produced.
kinase [1a] is inhibited by the substrate pyru-
vate and is activated by the products acetyl-
CoA and NADH+H+. The protein phosphatase
B. Uncouplers
[1b]—like isocitrate dehydrogenase [3] and the
Substances that functionally separate oxida-
ODH complex [4]—is activated by Ca2+. This is
tion and phosphorylation from one another
particularly important during muscle con-
are referred to as uncouplers. They break
traction, when large amounts of ATP are
down the proton gradient by allowing H+
needed. Insulin also activates the PDH com-
ions to pass from the intermembrane space
plex (through inhibition of phosphorylation)
back into the mitochondrial matrix without
and thereby promotes the breakdown of glu-
the involvement of ATP synthase. Uncoupling
cose and its conversion into fatty acids.
effects are produced by mechanical damage
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
145
A. Respiratory control
AH2
A
AH2
A
1
3
Coupling
1
1
points
N
A
N
A
N
A
N
A
Dehydro-
Dehydro-
genases
genases
2
2
H
Respira-
H
H
H
outside
tory chain
inside
ATP syn-
outside
inside
thesis
uncoupled
Heat
from proton
3
transport
3
P
A
ATP
A
A
P
A
P P
synthase
P
P
P
P P
P P
P
Endergonic
processes
Endergonic
processes
1. Coupled
2. Uncoupled
B. Uncouplers
C. Regulation of the
tricarboxylic acid cycle
1. Membrane
Insulin
Pyruvate
H
H
1. damage
1a
1b
Ca2
H
O
NO2
NADH
NO2
2. Mobile
Acetyl CoA
2. carriers
O2N
2
NO2
OH
Oxalo-
2,4-Dinitrophenol
Citrate
acetate
H
Inner
mitochochondrial
Most important
Isocitrate
factor: con-
N
A
membrane
centration ratio
NADH
[NADH] / [NAD
]
3
Norepinephrine
2-Oxo-
Succinyl
glutarate
Fatty
CoA
4
Fat
H
acids
Ca2
PDH kinase
Isocitrate dehydro-
1a
3
H
2.7.1.99
genase 1.1.1.42
Thermo-
genin
PDH phosphatase
2-Oxoglutarate
3. Gated proton
1b
(UCP1)
3.1.3.43
4
dehydrogenase 1.2.4.2,
3. channels
1.8.1.4, 2.3.1.61
Citrate synthase
2
4.1.3.7
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
146
Metabolism
ylic acid cycle and the pyruvate dehydro-
Respiration and fermentation
genase reaction
(see p. 144). β-Oxidation
and the malate shuttle, which are dependent
A. Aerobic and anaerobic oxidation of
on free NAD+, also come to a standstill. Since
glucose
amino acid degradation is also no longer able
In the presence of oxygen (i. e., in aerobic
to contribute to energy production, the cell
conditions), most animal cells are capable of
becomes totally dependent on ATP synthe-
“respiring” various types of nutrient (lipids,
sized via the degradation of glucose by
amino acids, and carbohydrates)—i. e., using
glycolysis. For this process to proceed contin-
oxidative processes to break them down com-
uously, the NADH+H+ formed in the cyto-
pletely. If oxygen is lacking (i. e., in anaerobic
plasm has to be constantly reoxidized. Since
conditions), only glucose can be used for ATP
this can no longer occur in the mitochondria,
synthesis. Although in these conditions glu-
in anaerobic conditions animal cells reduce
cose breakdown in animals already ends in
pyruvate to lactate and pass it into the blood.
lactate and only produces small quantities of
This type of process is called fermentation
ATP, it is decisively important for the survival
(see p. 148). The ATP yield is low, with only
of cells at times of oxygen deficiency.
two ATPs per glucose arising during lactate
synthesis.
In aerobic conditions (left), ATP is derived
almost exclusively from oxidative phosphor-
To estimate the number of ATP molecules
ylation (see p. 140). Fatty acids enter the mi-
formed in an aerobic state, it is necessary to
tochondria with the help of carnitine (see
know the P/O quotient—i. e., the molar ratio
p. 164), and are broken down there into CoA-
between synthesized ATP (“P”) and the water
bound acetyl residues. Glucose is converted
formed (“O”). During transport of two elec-
into pyruvate by glycolysis (see p. 150) in the
trons from NADH+H+ to oxygen, about 10 pro-
cytoplasm. Pyruvate is then also transported
tons are transported into the intermembrane
into the mitochondrial matrix, where it is
space, while from ubiquinol (QH2), the num-
oxidatively decarboxylated by the pyruvate
ber is only six. ATP synthase (see p. 142) prob-
dehydrogenase complex (see p. 134) to yield
ably requires three H+ to synthesize one ATP,
acetyl-CoA. The reducing equivalents
(2
so that maximum P/O quotients of around 3
NADH+H+ per glucose) that arise in glycolysis
or 2 are possible. This implies a yield of up to
enter the mitochondrial matrix via the malate
38 ATP per mol of glucose. However, the ac-
shuttle (see p. 212). The acetyl residues that
tual value is much lower. It needs to be taken
are formed are oxidized to CO2 in the tricar-
into account that the transport of specific me-
boxylic acid cycle (see p. 136). Breakdown of
tabolites into the mitochondrial matrix and
amino acids also produces acetyl residues or
the exchange of ATP4- for ADP3- are also
products that can directly enter the tricarbox-
driven by the proton gradient (see p. 212).
ylic acid cycle
(see p. 180). The reducing
The P/O quotients for the oxidation of
equivalents that are obtained are transferred
NADH+H+ and QH2 are therefore more in the
to oxygen via the respiratory chain as re-
range of 2.5 and 1.5. If the energy balance of
quired. In the process, chemical energy is re-
aerobic glycolysis is calculated on this basis,
leased, which is used (via a proton gradient)
the result is a yield of around 32 ATP per
to synthesize ATP (see p. 140).
glucose. However, this value is also not con-
stant, and can be adjusted as required by the
In the absence of oxygen—i. e., in anaerobic
cell’s own uncouplers (UCPs; see p. 144) and
conditions—the picture changes completely.
other mechanisms.
Since O2 is missing as the electron acceptor
for the respiratory chain, NADH+H+ and QH2
can no longer be reoxidized. Consequently,
not only is mitochondrial ATP synthesis
halted, but also almost the whole metabolism
in the mitochondrial matrix. The main reason
for this is the high NADH+H+ concentration
and lack of NAD+, which inhibit the tricarbox-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
147
A. Aerobic and anaerobic oxidation of glucose
Glucose
1
2
Glucose
1
2
A
A
A
A
P P P
P P P
P P P
P P
P
Glyco-
lysis
2
PEP
4
3
2
PEP
4
3
A
P
A
P
2
2
P
P
P P
N
A
N
A
N
A
N
A
5
2
2
5
2
2
A
A
2
P
P
P
2
P P P
2
Pyruvate
2
Pyruvate
6
2
Lactate
Malate shuttle
Pyruvate
Pyruvate
7
7
N
A
Acetyl
N
A
N
A
Acetyl
N
A
CoA
CoA
Tricarboxylic
H
H
acid cycle
8
inhibited due
8
to high ratio of
9
9
NADH/NAD
10
10
H
H
11
11
12
12
O2
H
O2
H
2 GTP
H2O
P
P
A
A
A
A
P P
P
P
P
P
P P
P
P
1. Aerobic
2. Anaerobic
ATP
Coenzymes
Enzymes
Coenzymes
ATP
-1
–1 ATP
1
Hexokinase
–1 ATP
-1
-2
–1 ATP
2
6-Phosphofructokinase
–1 ATP
-2
+3
+5 ATP
+2 NADH
3
Glyceraldehyde-3 P DH
+2 NADH
-2
+5
+2 ATP
4
Phosphoglycerate kinase
+2 ATP
0
NAD
+7
+2 ATP
5
Pyruvate kinase
+2 ATP
recycled
+2
6
Lactate dehydrogenase
-2 NADH
+12
+5 ATP
+2 NADH
7
Pyruvate dehydrogenase
+17
+5 ATP
+2 NADH
8
Isocitrate dehydrogenase
+22
+5 ATP
+2 NADH
9
Oxoglutarate dehydrogenase
+27
+5 ATP
+2 NADH
10
Malate dehydrogenase
+30
+3 ATP
+2 QH2
11
Succinate dehydrogenase
+32
+2 ATP
+2 GTP
12
Succinate-CoA ligase
DH = dehydrogenase
Sum: 32 ATP/glucose
Sum: 2 ATP/glucose
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
148
Metabolism
lated by pyruvate decarboxylase [2], which
Fermentations
does not occur in animal metabolism, to pro-
As discussed on p. 146, degradation of glucose
duce acetaldehyde (ethanal). When this is re-
to pyruvate is the only way for most organ-
duced by alcohol dehydrogenase [3], with
isms to synthesize ATP in the absence of oxy-
NADH being consumed, ethanol [3] is formed.
gen. The NADH+H+ that is also formed in this
Yeasts, unicellular fungi that belong to the
process has to be constantly reoxidized to
eukaryotes (3), rather than bacteria, are re-
NAD+ in order to maintain glycolysis and
sponsible for this type of fermentation. Yeasts
thus ATP synthesis. In the animal organism,
are also often used in baking. They produce
this is achieved by the reduction of pyruvate
CO2 and ethanol, which raise the dough.
to lactate. In microorganisms, there are many
Brewers’ and bakers’ yeasts (Saccharomyces
other forms of NAD+ regeneration. Processes
cerevisiae) are usually haploid and reproduce
of this type are referred to as fermentations.
asexually by budding (3). They can live both
Microbial fermentation processes are often
aerobically and anaerobically. Wine is pro-
used to produce foodstuffs and alcoholic bev-
duced by other types of yeast, some of which
erages, or to preserve food. Features common
already live on the grapes. To promote the
to all fermentation processes are that they
formation of ethanol, efforts are made to gen-
start with pyruvate and only occur under
erally exclude oxygen during alcoholic fer-
anaerobic conditions.
mentation—for example, by covering dough
with a cloth when it is rising and by ferment-
ing liquids in barrels that exclude air.
A. Lactic acid and propionic acid
fermentation
C. Beer brewing
Many milk products, such as sour milk, yo-
gurt, and cheese are made by bacterial lactic
Barley is the traditional starting material for
acid fermentation (1). The reaction is the same
the brewing of beer. Although cereal grains
as in animals. Pyruvate, which is mainly de-
contain starch, they hardly have any free sug-
rived from degradation of the disaccharide
ars. The barley grains are therefore first al-
lactose (see p. 38), is reduced to lactate by
lowed to germinate so that starch-cleaving
lactate dehydrogenase [1]. Lactic acid fermen-
amylases are formed. Careful warming of the
tation also plays an important role in the
sprouting grain produces malt. This is then
production of sauerkraut and silage. These
ground, soaked in water, and kept warm for
products usually keep for a long time, because
a certain time. In the process, a substantial
the pH reduction that occurs during fermen-
proportion of the starch is broken down into
tation inhibits the growth of putrefying bac-
the disaccharide maltose
(see p. 38). The
teria.
product (the wort) is then boiled, yeast and
Bacteria from the genera Lactobacillus and
hops are added, and the mixture is allowed to
Streptococcus are involved in the first steps of
ferment for several days. The addition of hops
dairy production (3). The raw materials pro-
makes the beer less perishable and gives it its
duced by their effects usually only acquire
slightly bitter taste. Other substances con-
their final properties after additional fermen-
tained in hops act as sedatives and diuretics.
tation processes. For example, the character-
istic taste of Swiss cheese develops during a
subsequent propionic acid fermentation. In
this process, bacteria from the genus
Propionibacterium convert pyruvate to propi-
onate in a complex series of reactions (2).
B. Alcoholic fermentation
Alcoholic beverages are produced by the fer-
mentation of plant products that have a high
carbohydrate content. Pyruvate, which is
formed from glucose, is initially decarboxy-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Energy Metabolism
149
A. Lactic acid and propionic acid fermentation
1.
Propionate
H
H
2.
Lactobacillus
3.
O
H
C
C C
O
H H
Propioni-
H
OH
bacterium
Lactate
O
H
C
C C
Streptococcus
H
H O
10 µm
i
N
A
2[H]
1
2[H]
N
A
DNA Cell wall
H
O
O
H
C
C C
B. Alcoholic fermentation
Lactate dehydrogenase
H
O
1
1.
2.
10 µm
3.
1.1.1.27
Pyruvate
Pyruvate decarboxylase
2
[TPP] 4.1.1.1
O
2
Alcohol dehydrogenase
3
Yeast
[Zn2 ] 1.1.1.1
CO2
C
(Saccharomyces
cerevisiae)
O
H
O
Vacuole
Septum Daughter cell
H
C C
Ethanal
(Acet-
H H
aldehyde)
N
A
3
Nucleus
N
A
ER
H
H
Cell wall
H
C C
OH
Tonoplast
Mitochondrium
H H
Ethanol
C. Beer brewing
CO
2
Germinate,
Malt
Grind,
Wort
Hops
dry
incubate
O2
in water
Barley
Fermentation
Starch
Maltose
Amylases
Amylases
Yeast
Glucose
formed in
Maltose
Ethanol
+ CO2
seedling
1.
2.
3.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
150
Metabolism
phorylation; see p. 124), and
1,3-bisphos-
Glycolysis
phoglycerate is produced. This intermediate
contains a mixed acid-anhydride bond, the
A. Balance
phosphate part of which is at a high chemical
Glycolysis is a catabolic pathway in the cyto-
potential.
plasm that is found in almost all organisms—
[7] Catalyzed by phosphoglycerate kinase,
irrespective of whether they live aerobically
this phosphate residue is transferred to ADP,
or anaerobically. The balance of glycolysis is
producing 3-phosphoglycerate and ATP. The
simple: glucose is broken down into two mol-
ATP balance is thus once again in equilibrium.
ecules of pyruvate, and in addition two mol-
[8] As a result of shifting of the remaining
ecules of ATP and two of NADH+H+ are
phosphate residue within the molecule, the
formed.
isomer 2-phosphoglycerate is formed.
In the presence of oxygen, pyruvate and
[9] Elimination of water from 2-phospho-
NADH+H+ reach the mitochondria, where
glycerate produces the phosphate ester of the
they undergo further transformation (aerobic
enol form of pyruvate—phosphoenolpyruvate
glycolysis; see p. 146). In anaerobic condi-
(PEP). This reaction also raises the second
tions, fermentation products such as lactate
phosphate residue to a high potential.
or ethanol have to be formed in the cytoplasm
[10] In the last step, pyruvate kinase trans-
from pyruvate and NADH+H+, in order to re-
fers this residue to ADP. The remaining enol
generate NAD+ so that glycolysis can continue
pyruvate is immediately rearranged into
(anaerobic glycolysis; see p. 146). In the anae-
pyruvate, which is much more stable. Along
robic state, glycolysis is the only means of
with step [7] and the thiokinase reaction in
obtaining ATP that animal cells have.
the tricarboxylic acid cycle (see p. 136), the
pyruvate kinase reaction is one of the three
reactions in animal metabolism that are able
B. Reactions
to produce ATP independently of the respira-
Glycolysis involves ten individual steps, in-
tory chain.
cluding three isomerizations and four phos-
In glycolysis, two molecules of ATP are ini-
phate transfers. The only redox reaction takes
tially used for activation ([1], [3]). Later, two
place in step [6].
ATPs are formed per C3 fragment. Overall,
[1] Glucose, which is taken up by animal
therefore, there is a small net gain of 2 mol
cells from the blood and other sources, is first
ATP per mol of glucose.
phosphorylated to glucose 6-phosphate, with
ATP being consumed. The glucose 6-phos-
C. Energy profile
phate is not capable of leaving the cell.
[2] In the next step, glucose 6-phosphate is
The energy balance of metabolic pathways de-
isomerized into fructose 6-phosphate.
pends not only on the standard changes in
[3] Using ATP again, another phosphoryla-
enthalpy ∆G0, but also on the concentrations
tion takes place, giving rise to fructose 1,6-
of the metabolites (see p. 18). Fig. C shows the
bisphosphate. Phosphofructokinase is the
actual enthalpy changes ∆G for the individual
most important key enzyme in glycolysis
steps of glycolysis in erythrocytes.
(see p. 144).
As can be seen, only three reactions ([1],
[4] Fructose
1,6-bisphosphate is broken
[3], and
[10]), are associated with large
down by aldolase into the C3 compounds glyc-
changes in free enthalpy. In these cases, the
eraldehyde 3-phosphate (also known as glyc-
equilibrium lies well on the side of the prod-
eral 3-phosphate) and glycerone 3-phosphate
ucts (see p. 18). All of the other steps are
(dihydroxyacetone 3-phosphate).
freely reversible. The same steps are also fol-
[5] The latter two products are placed in
lowed—in the reverse direction—in gluconeo-
fast equilibrium by triosephosphate isomerase.
genesis (see p. 154), with the same enzymes
[6] Glyceraldehyde
3-phosphate is now
being activated as in glucose degradation. The
oxidized by glyceraldehyde-3-phosphate de-
non-reversible steps [1], [3], and [10] are by-
hydrogenase, with NADH+H+ being formed.
passed in glucose biosynthesis (see p. 154).
In this reaction, inorganic phosphate is taken
up into the molecule (substrate-level phos-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrate Metabolism
151
A. Glycolysis: balance
2
N
A
2
N
A
HO
CH
2
COO
COO
H
O
OH
Glycolysis
C O
C O
OH H
HO
H
∆GOI = -35 kJ · mol-1
CH
CH3
3
H
OH
2
A
2
A
P P
P P P
Glucose
Pyruvate
Pyruvate
2
P
B. Reactions
Glucose
Fructose
Fructose
Glucose
6-phosphate
6-phosphate
1,6-bisphosphate
HO
CH2
P
O
CH2
P
O
CH2
P
O
CH2
O OH
O OH
H
O
OH
H
O
OH
H HO
H HO
OH
H H
1
OH
H H
2
3
H
CH2OH
H
CH2
O
P
HO
H
HO
H
OH H
OH H
H
OH
H
OH
A
A
A
A
P
P
P
P P
P P
P
P P
2
2
4
A
A
N
A
N
A
2
P
P P
2
P P
O
O
O
O
O
O P
O
H
C
C
C
C
H2C OH
HC O
P
8
7
HC OH
6
HC OH
5
HC OH
C O
H2C OH
H2C O
P
H2C O
P
H2C O
P
P
O
CH2
2 P
2-Phospho-
3-Phospho-
1,3-Bisphospho-
Glyceraldehyde
Glycerone
2
2
2
glycerate
glycerate
glycerate
3-phosphate
3-phosphate
9
2 H2O
C. Energy profile
0
O
O
O O
A
Steps 1, 3 and 10
C
C
P P P
are bypassed in
gluconeogenesis
C O P
10
C O
1
CH2
CH3
-20
A
P P
Phospho-
2
2 Pyruvate
enolpyruvate
A
A
2
2
P P
2
P P P
A
A
P P
P P P
A
-40
P P P
Glyceraldehyde-
1
Hexokinase 2.7.1.1
6
P
3-
dehydro-
genase 1.2.1.12
3
Glucose 6-phosphate
2
5
8
9
Isomerase 5.3.1.9
A
Phosphoglycerate
P P
7
kinase 2.7.2.3
6-Phosphofructo-
3
-60
kinase 2.7.1.11
Phosphoglycerate
4
A
8
6
7
P P
Fructose bisphosphate
mutase 5.4.2.1
4
10
aldolase 4.1.2.13
Phosphopyruvate
A
9
P P P
hydratase 4.2.1.11
Triose-phosphate
5
isomerase 5.3.1.1
-80
Pyruvate kinase
10
Pyruvate
2.7.1.40
∆G' (kJ · mol-1)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
152
Metabolism
of the regenerative branch is to adjust the net
Pentose phosphate pathway
production of NADPH+H+ and pentose phos-
The pentose phosphate pathway (PPP, also
phates to the cell’s current requirements. Nor-
known as the hexose monophosphate
mally, the demand for NADPH+H+ is much
pathway) is an oxidative metabolic pathway
higher than that for pentose phosphates. In
located in the cytoplasm, which, like glycoly-
these conditions, the reaction steps shown
sis, starts from glucose 6-phosphate. It sup-
first convert six ribulose
5-phosphates to
plies two important precursors for anabolic
five molecules of fructose 6-phosphate and
pathways: NADPH+H+, which is required for
then, by isomerization, regenerate five glu-
the biosynthesis of fatty acids and isopren-
cose 6-phosphates. These can once again sup-
oids, for example (see p. 168), and ribose 5-
ply NADPH+H+ to the oxidative part of the
phosphate, a precursor in nucleotide biosyn-
PPP. Repeating these reactions finally results
thesis (see p. 188).
in the oxidation of one glucose 6-phosphate
into six CO2. Twelve NADPH+H+ arise in the
same process. In sum, no pentose phosphates
A. Pentose phosphate pathway:
are produced via this pathway.
oxidative part
In the recombination of sugar phosphates
The oxidative segment of the PPP converts
in the regenerative part of the PPP, there are
glucose 6-phosphate to ribulose 5-phosphate.
two enzymes that are particularly important:
One CO2 and two NADPH+H+ are formed in
[5] Transaldolase transfers C3 units from
the process. Depending on the metabolic
sedoheptulose 7-phosphate, a ketose with
state, the much more complex regenerative
seven C atoms, to the aldehyde group of glyc-
part of the pathway (see B) can convert some
eraldehyde 3-phosphate.
of the pentose phosphates back to hexose
[4] Transketolase, which contains thiamine
phosphates, or it can pass them on to glycol-
diphosphate, transfers C2 fragments from one
ysis for breakdown. In most cells, less than
sugar phosphate to another.
10% of glucose 6-phosphate is degraded via
The reactions in the regenerative segment
the pentose phosphate pathway.
of the PPP are freely reversible. It is therefore
easily possible to use the regenerative part of
the pathway to convert hexose phosphates
B. Reactions
into pentose phosphates. This can occur
[1] The oxidative part starts with the oxida-
when there is a high demand for pentose
tion of glucose
6-phosphate by glucose-6-
phosphates—e. g., during DNA replication in
phosphate dehydrogenase. This forms
the S phase of the cell cycle (see p. 394).
NADPH+H+ for the first time. The second
product, 6-phosphogluconolactone, is an in-
Additional information
tramolecular ester
(lactone) of
6-phospho-
gluconate.
When energy in the form of ATP is required in
[2] A specific hydrolase then cleaves the
addition to NADPH+H+, the cell is able to
lactone, exposing the carboxyl group of
channel the products of the regenerative
6-phosphogluconate.
part of the PPP (fructose 6-phosphate and
[3] The last enzyme in the oxidative part is
glyceraldehyde 3-phosphate) into glycolysis.
phosphogluconate dehydrogenase [3], which
Further degradation is carried out via the tri-
releases the carboxylate group of 6-phospho-
carboxylic acid cycle and the respiratory chain
gluconate as CO2 and at the same time oxi-
to CO2 and water. Overall, the cell in this way
dizes the hydroxyl group at C3 to an oxo
obtains
12 mol NADPH+H+ and around
group. In addition to a second NADPH+H+,
150 mol ATP from 6 mol glucose 6-phos-
this also produces the ketopentose ribulose
phate. PPP activity is stimulated by insulin
5-phosphate. This is converted by an isomer-
(see p. 388). This not only increases the rate
ase to ribose 5-phosphate, the initial com-
of glucose degradation, but also produces ad-
pound for nucleotide synthesis (top).
ditional NADPH+H+ for fatty acid synthesis
The regenerative part of the PPP is only
(see p. 168).
shown here schematically. A complete reac-
tion scheme is given on p. 408. The function
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrate Metabolism
153
A. Pentose phosphate pathway: oxidative part
CO
CH2
P
O
2
P
O
H H O H
H
O
OH
PPP -
oxidative
H
C C
C C C H
OH
H H
part
H OH OH OH
HO
H
N
A
N
A
2
2
H
OH
P
P
Glucose 6-phosphate
NADP
NADPH + H
Ribulose 5-phosphate
B. Reactions
P
O H H O H
P P
P P
P P
Ribulose
Ribulose
6
H
C C
C C C H
5-phosphate
5-
P
H OH OH OH
N
A
6
P
6 CO
2
3
P P
P P
P P
a
Ribose
N
A
6
5-
P
P
Xylulose
b
P
O H H OH H
5-
P
O
a
b
b
6-Phospho-
H
C C
C C C
C
6
4
gluconate
H OH OH H OH
O
P
P
Sedo-
Anabolic
a
heptulose
2
P P
7-
P
pathways
H2O
P
O
CH
Glyceral
2
b
3-
P
6-Phospho-
a
b
H
O
6
glucono-
H
lactone
OH H
C
O
5
HO
H
OH
P
P
Fructose
a
P P
6-
P
N
A
6
P
Erythrose
b
1
4-
P
N
A
a
b
6
P
4
P
O
CH2
P
P
P
Fructose
a
Glucose
H
O OH
6-
P
6
6-phosphate
P P
Glycer-
OH
H H
b
aldehyde
HO
H
3-
P
a
a
b
H
OH
1
Glucose 6-phosphate dehydrogenase 1.1.1.49
4
Transketolase 2.2.1.1
2
Gluconolactonase 3.1.1.17
5
Transaldolase 2.2.1.2
3
Phosphogluconate dehydrogenase (decarboxylating) 1.1.1.44
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
154
Metabolism
from lactate or amino acids is therefore ini-
Gluconeogenesis
tially transported into the mitochondrial ma-
Some tissues, such as brain and erythrocytes,
trix, and—in a biotin-dependent reaction cat-
depend on a constant supply of glucose. If the
alyzed by pyruvate carboxylase—is carboxy-
amount of carbohydrate taken up in food is
lated there to oxaloacetate. Oxaloacetate is
not suf cient, the blood sugar level can be
also an intermediate in the tricarboxylic acid
maintained for a limited time by degradation
cycle. Amino acids with breakdown products
of hepatic glycogen (see p. 156). If these re-
that enter the cycle or supply pyruvate can
serves are also exhausted, de-novo synthesis
therefore be converted into glucose
(see
of glucose (gluconeogenesis) begins. The liver
p. 180).
is also mainly responsible for this (see p. 310),
[3] The oxaloacetate formed in the mito-
but the tubular cells of the kidney also show a
chondrial matrix is initially reduced to ma-
high level of gluconeogenetic activity (see
late, which can leave the mitochondria via
p. 328). The main precursors for gluconeo-
inner membrane transport systems
(see
genesis are amino acids derived from muscle
p. 212).
proteins. Another important precursor is
[4] In the cytoplasm, oxaloacetate is re-
lactate, which is formed in erythrocytes and
formed and then converted into phospho-
muscle proteins when there is oxygen de-
enol pyruvate by a GTP-dependent PEP car-
ficiency. Glycerol produced from the degrada-
boxykinase. The subsequent steps up to fruc-
tion of fats can also be used for gluconeogen-
tose 1,6-bisphosphate represent the reverse
esis. However, the conversion of fatty acids
of the corresponding reactions involved in
into glucose is not possible in animal metab-
glycolysis. One additional ATP per C3 frag-
olism (see p. 138). The human organism can
ment is used for the synthesis of 1,3-bisphos-
synthesize several hundred grams of glucose
phoglycerate.
per day by gluconeogenesis.
Two gluconeogenesis-specific phosphat-
ases then successively cleave off the phos-
phate residues from fructose
1,6-bisphos-
A. Gluconeogenesis
phate. In between these reactions lies the
Many of the reaction steps involved in gluco-
isomerization of fructose 6-phosphate to glu-
neogenesis are catalyzed by the same en-
cose
6-phosphate—another glycolytic reac-
zymes that are used in glycolysis
(see
tion.
p. 150). Other enzymes are specific to gluco-
[5] The reaction catalyzed by fructose
neogenesis and are only synthesized, under
1,6-bisphosphatase is an important regulation
the influence of cortisol and glucagon when
point in gluconeogenesis (see p. 158).
needed (see p. 158). Glycolysis takes place
[6] The last enzyme in the pathway, glucose
exclusively when needed in the cytoplasm,
6-phosphatase, occurs in the liver, but not in
but gluconeogenesis also involves the mito-
muscle. It is located in the interior of the
chondria and the endoplasmic reticulum (ER).
smooth endoplasmic reticulum. Specific
Gluconeogenesis consumes 4 ATP (3 ATP + 1
transporters allow glucose
6-phosphate to
GTP) per glucose—i. e., twice as many as gly-
enter the ER and allow the glucose formed
colysis produces.
there to return to the cytoplasm. From there,
it is ultimately released into the blood.
[1] Lactate as a precursor for gluconeogen-
esis is mainly derived from muscle (see Cori
Glycerol initially undergoes phosphoryla-
cycle, p. 338) and erythrocytes. LDH
(see
tion at C-3
[7]. The glycerol
3-phosphate
p. 98) oxidizes lactate to pyruvate, with
formed is then oxidized by an NAD+-depen-
NADH+H+ formation.
dent dehydrogenase to form glycerone 3-
[2] The first steps of actual gluconeogenesis
phosphate [8] and thereby channeled into
take place in the mitochondria. The reason for
gluconeogenesis. An FAD-dependent mito-
this “detour” is the equilibrium state of the
chondrial enzyme is also able to catalyze
pyruvate kinase reaction (see p. 150). Even
this reaction
(known as the “glycerophos-
coupling to ATP hydrolysis would not be suf-
phate shuttle”; see p. 212).
ficient to convert pyruvate directly into phos-
phoenol pyruvate
(PEP). Pyruvate derived
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrate Metabolism
155
A. Gluconeogenesis
Lactate
Glucose
1
dehydrogenase
1.1.1.27
Glucose 6-phosphate
Glucose
Pyruvate
P
O
CH2
H2O
HO
CH2
2
carboxylase
(biotin)6.4.1.1
H
O
OH
H
OOH
6
OH
H H
OH
H H
Malate
3
dehydrogenase
HO
H
HO
H
1.1.1.37
P
H
OH
H
OH
gER
PEP
4
carboxykinase
4.1.1.32
Glucose 6-phosphate
Fructose 1,6-
5
bisphosphate
P
O
CH2
P
O
CH2
OH
P
3.1.3.11
O
O OH
H HO
H HO
Glucose 6-
5
6
phosphatase
H
CH2OH
H
CH2
O
P
3.1.3.9
H2O
OH H
OH H
Fructose 6-phosphate
Fructose 1,6-bisphosphate
Glycerol kinase
7
2.7.1.30
ATP ADP
Glycerol 3-
3-Phospho-
1,3-Bisphospho- Glyceraldehyde
Glycerone
phosphate
8
glycerate
glycerate
3-phosphate
3-phosphate
dehydrogenase
1.1.1.8
NADH
+H
8
NAD NADH
NAD
2-Phospho-
NADH+H NAD
Glycerol
3
glycerate
3-phosphate
ADP
7
GTP GDP
ATP
COO
COO
COO
COO
H2C OH
4
1
C
O
C O
P
C O
H
C
OH
HC OH
Malate
CH2
CO2
CH2
CH3
CH3
H2C OH
COO
Pyruvate
Lactate
Glycerol
Phospho-
Oxaloacetate
enolpyruvate
COO
COO
CO2
COO
H
C
OH
C O
3
2
C O
CH2
CH2
NAD
NADH
ADP ATP
CH3
COO
+H
COO
P
Pyruvate
Malate
Oxaloacetate
Mitochrondrion
Amino acids
Glycerol
Cytoplasm
Lactate
Amino acids
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
156
Metabolism
important degradative enzyme, glycogen
Glycogen metabolism
phosphorylase, cleaves residues from a non-
Glycogen (see p. 40) is used in animals as a
reducing end one after another as glucose
carbohydrate reserve, from which glucose
1-phosphate. The larger the number of these
phosphates and glucose can be released
ends, the more phosphorylase molecules can
when needed. Glucose storage itself would
attack simultaneously. The formation of glu-
not be useful, as high concentrations within
cose 1-phosphate instead of glucose has the
cells would make them strongly hypertonic
advantage that no ATP is needed to channel
and would therefore cause an influx of water.
the released residues into glycolysis or the
By contrast, insoluble glycogen has only low
PPP.
osmotic activity.
[5] [6] Due to the structure of glycogen
phosphorylase, degradation comes to a halt
four residues away from each branching
A. Glycogen balance
point. Two more enzymes overcome this
Animal glycogen, like amylopectin in plants, is
blockage. First, a glucanotransferase moves a
a branched homopolymer of glucose. The glu-
trisaccharide from the side chain to the end of
cose residues are linked by an α1
4-glyco-
the main chain [5]. A 1,6-glucosidase [6] then
sidic bond. Every tenth or so glucose residue
cleaves the single remaining residue as a free
has an additional α1
6 bond to another
glucose and leaves behind an unbranched
glucose. These branches are extended by
chain that is once again accessible to phos-
additional α1
4-linked glucose residues.
phorylase.
This structure produces tree-shaped mole-
The regulation of glycogen metabolism by
cules consisting of up to
50000 residues
interconversion, and the role of hormones in
(M > 1
107 Da).
these processes, are discussed on p. 120.
Hepatic glycogen is never completely de-
graded. In general, only the nonreducing ends
B. Glycogen balance
of the “tree” are shortened, or—when glucose
is abundant—elongated. The reducing end of
The human organism can store up to 450 g of
the tree is linked to a special protein, glyco-
glycogen—one-third in the liver and almost all
genin. Glycogenin carries out autocatalytic
of the remainder in muscle. The glycogen
covalent bonding of the first glucose at one
content of the other organs is low.
of its tyrosine residues and elongation of this
Hepatic glycogen is mainly used to main-
by up to seven additional glucose residues. It
tain the blood glucose level in the postresorp-
is only at this point that glycogen synthase
tive phase (see p. 308). The glycogen content
becomes active to supply further elongation.
of the liver therefore varies widely, and can
[1] The formation of glycosidic bonds be-
decline to almost zero in periods of extended
tween sugars is endergonic. Initially, there-
hunger. After this, gluconeogenesis
(see
fore, the activated form—UDP-glucose—is
p. 154) takes over the glucose supply for the
synthesized by reaction of glucose 1-phos-
organism. Muscle glycogen serves as an energy
phate with UTP (see p. 110).
reserve and is not involved in blood glucose
[2] Glycogen synthase now transfers glu-
regulation. Muscle does not contain any glu-
cose residues one by one from UDP-glucose
cose 6-phosphatase and is therefore unable to
to the non-reducing ends of the available
release glucose into the blood. The glycogen
“branches.”
content of muscle therefore does not fluctuate
[3] Once the growing chain has reached a
as widely as that of the liver.
specific length (> 11 residues), the branching
enzyme cleaves an oligosaccharide consisting
of 6-7 residues from the end of it, and adds
this into the interior of the same chain or a
neighboring one with α1
6 linkage. These
branches are then further extended by glyco-
gen synthase.
[4] The branched structure of glycogen al-
lows rapid release of sugar residues. The most
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrate Metabolism
157
A. Glycogen metabolism
α-1,4-Glucan chain
α-1,4-Glucan chain
(unbranched)
(α-1,6-branched)
3
R
R
Reducing
end
U
P
P P
4
UDP-
Degra-
2
4
dation
glucose
U
P
P P
Glucose 1-
phosphate
P P
4
R
1
UTP
Further
R
degradation
P
5
Glucose
1-phosphate
6
Building block
synthesis
R
Glucose
UTP-glucose 1-phosphate
1
4
Phosphorylase 2.4.1.1
uridyltransferase 2.7.7.9
2
Glycogen synthase 2.4.1.11
5
4-α-Glucanotransferase 2.4.1.25
Glucan branching enzyme
Amylo-1,6-glucosidase
3
6
2.4.1.18
3.2.1.33
B. Glycogen balance
Liver
Blood
Muscle
Liver
Muscle
glycogen
glycogen
4 - 6 g · l-1
300 g
Blood
Glucose
Glucose
glucose
1-phosphate
1-phosphate
200 g
150 g
De-
Glucose
Glucose
Degra-
grad-
6-phosphate
6-phosphate
dation
ation
1g
Glucose
Glucose
Glucose
Liver
Muscle
glycogen
glycogen
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
158
Metabolism
Glucocorticoids—mainly cortisol
(see
Regulation
p. 374)—induce all of the key enzymes in-
volved in gluconeogenesis [4, 6, 8, 9]. At the
A. Regulation of carbohydrate metabolism
same time, they also induce enzymes in-
In all organisms, carbohydrate metabolism is
volved in amino acid degradation and thereby
subject to complex regulatory mechanisms
provide precursors for gluconeogenesis. Reg-
involving hormones, metabolites, and
ulation of the expression of PEP carboxy-
coenzymes. The scheme shown here (still a
kinase, a key enzyme in gluconeogenesis, is
simplified one) applies to the liver, which
discussed in detail on p. 244.
has central functions in carbohydrate metab-
Metabolites. High concentrations of ATP
olism (see p. 306). Some of the control mech-
and citrate inhibit glycolysis by allosteric reg-
anisms shown here are not effective in other
ulation of phosphofructokinase. ATP also
tissues.
inhibits pyruvate kinase. Acetyl-CoA, an inhib-
One of the liver’s most important tasks is to
itor of pyruvate kinase, has a similar effect. All
store excess glucose in the form of glycogen
of these metabolites arise from glucose
and to release glucose from glycogen when
degradation (feedback inhibition). AMP and
required (buffer function). When the glycogen
ADP, signals for ATP deficiency, activate gly-
reserves are exhausted, the liver can provide
cogen degradation and inhibit gluconeogene-
glucose by de novo synthesis (gluconeogene-
sis.
sis; see p. 154). In addition, like all tissues, the
liver breaks glucose down via glycolysis.
B. Fructose 2,6-bisphosphate
These functions have to be coordinated with
each other. For example, there is no point in
Fructose 2,6-bisphosphate (Fru-2,6-bP) plays
glycolysis and gluconeogenesis taking place
an important part in carbohydrate metabo-
simultaneously, and glycogen synthesis and
lism. This metabolite is formed in small quan-
glycogen degradation should not occur simul-
tities from fructose
6-phosphate and has
taneously either. This is ensured by the fact
purely regulatory functions. It stimulates gly-
that two different enzymes exist for important
colysis by allosteric activation of phospho-
steps in both pathways, each of which cata-
fructokinase and inhibits gluconeogenesis by
lyzes only the anabolic or the catabolic reac-
inhibition of fructose 1,6-bisphosphatase.
tion. The enzymes are also regulated differ-
The synthesis and degradation of Fru-2,6-
ently. Only these key enzymes are shown
bP are catalyzed by one and the same protein
here.
[10a, 10b]. If the enzyme is present in an un-
Hormones. The hormones that influence
phosphorylated form [10a], it acts as a kinase
carbohydrate metabolism include the pepti-
and leads to the formation of Fru-2,6-bP. After
des insulin and glucagon; a glucocorticoid,
phosphorylation by cAMP-dependent protein
cortisol; and a catecholamine, epinephrine
kinase A (PK-A), it acts as a phosphatase [10b]
(see p. 380). Insulin activates glycogen
and now catalyzes the degradation of Fru-2,6-
synthase ([1]; see p. 388), and induces several
bP to fructose 6-phosphate. The equilibrium
enzymes involved in glycolysis [3, 5, 7]. At the
between [10a] and [10b] is regulated by hor-
same time, insulin inhibits the synthesis of
mones. Epinephrine and glucagon increase
enzymes involved in gluconeogenesis
the cAMP level (see p. 120). As a result of
(repression; [4, 6, 8, 9]). Glucagon, the antag-
increased PK-A activity, this reduces the Fru-
onist of insulin, has the opposite effect. It
2,6-bP concentration and inhibits glycolysis,
induces gluconeogenesis enzymes [4, 6, 8, 9]
while at the same time activating gluconeo-
and represses pyruvate kinase [7], a key en-
genesis. Conversely, via [10a], insulin acti-
zyme of glycolysis. Additional effects of glu-
vates the synthesis of Fru-2,6-bP and thus
cagon are based on the interconversion of en-
glycolysis. In addition, insulin also inhibits
zymes and are mediated by the second mes-
the action of glucagon by reducing the cAMP
senger cAMP. This inhibits glycogen synthesis
level (see p. 120).
[1] and activates glycogenolysis
[2].
Epinephrine acts in a similar fashion. The in-
hibition of pyruvate kinase [7] by glucagon is
also due to interconversion.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrate Metabolism
159
A. Regulation of carbohydrate metabolism
Epinephrine
Glycogen
Epinephrine
Glucagon
Glucagon
Insulin
1
2
AMP
Fru-2,6-bis P
ATP
Insulin
Insulin
AMP
Citrate
Glucose
1-phosphate
3
5
3'
Glucose
Fructose
Fructose 1,6-
Glucose
6-phosphate
6-phosphate
bisphosphate
4
6
R
Insulin
R
Cortisol
Cortisol
Fru 2,6-bis P
Insulin
Glucagon
Glucagon
AMP
Glucagon
ATP
Acetyl CoA
1
Glycogen synthase 2.4.1.11
Insulin
R
2
Glycogen phosphorylase 2.4.1.1
3
Hexokinase 2.7.1.1
3'
Hexokinase (liver) 2.7.1.1
PEP
7
Pyruvate
4
Glucose 6-phosphatase 3.1.3.9
Acetyl
R
Insulin
R
CoA
5
6-Phosphofructokinase 2.7.1.11
9
8
6
Fructose 1,6-bisphosphatase 3.1.3.11
Cortisol
7
Pyruvate kinase 2.7.1.40
Glucagon
8
Pyruvate carboxylase 6.4.1.1
Oxalo-
Oxalo-
9
PEP carboxykinase (GTP) 4.1.1.32
acetate
acetate
B. Fructose 2,6-bisphosphate
Insulin
Citrate, PEP
5
10a
P
ATP
Fructose 6-
Fructose 2,6-
Fructose 1,6-
phosphate
bisphosphate
bisphosphate
ADP
10b
6
Insulin
cAMP
Glucagon
10a
6-Phosphofructo-2-kinase 2.7.1.105
10b
Fructose 2,6-bisphosphatase 3.1.3.46
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
160
Metabolism
sensitive lipases (see p. 162) and prevents the
Diabetes mellitus
breakdown of muscle protein.
Diabetes mellitus is a very common metabolic
The effects of insulin deficiency on metab-
disease that is caused by absolute or relative
olism are shown by arrows in the illustration.
insulin deficiency. The lack of this peptide
Particularly noticeable is the increase in the
hormone (see p. 76) mainly affects carbohy-
glucose concentration in the blood, from
drate and lipid metabolism. Diabetes mellitus
5 mM to 9 mM (90 mg dL-1) or more
occurs in two forms. In type 1 diabetes (in-
(hyperglycemia, elevated blood glucose
sulin-dependent diabetes mellitus, IDDM),
level). In muscle and adipose tissue - the two
the insulin-forming cells are destroyed in
most important glucose consumers—glucose
young individuals by an autoimmune reac-
uptake and glucose utilization are impaired
tion. The less severe type 2 diabetes (non-
by insulin deficiency. Glucose utilization in
insulin-dependent diabetes mellitus, NIDDM)
the liver is also reduced. At the same time,
usually has its first onset in elderly individu-
gluconeogenesis is stimulated, partly due to
als. The causes have not yet been explained in
increased proteolysis in the muscles. This in-
detail in this type.
creases the blood sugar level still further.
When the capacity of the kidneys to resorb
glucose is exceeded (at plasma concentrations
A. Insulin biosynthesis
of 9 mM or more), glucose is excreted in the
Insulin is produced by the B cells of the islets
urine (glucosuria).
of Langerhans in the pancreas. As is usual with
The increased degradation of fat that oc-
secretory proteins, the hormone’s precursor
curs in insulin deficiency also has serious ef-
(preproinsulin) carries a signal peptide that
fects. Some of the fatty acids that accumulate
directs the peptide chain to the interior of
in large quantities are taken up by the liver
the endoplasmic reticulum (see p. 210). Proin-
and used for lipoprotein synthesis (hyperlipi-
sulin is produced in the ER by cleavage of the
demia), and the rest are broken down into
signal peptide and formation of disulfide
acetyl CoA. As the tricarboxylic acid cycle is
bonds. Proinsulin passes to the Golgi appara-
not capable of taking up such large quantities
tus, where it is packed into vesicles—the β-
of acetyl CoA, the excess is used to form ke-
granules. After cleavage of the C peptide, ma-
tone bodies
(acetoacetate and E-hydroxy-
ture insulin is formed in the β-granules and is
butyrate see p. 312). As H+ ions are released
stored in the form of zinc-containing hexam-
in this process, diabetics not receiving ad-
ers until secretion.
equate treatment can suffer severe metabolic
acidosis (diabetic coma). The acetone that is
also formed gives these patients’ breath a
B. Effects of insulin deficiency
characteristic odor. In addition, large amounts
The effects of insulin on carbohydrate
of ketone body anions appear in the urine
metabolism are discussed on p. 158. In sim-
(ketonuria).
plified terms, they can be described as stim-
Diabetes mellitus can have serious secon-
ulation of glucose utilization and inhibition of
dary effects. A constantly raised blood sugar
gluconeogenesis. In addition, the transport of
level can lead in the long term to changes in
glucose from the blood into most tissues is
the blood vessels (diabetic angiopathy), kid-
also insulin-dependent (exceptions to this in-
ney damage (nephropathy) and damage to
clude the liver, CNS, and erythrocytes).
the nervous system (neuropathy), as well as
The lipid metabolism of adipose tissue is
to cataracts in the eyes.
also influenced by the hormone. In these cells,
insulin stimulates the reorganization of glu-
cose into fatty acids. This is mainly based on
activation of acetyl CoA carboxylase
(see
p. 162) and increased availability of
NADPH+H+ due to increased PPP activity
(see p. 152). On the other hand, insulin also
inhibits the degradation of fat by hormone-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Carbohydrate Metabolism
161
A. Insulin biosynthesis
B cells
Preproinsulin
1
Proinsulin
2
Insulin
3
104 AS
84 AS
51 AS
Nucleus
H3N
4
Zn2
3
Exo-
1
cytosis
β-Granules
Rough
2
Signal
ER
Golgi apparatus
peptide
C peptide
Insulin
4
hexamer
OOC
B. Effects of insulin deficiency
Muscle and
Blood
Adipose
other insulin-
tissue
Fat
dependent tissues
Glucose
uptake
impaired
Glucose
C
Glucose
C
Glucose
Lipolysis
Amino
Protein
Fatty acids
acids
Proteo-
Maximal
lysis
resorption
capacity
Hyper-
exceeded
glycemia
Glucose
Amino
acids
Fatty acids
Hyper-
lipid-
Glyco-
Glucose
emia
gen
Lipo-
Kidney
proteins
Fatty
acids
Anions
of ketone
bodies
Urine
Acetyl
Ketone
Pyruvate
Liver
CoA
bodies
Glucose
Anions
H
of ketone
Metabolic
bodies
Insulin deficiency
acidosis
Water
Electrolytes
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
162
Metabolism
coenzyme and ATP derived from it by oxida-
Overview
tive phosphorylation. If acetyl CoA production
exceeds the energy requirements of the hepa-
A. Fat metabolism
tocytes—as is the case when there is a high
Fat metabolism in adipose tissue (top). Fats
level of fatty acids in the blood plasma (typical
(triacylglycerols) are the most important en-
in hunger and diabetes mellitus)—then the
ergy reserve in the animal organism. They are
excess is converted into ketone bodies (see
mostly stored in insoluble form in the cells of
p. 312). These serve exclusively to supply
adipose tissue—the adipocytes—where they
other tissues with energy.
are constantly being synthesized and broken
down again.
Fat synthesis in the liver (right). Fatty acids
and fats are mainly synthesized in the liver
As precursors for the biosynthesis of fats
and in adipose tissue, as well as in the kid-
(lipogenesis), the adipocytes use triacylgly-
neys, lungs, and mammary glands. Fatty acid
cerols from lipoproteins (VLDLs and chylomi-
biosynthesis occurs in the cytoplasm—in con-
crons; see p. 278), which are formed in the
trast to fatty acid degradation. The most im-
liver and intestines and delivered by the
portant precursor is glucose, but certain
blood. Lipoprotein lipase [1], which is located
amino acids can also be used.
on the inner surface of the blood capillaries,
The first step is carboxylation of acetyl CoA
cleaves these triacylglycerols into glycerol
to malonyl CoA. This reaction is catalyzed by
and fatty acids, which are taken up by the
acetyl-CoA carboxylase [5], which is the key
adipocytes and converted back into fats.
enzyme in fatty acid biosynthesis. Synthesis
The degradation of fats (lipolysis) is cata-
into fatty acids is carried out by fatty acid
lyzed in adipocytes by hormone-sensitive
synthase
[6]. This multifunctional enzyme
lipase
[2]—an enzyme that is regulated by
(see p. 168) starts with one molecule of ace-
various hormones by cAMP-dependent inter-
tyl-CoA and elongates it by adding malonyl
conversion (see p. 120). The amount of fatty
groups in seven reaction cycles until palmi-
acids released depends on the activity of this
tate is reached. One CO2 molecule is released
lipase; in this way, the enzyme regulates the
in each reaction cycle. The fatty acid therefore
plasma levels of fatty acids.
grows by two carbon units each time.
In the blood plasma, fatty acids are trans-
NADPH+H+ is used as the reducing agent
ported in free form—i. e., non-esterified. Only
and is derived either from the pentose phos-
short-chain fatty acids are soluble in the
phate pathway (see p. 152) or from isocitrate
blood; longer, less water-soluble fatty acids
dehydrogenase and malic enzyme reactions.
are transported bound to albumin.
The elongation of the fatty acid by fatty acid
synthase concludes at C16, and the product,
Degradation of fatty acids in the liver (left).
palmitate
(16:0), is released. Unsaturated
Many tissues take up fatty acids from the
fatty acids and long-chain fatty acids can arise
blood plasma in order to synthesize fats or
from palmitate in subsequent reactions. Fats
to obtain energy by oxidizing them. The me-
are finally synthesized from activated fatty
tabolism of fatty acids is particularly intensive
acids
(acyl CoA) and glycerol 3-phosphate
in the hepatocytes in the liver.
(see p. 170). To supply peripheral tissues,
The most important process in the degra-
fats are packed by the hepatocytes into lipo-
dation of fatty acids is E-oxidation—a meta-
protein complexes of the VLDL type and re-
bolic pathway in the mitochondrial matrix
leased into the blood in this form (see p. 278).
(see p. 164). Initially, the fatty acids in the
cytoplasm are activated by binding to coen-
zyme A into acyl CoA [3]. Then, with the help
of a transport system (the carnitine shuttle
[4]; see p. 164), the activated fatty acids enter
the mitochondrial matrix, where they are
broken down into acetyl CoA. The resulting
acetyl residues can be oxidized to CO2 in the
tricarboxylic acid cycle, producing reduced
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipid Metabolism
163
A. Fat metabolism
Lipoprotein
Acetyl-CoA
1
5
lipase 3.1.1.34
carboxylase
Epinephrine
Largest
6.4.1.2 [Biotin]
Hormone-sensitive
2
Norepinephrine
Adipocyte
energy
lipase 3.1.1.3
Fatty acid
Glucagon etc.
store
6
synthase
Fatty acid-CoA
3
2.3.1.85
Triacylglycerols
ligase 6.2.1.3
Carnitine O-
Insulin
2
4
palmitoyltrans-
COO
ferase 2.3.1.21
Free fatty acids
From the
intestine
1
Insulin
COO
Chylo-
microns
Fatty acid-albumin complex
Blood
VLDL
Hepatocyte
Free fatty acids
CoA
ATP
3
Triacylglycerols
AMP+
P
P
Acyl CoA
Fat biosynthesis
4
Mitochondrion
Acyl CoA
Acyl CoA
Unsaturated
Elongated
fatty acids
fatty acids
NADH
Palmitate
β-Oxidation
+H
NADP
CO2
7x
6
Acetyl CoA
Acetyl CoA
NADPH
or acyl CoA
+H
Malonyl CoA
ADP+
P
Ketone bodies
Citrate
Insulin
5
Glucagon
ATP
HCO3
Citric acid
cycle
Acetyl CoA
Respiratory
chain
Amino acid
Glycolysis
degradation
Supply
of energy
to other
tissues
CO2+ATP
Amino acids
Glucose
Most important
Fatty acid
Fatty acid
precursor
degradation
biosynthesis
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
164
Metabolism
[4] β-Ketoacyl-CoA is now broken down by
Fatty acid degradation
an acyl transferase into acetyl CoA and an acyl
CoA shortened by 2 C atoms (“thioclastic
A. Fatty acid degradation: E-oxidation
cleavage”).
After uptake by the cell, fatty acids are
Several cycles are required for complete
activated by conversion into their CoA deriva-
degradation of long-chain fatty acids—eight
tives—acyl CoA is formed. This uses up two
cycles in the case of stearyl-CoA (C18:0), for
energy-rich anhydride bonds of ATP per fatty
example. The acetyl CoA formed can then
acid (see p. 162). For channeling into the mi-
undergo further metabolism in the tricarbox-
tochondria, the acyl residues are first trans-
ylic acid cycle (see p. 136), or can be used for
ferred to carnitine and then transported
biosynthesis. When there is an excess of ace-
across the inner membrane as acyl carnitine
tyl CoA, the liver can also form ketone bodies
(see B).
(see p. 312).
The degradation of the fatty acids occurs in
When oxidative degradation is complete,
the mitochondrial matrix through an oxida-
one molecule of palmitic acid supplies around
tive cycle in which C2 units are successively
106 molecules of ATP, corresponding to an
cleaved off as acetyl CoA (activated acetic
energy of 3300 kJ mol-1. This high energy
acid). Before the release of the acetyl groups,
yield makes fats an ideal form of storage for
each CH2 group at C-3 of the acyl residue (the
metabolic energy. Hibernating animals such
β-C atom) is oxidized to the keto group—
as polar bears can meet their own energy
hence the term
-oxidation for this metabolic
requirements for up to 6 months solely by
pathway. Both spatially and functionally, it is
fat degradation, while at the same time pro-
closely linked to the tricarboxylic acid cycle
ducing the vital water they need via the res-
(see p. 136) and to the respiratory chain (see
piratory chain (“respiratory water”).
p. 140).
B. Fatty acid transport
[1] The first step is dehydrogenation of acyl
CoA at C-2 and C-3. This yields an unsaturated
The inner mitochondrial membrane has a
∆2-enoyl-CoA derivative with a trans-config-
group-specific transport system for fatty
ured double bond. The two hydrogen atoms
acids. In the cytoplasm, the acyl groups of
are initially transferred from FAD-containing
activated fatty acids are transferred to carni-
acyl CoA dehydrogenase to the electron-trans-
tine by carnitine acyltransferase [1]. They are
ferring flavoprotein (ETF). ETF dehydrogenase
then channeled into the matrix by an acylcar-
[5] passes them on from ETF to ubiquinone
nitine/carnitine antiport as acyl carnitine, in
(coenzyme Q), a component of the respiratory
exchange for free carnitine. In the matrix, the
chain (see p. 140). Other FAD-containing mi-
mitochondrial enzyme carnitine acyltransfer-
tochondrial dehydrogenases are also able to
ase catalyzes the return transfer of the acyl
supply the respiratory chain with electrons in
residue to CoA.
this fashion.
The carnitine shuttle is the rate-determin-
There are three isoenzymes (see p. 98) of
ing step in mitochondrial fatty acid degrada-
acyl CoA dehydrogenase that are specialized
tion. Malonyl CoA, a precursor of fatty acid
for long-chain fatty acids (12-18 C atoms),
biosynthesis, inhibits carnitine acyltransferase
medium-chain fatty acids (4-14), and short-
(see p. 162), and therefore also inhibits uptake
chain fatty acids (4-8).
of fatty acids into the mitochondrial matrix.
[2] The next step in fatty acid degradation
The most important regulator of β-oxida-
is the addition of a water molecule to the
tion is the NAD+/NADH+H+ ratio. If the respi-
double bond of the enoyl CoA (hydration),
ratory chain is not using any NADH+H+, then
with formation of
-hydroxyacyl CoA.
not only the tricarboxylic acid cycle
(see
[3] In the next reaction, the OH group at C-
p. 136) but also β-oxidation come to a stand-
3 is oxidized to a carbonyl group (dehydro-
still due to the lack of NAD+.
genation). This gives rise to
-ketoacyl CoA,
and the reduction equivalents are transferred
to NAD+, which also passes them on to the
respiratory chain.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipid Metabolism
165
A. Fatty acid degradation: β-oxidation
Electron-
transferring
F
A
QH2
flavoprotein (ETF)
Respiratory
5
chain
β-Carbon
F
A
1
Q
H
H
H
O
C C C
C
S
H
O
A
H
H
H
C C
S
A
Acyl CoA
H
Acyl CoA
S
(n - 2 carbons)
4
A
H
O
H
C
C
S
A
H
O
H
H
C C
S
A
H
O
H
H
O
H
C
C
S
H
O
H O
H
C C
S
A
H
H
O
H
A
C C C
C
S
H
C
C
C
C
S
A
Acetyl CoA
A
H
H
H
H
H2O
N
A
Tricarboxylic acid cycle
2
3
1
Acyl-CoA dehydrogenase 1.3.99.3
H
OH
H O
2
Enoyl-CoA hydratase 4.2.1.17
N
A
Respiratory
C C C
C
S
chain
3-Hydroxyacyl-CoA dehydrogenase
A
3
H
H
H
1.1.1.35
4
Acetyl-CoA acyltransferase 2.3.1.16
5
ETF dehydrogenase [FAD, Fe4S4] 1.5.5.1
B. Fatty acid transport
Carnitine
O
1
Acyl
O-palmitoyltransferase
C
carnitine
2.3.1.21
O
CH3
Fat degradation
β-Oxidation
OOC
CH2
C
CH2
N CH3
H
CH3
S
Acyl
Acyl
S
A
Carnitine
Carnitine
1
1
S
S
Acyl
Acyl
A
carnitine
carnitine
A
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
166
Metabolism
boxylic acid cycle and is available for gluco-
Minor pathways of fatty acid
neogenesis through conversion into oxaloace-
degradation
tate. Odd-numbered fatty acids from pro-
pionyl-CoA can therefore be used to synthe-
Most fatty acids are saturated and even-num-
size glucose.
bered. They are broken down via E-oxidation
This pathway is also important for rumi-
(see p.164). In addition, there are special path-
nant animals, which are dependent on sym-
ways involving degradation of unsaturated
biotic microorganisms to break down their
fatty acids
(A), degradation of fatty acids
food. The microorganisms produce large
with an odd number of C atoms (B), α and ω
amounts of propionic acid as a degradation
oxidation of fatty acids, and degradation in
product, which the host can channel into the
peroxisomes.
metabolism in the way described.
A. Degradation of unsaturated fatty acids
Further information
Unsaturated fatty acids usually contain a cis
In addition to the degradation pathways de-
double bond at position 9 or 12—e. g., linoleic
scribed above, there are also additional spe-
acid (18:2; 9,12). As with saturated fatty acids,
cial pathways for particular fatty acids found
degradation in this case occurs via β-oxida-
in food.
tion until the C-9-cis double bond is reached.
a Oxidation is used to break down methyl-
Since enoyl-CoA hydratase only accepts sub-
branched fatty acids. It takes place through
strates with trans double bonds, the corre-
step-by-step removal of C1 residues, begins
sponding enoyl-CoA is converted by an iso-
with a hydroxylation, does not require coen-
merase from the cis-∆3, cis- ∆6 isomer into the
zyme A, and does not produce any ATP.
trans-∆3,cis-∆6 isomer [1]. Degradation by β-
w Oxidation—i. e., oxidation starting at the
oxidation can now continue until a shortened
end of the fatty acid—also starts with a hy-
trans-∆2, cis-∆4 derivative occurs in the next
droxylation catalyzed by a monooxygenase
cycle. This cannot be isomerized in the same
(see p. 316), and leads via subsequent oxida-
way as before, and instead is reduced in an
tion to fatty acids with two carboxyl groups,
NADPH-dependent way to the trans-∆3 com-
which can undergo β-oxidation from both
pound [2]. After rearrangement by enoyl-CoA
ends until C8 or C6 dicarboxylic acids are
isomerase [1], degradation can finally be com-
reached, which can be excreted in the urine
pleted via normal β-oxidation.
in this form.
Degradation of unusually long fatty acids.
B. Degradation of oddnumbered fatty acids
An alternative form of β-oxidation takes place
in hepatic peroxisomes, which are specialized
Fatty acids with an odd number of C atoms are
for the degradation of particularly long fatty
treated in the same way as “normal” fatty
acids (n > 20). The degradation products are
acids—i. e., they are taken up by the cell with
acetyl-CoA and hydrogen peroxide (H2O2),
ATP-dependent activation to acyl CoA and are
which is detoxified by the catalase
(see
transported into the mitochondria with the
p. 32) common in peroxisomes.
help of the carnitine shuttle and broken
down there by β−oxidation (see p. 164). In
Enzyme defects are also known to exist in
the last step, propionyl CoA arises instead of
the minor pathways of fatty acid degradation.
acetyl CoA. This is first carboxylated by pro-
In Refsum disease, the methyl-branched phy-
pionyl CoA carboxylase into (S)-methylmalonyl
tanic acid (obtained from vegetable foods)
CoA [3], which—after isomerization into the
cannot be degraded by α-oxidation. In Zell-
(R) enantiomer (not shown; see p. 411)—is
weger syndrome, a peroxisomal defect means
isomerized into succinyl CoA [4].
that long-chain fatty acids cannot be de-
Various coenzymes are involved in these
graded.
reactions. The carboxylase [3] requires biotin,
and the mutase [4] is dependent on coenzyme
B12 (5 -deoxyadenosyl cobalamin; see p. 108).
Succinyl-CoA is an intermediate in the tricar-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipid Metabolism
167
A. Degradation of unsaturated fatty acids
Linoleoyl CoA (18 : 2; 9,12)
O
9
C
S
1
2
A
Shift and
Reduction and
12
izomerization
shift of the
cis
3 CoA
of the
marked
marked
double bonds
double bond
β-Oxidation
3 Acetyl CoA
O
O
O
3
S
S
S
C
2
C
2
C
A
A
A
6
4
trans
trans
cis
β-Oxidation
6
1
cis
cis
CoA
Acetyl CoA
NADPH
N
A
O
O
P
S
S
2
2
C
C
4 CoA
A
A
N
A
3
P
trans
trans
1
NADP
β-Oxidation
Enoyl-CoA isomerase
1
5.3.3.8
5 Acetyl CoA
2,4-Dienoyl-CoA reductase
2
1.3.1.34
B. Degradation of odd-numbered fatty acids
Propionyl-CoA carboxylase
Odd-numbered
3
6.4.1.3 [biotin]
fatty acids
Methylmalonyl-CoA mutase
4
5.4.99.2 [cobamide]
H
O
β-Oxidation
H C C
S
A
A
A
P P
P
Tricarboxylic acid cycle
P P
H
P
n Acetyl CoA
H
H
O
H
H
O
O
H
H
H
C C C
S
3
H
C
C C
S
4
S
C C
C H
A
A
A
S
H
H
A
H
COO
H COO
Propionyl CoA
CO2
Methylmalonyl CoA
Succinyl CoA
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
168
Metabolism
ductase [6]. Finally, domain 3 serves to release
Fatty acid synthesis
the finished product by acyl-[ACP]-hydrolase
In the vertebrates, biosynthesis of fatty acids
[7] after seven steps of chain elongation.
is catalyzed by fatty acid synthase, a multi-
functional enzyme. Located in the cytoplasm,
B. Reactions of fatty acid synthase
the enzyme requires acetyl CoA as a starter
molecule. In a cyclic reaction, the acetyl resi-
The key enzyme in fatty acid synthesis is ace-
due is elongated by one C2 unit at a time for
tyl CoA carboxylase (see p. 162), which pre-
seven cycles. NADPH+H+ is used as a reducing
cedes the synthase and supplies the malonyl-
agent in the process. The end product of the
CoA required for elongation. Like all carbox-
reaction is the saturated C16 acid, palmitic
ylases, the enzyme contains covalently bound
acid.
biotin as a prosthetic group and is hormone-
dependently inactivated by phosphorylation
or activated by dephosphorylation
(see
A. Fatty acid synthase
p. 120). The precursor citrate (see p. 138) is
Fatty acid synthase in vertebrates consists of
an allosteric activator, while palmitoyl-CoA
two identical peptide chains—i. e., it is a ho-
inhibits the end product of the synthesis
modimer. Each of the two peptide chains,
pathway.
which are shown here as hemispheres, cata-
[1] The first cycle (n = 1) starts with the
lyzes all seven of the partial reactions re-
transfer of an acetyl residue from acetyl CoA
quired to synthesize palmitate. The spatial
to the peripheral cysteine residue (Cys-SH). At
compression of several successive reactions
the same time,
into a single multifunctional enzyme has ad-
[2] a malonyl residue is transferred from
vantages in comparison with separate en-
malonyl CoA to 4-phosphopantetheine (Pan-
zymes. Competing reactions are prevented,
SH).
the individual reactions proceed in a coordi-
[3] By condensation of the acetyl resi-
nated way as if on a production line, and due
due—or (in later cycles) the acyl residue—with
to low diffusion losses they are particularly
the malonyl group, with simultaneous decar-
ef cient.
boxylation, the chain is elongated.
Each subunit of the enzyme binds acetyl
[4]-[6] The following three reactions (re-
residues as thioesters at two different SH
duction of the 3-oxo group, dehydrogenation
groups: at one peripheral cysteine residue
of the 3-hydroxyl derivative, and renewed
(CysSH) and one central
4 -phosphopante-
reduction of it) correspond in principle to a
theine group (Pan-SH). Pan-SH, which is very
reversal of β-oxidation, but they are catalyzed
similar to coenzyme A (see p. 12), is cova-
by other enzymes and use NADPH+H+ instead
lently bound to a protein segment of the syn-
of NADH+H+ for reduction. They lead to an
thase known as the acyl-carrier protein (ACP).
acyl residue bound at Pan-SH with 2n + 2 C
This part functions like a long arm that passes
atoms (n = the number of the cycle). Finally,
the substrate from one reaction center to the
depending on the length of the product,
next. The two subunits of fatty acid synthase
[1 ] The acyl residue is transferred back to
cooperate in this process; the enzyme is
the peripheral cysteine, so that the next cycle
therefore only capable of functioning as a
can begin again with renewed loading of the
dimer.
ACP with a malonyl residue, or:
Spatially, the enzyme activities are ar-
[7] After seven cycles, the completed pal-
ranged into three different domains.
mitic acid is hydrolytically released.
Domain 1 catalyzes the entry of the substrates
acetyl CoA and malonyl CoA by
[ACP]-S-
In all, one acetyl-CoA and seven malonyl-
acetyltransferase
[1] and
[ACP]-Smalonyl
CoA are converted with the help of
14
transferase [2] and subsequent condensation
NADPH+H+ into one palmitic acid, 7 CO2,
of the two partners by
3-oxoacyl-[ACP]-
6 H2O, 8 CoA and 14 NADP+. Acetyl CoA car-
synthase [3]. Domain 2 catalyzes the conver-
boxylase also uses up seven ATP.
sion of the 3-oxo group to a CH2 group by 3-
oxoacyl-[ACP]-reductase
[4],
3-hydroxyacyl-
[ACP]-dehydratase
[5], and enoyl-[ACP]-re-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipid Metabolism
169
A. Fatty acid synthase
B. Reactions of fatty acid synthesis
1
2
Substrate entry
5
Water cleavage
1
7
Palmitate
3
Chain elongation
6
Reduction
H2O
Product
release
4
Reduction
7
Product release
O
H
H
H
Pan-S
C
C
C
C H
Acyl-
Palmitate
Cys-SH
H H H
N
A
6
NADP
5
P
6
1
2
4
N
A
7
ACP
NADP+H
P
-Pantethein
P
3
O
H
H
Cys
SH
SH
SH
Cys
SH
trans-Enoyl-
Pan-S
C
C
C C H
3
ACP
Cys-SH
H H
7
4
2
1
H2O
5
6
5
O
H
OH
H
Pan-S
3-Hydroxy-
C
C
C C H
Acetyl
S
acyl-
A
Cys-SH
H
H H
Malonyl
S
N
A
A
NADP
P
4
N
A
P
NADP+H
[ACP]-S-Acetyl-
1
transferase 2.3.1.38
O
H
O
H
[ACP]-S-Malonyl-
2
3-Oxoacyl-
Pan-S
C
C
C C H
transferase 2.3.1.39
Cys-SH
H
H
3-Oxoacyl-[ACP]
3
7x
synthase 2.3.1.41
CO2
3
3-Oxoacyl-[ACP]
4
reductase 1.1.1.100
H
O
H
3-Hydroxypalmitoyl-[ACP]
5
Pan-S
C
C
COO
dehydratase 4.2.1.61
H
Enoyl-[ACP]
6
reductase (NADPH) 1.3.1.10
O
H
S
A
7
Acyl-[ACP]
S
Cys-S
C
C
H
hydrolase 3.1.2.14
A
2
H
1
Acetyl or
Malonyl
S
Starting reaction
Acetyl
S
acyl residue
A
A
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
170
Metabolism
The biosynthesis of most phospholipids
Biosynthesis of complex lipids
also starts from DAG.
[7] Transfer of a phosphocholine residue to
A. Biosynthesis of fats and phospholipids
the free OH group gives rise to phosphatidyl-
Complex lipids, such as neutral fats (triacyl-
choline
(lecithin; enzyme:
1-alkyl-2-acetyl-
glycerols), phospholipids, and glycolipids, are
glycerolcholine phosphotransferase
2.7.8.16).
synthesized via common reaction pathways.
The phosphocholine residue is derived from
Most of the enzymes involved are associated
the precursor CDP-choline (see p. 110). Phos-
with the membranes of the smooth endoplas-
phatidylethanolamine is similarly formed
mic reticulum.
from CDP-ethanolamine and DAG. By
contrast, phosphatidylserine is derived from
The synthesis of fats and phospholipids
phosphatidylethanolamine by an exchange of
starts with glycerol 3-phosphate. This com-
the amino alcohol. Further reactions serve to
pound can arise via two pathways:
interconvert the phospholipids—e. g., phos-
[1] By reduction from the glycolytic
phatidylserine can be converted into phos-
intermediate glycerone 3-phosphate (dihy-
phatidylethanolamine by decarboxylation,
droxyacetone
3-phosphate; enzyme: glyc-
and the latter can then be converted into
erol-3-phosphate dehydrogenase
(NAD+)
phosphatidylcholine by methylation with S-
1.1.1.8), or:
adenosyl methionine (not shown; see also
[2] By phosphorylation of glycerol deriving
p. 409). The biosynthesis of phosphatidylino-
from fat degradation (enzyme: glycerol kinase
sitol starts from phosphatidate rather than
2.7.1.30).
DAG.
[3] Esterification of glycerol 3-phosphate
[8] In the lumen of the intestine, fats from
with a long-chain fatty acid produces a
food are mainly broken down into monoacyl-
strongly amphipathic lysophosphatidate (en-
glycerols (see p. 270). The cells of the intesti-
zyme: glycerol-3-phosphate acyltransferase
nal mucosa re-synthesize these into neutral
2.3.1.15). In this reaction, an acyl residue is
fats. This pathway also passes via DAG
transferred from the activated precursor
(enzyme: acylglycerolpalmitoyl transferase
acyl-CoA to the hydroxy group at C-1.
2.3.1.22).
[4] A second esterification of this type leads
[9] Transfer of a CMP residue gives rise first
to a phosphatidate (enzyme: 1-acylglycerol-3-
to CDP-diacylglycerol (enzyme: phosphatida-
phosphate acyltransferase
2.3.1.51). Unsatu-
tecytidyl transferase 2.3.1.22).
rated acyl residues, particularly oleic acid,
[10] Substitution of the CMP residue by
are usually incorporated at C-2 of the glycerol.
inositol then provides phosphatidylinositol
Phosphatidates (anions of phosphatidic acids)
(PtdIns; enzyme: CDPdiacylglycerolinositol-3-
are the key molecules in the biosynthesis of
phosphatidyl transferase 2.7.8.11).
fats, phospholipids, and glycolipids.
[12] An additional phosphorylation (en-
[5] To biosynthesize fats (triacylglycerols),
zyme: phosphatidylinositol-4-phosphate kin-
the phosphate residue is again removed by
ase 2.7.1.68) finally provides phosphaditylino-
hydrolysis (enzyme: phosphatidate phospha-
sitol-4,5-bisphosphate
(PIP2, PtdIns(4,5)P2).
tase
3.1.3.4). This produces diacylglycerols
PIP2 is the precursor for the second messen-
(DAG).
gers
2,3-diacylglycerol
(DAG) and inositol-
[6] Transfer of an additional acyl residue to
1,4,5-trisphosphate (InsP3, IP3; see p. 367).
DAG forms triacylglycerols (enzyme: diacyl-
The biosynthesis of the sphingolipids is
glycerol acyltransferase
2.3.1.20). This com-
shown in schematic form on p. 409.
pletes the biosynthesis of neutral fats. They
are packaged into VLDLs by the liver and re-
leased into the blood. Finally, they are stored
by adipocytes in the form of insoluble fat
droplets.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipid Metabolism
171
A. Biosynthesis of fats and phospholipids
Triacylglycerol
Phosphatidylcholine
P
Inositol
P P
Phosphatidylinositol 4,5-bisphosphate
P
Choline
S
A
ADP
6
12
S
ATP
A
C
P
7
or
ethanol-
P
Inositol
C
amine
P
P P
Choline
Phosphatidylinositol 4-phosphate
Diacylglycerol
S
P
A
ADP
11
5
8
ATP
S
A
P
Inositol
P
Phosphatidylinositol
Phosphatidate
CMP
P
P
9
S
A
2-Monoacylglycerol
10
4
Inositol
C
P P P
S
A
Acyl CoA
C
P
P
P
Food
CDP-diacylglycerol
Lysophosphatidate
S
A
Glycolysis
3
S
A
Acyl CoA
N
A
N
A
O
1
2
P
P
A
A
P P
P P P
Glycerone 3-phosphate
Glycerol
Glycerol
3-phosphate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
172
Metabolism
(effectors: hormones). Insulin and thyroxine
Biosynthesis of cholesterol
stimulate the enzyme and glucagon inhibits it
Cholesterol is a major constituent of the cell
by cAMP-dependent phosphorylation. A large
membranes of animal cells (see p. 216). It
supply of cholesterol from food also inhibits
would be possible for the body to provide its
3-HMG-CoA reductase.
full daily cholesterol requirement (ca. 1 g) by
(2) Formation of isopentenyl diphosphate.
synthesizing it itself. However, with a mixed
After phosphorylation, mevalonate is decar-
diet, only about half of the cholesterol is de-
boxylated to isopentenyl diphosphate, with
rived from endogenous biosynthesis, which
consumption of ATP. This is the component
takes place in the intestine and skin, and
from which all of the isoprenoids are built
mainly in the liver (about 50%). The rest is
(see p. 53).
taken up from food. Most of the cholesterol
(3) Formation of squalene. Isopentenyl
is incorporated into the lipid layer of plasma
diphosphate undergoes isomerization to
membranes, or converted into bile acids (see
form dimethylallyl diphosphate. The two C5
p. 314). A very small amount of cholesterol is
molecules condense to yield geranyl diphos-
used for biosynthesis of the steroid hormones
phate, and the addition of another isopen-
(see p. 376). In addition, up to 1 g cholesterol
tenyl diphosphate produces farnesyl diphos-
per day is released into the bile and thus
phate. This can then undergo dimerization, in
excreted.
a head-to-head reaction, to yield squalene.
Farnesyl diphosphate is also the starting-
point for other polyisoprenoids, such as doli-
A. Cholesterol biosynthesis
chol (see p. 230) and ubiquinone (see p. 52).
Cholesterol is one of the isoprenoids, synthe-
(4) Formation of cholesterol. Squalene, a
sis of which starts from acetyl CoA (see p. 52).
linear isoprenoid, is cyclized, with O2 being
In a long and complex reaction chain, the C27
consumed, to form lanosterol, a C30 sterol.
sterol is built up from C2 components. The
Three methyl groups are cleaved from this
biosynthesis of cholesterol can be divided
in the subsequent reaction steps, to yield the
into four sections. In the first
(1),
end product cholesterol. Some of these reac-
mevalonate, a C6
compound, arises from
tions are catalyzed by cytochrome P450 sys-
three molecules of acetyl CoA. In the second
tems (see p. 318).
part (2), mevalonate is converted into isopen-
The endergonic biosynthetic pathway de-
tenyl diphosphate, the “active isoprene.” In
scribed above is located entirely in the smooth
the third part (3), six of these C5 molecules
endoplasmic reticulum. The energy needed
are linked to produce squalene, a C30 com-
comes from the CoA derivatives used and
pound. Finally, squalene undergoes cycliza-
from ATP. The reducing agent in the formation
tion, with three C atoms being removed, to
of mevalonate and squalene, as well as in the
yield cholesterol
(4). The illustration only
final steps of cholesterol biosynthesis, is
shows the most important intermediates in
NADPH+H+.
biosynthesis.
The division of the intermediates of the
(1) Formation of mevalonate. The conver-
reaction pathway into three groups is charac-
sion of acetyl CoA to acetoacetyl CoA and then
teristic: CoA compounds, diphosphates, and
to
3-hydroxy-3-methylglutaryl CoA
(3-HMG
highly lipophilic, poorly soluble compounds
CoA) corresponds to the biosynthetic path-
(squalene to cholesterol), which are bound to
way for ketone bodies (details on p. 312). In
sterol carriers in the cell.
this case, however, the synthesis occurs not in
the mitochondria as in ketone body synthesis,
but in the smooth endoplasmic reticulum. In
the next step, the 3-HMG group is cleaved
from the CoA and at the same time reduced
to mevalonate with the help of NADPH+H+. 3-
HMG CoA reductase is the key enzyme in cho-
lesterol biosynthesis. It is regulated by repres-
sion of transcription
(effectors: oxysterols
such as cholesterol) and by interconversion
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipid Metabolism
173
A. Cholesterol biosynthesis
Acetyl CoA
1
Mevalonate
2
Isopentenyl diphosphate
3
Squalene
4
Cholesterol
C2
C6
C
C30
C27
5
3x
1C
6x
3C
O
CH3
3x H3C C
S
Acetyl CoA
A
OOC
CH2
C CH
2
CH2OH
Mevalonate
OH
3-Hydroxy-
3-methyl-
A
2
P P P
glutaryl CoA
CH3
O
A
2
P P
OOC CH2
C CH2
C
S
A
OH
2
N
A
CH
3
P
OOC
CH2
C CH2
CH2
O P P
Insulin
OH
HMG-CoA
Mevalonyl
Thyroxin
R
Cholesterol
reductase
diphosphate
A
1.1.1.34
Glucagon
P
P
P
Key enzyme
A
N
A
CO2
P
P P
S
2
A
P
CH3
CH3
C
OOC
CH2
C CH2
CH2OH
H2C
O
P P
OH
Mevalonate
Isopentenyl diphosphate
1
2
3
4
Dimethylallyl
Isopentenyl diphosphate
diphosphate
Squalene
O
P
P
O
P P
O2 +
N
A
P
Two steps
N
A
H2O +
Geranyl
P
P P
diphosphate
O
P P
Lanosterol
Farnesyl
diphosphate
P P
HO
N
A
x O2 + x
P
O P P
2 CO2
N
A
x H2O + x
1 HCOOH
P
Multiple steps
NADPH+H
P P
NADP
Squalene
Cholesterol
HO
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
174
Metabolism
is reincorporated into proteins (protein bio-
Protein metabolism: overview
synthesis). The body’s high level of protein
Quantitatively, proteins are the most impor-
turnover is due to the fact that many proteins
tant group of endogenous macromolecules. A
are relatively short-lived. On average, their
person weighing 70 kg contains about 10 kg
half-lives amount to 2-8 days. The key en-
protein, with most of it located in muscle. By
zymes of the intermediary metabolism have
comparison, the proportion made up by other
even shorter half-lives. They are sometimes
nitrogencontaining compounds is minor. The
broken down only a few hours after being
organism’s nitrogen balance is therefore pri-
synthesized, and are replaced by new mole-
marily determined by protein metabolism.
cules. This constant process of synthesis and
Several hormones—mainly testosterone and
degradation makes it possible for the cells to
cortisol—regulate the nitrogen balance (see
quickly adjust the quantities, and therefore
p. 374).
the activity, of important enzymes in order
to meet current requirements. By contrast,
structural proteins such as the histones, he-
A. Protein metabolism: overview
moglobin, and the components of the cyto-
In adults, the nitrogen balance is generally in
skeleton are particularly long-lived.
equilibrium—i. e., the quantities of protein ni-
Almost all cells are capable of carrying out
trogen taken in and excreted per day are ap-
biosynthesis of proteins (top left). The forma-
proximately equal. If only some of the nitro-
tion of peptide chains by translation at the
gen taken in is excreted again, then the bal-
ribosome is described in greater detail on
ance is positive. This is the case during growth,
pp. 250-253. However, the functional forms
for example. Negative balances are rare and
of most proteins arise only after a series of
usually occur due to disease.
additional steps. To begin with, supported by
Proteins taken up in food are initially bro-
auxiliary proteins, the biologically active con-
ken down in the gastrointestinal tract into
formation of the peptide chain has to be
amino acids, which are resorbed and distrib-
formed (folding; see pp. 74,
232). During
uted in the organism via the blood
(see
subsequent “post-translational” maturation,
p. 266). The human body is not capable of
many proteins remove part of the peptide
synthesizing 8-10 of the 20 proteinogenic
chain again and attach additional groups—
amino acids it requires (see p. 60). These
e. g., oligosaccharides or lipids. These pro-
amino acids are essential, and have to be sup-
cesses take place in the endoplasmic reticu-
plied from food (see p. 184).
lum and in the Golgi apparatus (see p. 232).
Proteins are constantly being lost via the
Finally, the proteins have to be transported to
intestine and, to a lesser extent, via the kid-
their site of action (sorting; see p. 228).
neys. To balance these inevitable losses, at
Some intracellular protein degradation
least 30 g of protein have to be taken up
(proteolysis) takes place in the lysosomes
with food every day. Although this minimum
(see p. 234). In addition, there are protein
value is barely reached in some countries, in
complexes in the cytoplasm, known as pro-
the industrial nations the protein content of
teasomes, in which incorrectly folded or old
food is usually much higher than necessary.
proteins are degraded. These molecules are
As it is not possible to store amino acids, up to
recognized by a special marking (see p. 176).
100 g of excess amino acids per day are used
The proteasome also plays an important part
for biosynthesis or degraded in the liver in
in the presentation of antigens by immune
this situation. The nitrogen from this excess
cells (see p. 296).
is converted into urea (see p. 182) and ex-
creted in the urine in this form. The carbon
skeletons are used to synthesize carbohy-
drates or lipids (see p. 180), or are used to
form ATP.
It is thought that adults break down
300-400 g of protein per day into amino
acids (proteolysis). On the other hand, ap-
proximately the same amount of amino acids
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Protein Metabolism
175
A. Protein metabolism: overview
Functional
proteins
10 000 g
Maturation
Marking
Sugars
Folding
Degradation
Proteasome
Lysosome
Peptide
Translation
Degradation
Amino acid pool
100 g
Excess
amino
acids
Protein
Food
secretion
Glucose
Lipids
50-100 g/day
50 g/day
De-novo
Resorption
synthesis
Degradation
Up to 150 g/day
As required
50-100 g/day
2-Oxo-
Ketone
acids
bodies
Digestion
A
Pyruvate
NH3
P P P
Amino
Phosphoenolpyruvate
ATP
acids
3-Phosphoglycerate
2-Oxoacids
Urea
CO2, H2O
Sugar phosphates
Intestine
10g/day
Kidney
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
176
Metabolism
molecules) are marked by covalent linkage
Proteolysis
with chains of the small protein ubiquitin.
The ubiquitin is previously activated by the
A. Proteolytic enzymes
introduction of reactive thioester groups.
Combinations of several enzymes with differ-
Molecules marked with ubiquitin (“ubiquiti-
ent specificities are required for complete
nated”) are recognized by the 19S particle,
degradation of proteins into free amino
unfolded using ATP, and then shifted into
acids. Proteinases and peptidases are found
the interior of the nucleus, where degradation
not only in the gastrointestinal tract
(see
takes place. Ubiquitin is not degraded, but is
p. 268), but also inside the cell (see below).
reused after renewed activation.
The proteolytic enzymes are classified into
endopeptidases and exopeptidases, according
C. Serine proteases
to their site of attack in the substrate mole-
cule. The endopeptidases or proteinases cleave
A large group of proteinases contain serine in
peptide bonds inside peptide chains. They
their active center. The serine proteases in-
“recognize” and bind to short sections of the
clude, for example, the digestive enzymes
substrate’s sequence, and then hydrolyze
trypsin, chymotrypsin, and elastase
(see
bonds between particular amino acid residues
pp. 94 and 268), many coagulation factors
in a relatively specific way (see p. 94). The
(see p. 290), and the fibrinolytic enzyme plas-
proteinases are classified according to their
min and its activators (see p. 292).
reaction mechanism. In serine proteinases,
As described on p. 270, pancreatic protein-
for example (see C), a serine residue in the
ases are secreted as proenzymes (zymogens).
enzyme is important for catalysis, while in
Activation of these is also based on proteolytic
cysteine proteinases, it is a cysteine residue,
cleavages. This is illustrated here in detail us-
and so on.
ing the example of trypsinogen, the precursor
The exopeptidases attack peptides from
of trypsin (1). Activation of trypsinogen starts
their termini. Peptidases that act at the N
with cleavage of an N-terminal hexapeptide
terminus are known as aminopeptidases,
by enteropeptidase (enterokinase), a specific
while those that recognize the C terminus
serine proteinase that is located in the mem-
are called carboxypeptidases. The dipepti-
brane of the intestinal epithelium. The cleav-
dases only hydrolyze dipeptides.
age product (β-trypsin) is already catalytically
active, and it cleaves additional trypsinogen
molecules at the sites marked in red in the
B. Proteasome
illustration (autocatalytic cleavage). The pre-
The functional proteins in the cell have to be
cursors of chymotrypsin, elastase, and car-
protected in order to prevent premature deg-
boxypeptidase A, among others, are also acti-
radation. Some of the intracellularly active
vated by trypsin.
proteolytic enzymes are therefore enclosed
The active center of trypsin is shown in
in lysosomes (see p. 234). The proteinases
Fig. 2. A serine residue in the enzyme (Ser-
that act there are also known as cathepsins.
195), supported by a histidine residue and
Another carefully regulated system for pro-
an aspartate residue (His-57, Asp-102), nucle-
tein degradation is located in the cytoplasm.
ophilically attacks the bond that is to be
This consists of large protein complexes (mass
cleaved (red arrow). The cleavage site in the
2
106 Da), the proteasomes. Proteasomes
substrate peptide is located on the C-terminal
contain a barrel-shaped core consisting of 28
side of a lysine residue, the side chain of
subunits that has a sedimentation coef cient
which is fixed in a special “binding pocket”
(see p. 200) of
20 S.Proteolytic activity
of the enzyme (left) during catalysis
(see
(shown here by the scissors) is localized in
p. 94).
the interior of the 20-S core and is therefore
protected. The openings in the barrel are
sealed by 19-S particles with a complex struc-
ture that control access to the core.
Proteins destined for degradation in the
proteasome (e. g., incorrectly folded or old
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Protein Metabolism
177
A. Proteolytic enzymes
C-Terminus
COO
Serine proteinase
H3N
N-Terminus
3.4.21.n
Aminopeptidase
H3N
COO
Cysteine proteinase
Exopeptidase
[Zn2
] 3.4.11.n
3.4.22.n
Endopeptidase
Aspartate proteinase
3.4.23.n
Metalloproteinase
Dipeptidase
H3N
3.4.24.n
[Zn2
] 3.4.13.n
Carboxy-
peptidase
3.4.17.n
Amino acid residue
COO
B. Proteasome
C. Serine proteases
Folded
Activated
Disulfide Autocatalytic
protein
ubiquitin
bond
cleavage
Auto-
cata-
lytic
cleavage
Marking
Activation
with
of
ubiquitin
ubiquitin
Ubiquitinated
protein
Cleavage
by entero-
peptidase
Active
19 S
and trypsin
center
particle
1
Enteropeptidase
2
Trypsin
Binding
3.4.21.9
3.4.21.4
20 S
1. Trypsinogen activation
nucleus
19 S
particle
Asp-102
Ser-195
Lysine residue
A
His-57
Unfolding
Degradation
Substrate
A
2. Trypsin: active center
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
178
Metabolism
The ketimine (3) is hydrolyzed to yield the 2-
Transamination and deamination
oxoacid and pyridoxamine phosphate (4).
Amino nitrogen accumulates during protein
In the second part of the reaction (see A,
degradation. In contrast to carbon, amino ni-
1b), these steps take place in the opposite
trogen is not suitable for oxidative energy
direction: pyridoxamine phosphate and the
production. If they are not being reused for
second 2-oxoacid form a ketimine, which is
biosynthesis, the amino groups of amino acids
isomerized into aldimine. Finally, the second
are therefore incorporated into urea
(see
amino acid is cleaved and the coenzyme is
p. 182) and excreted in this form.
regenerated.
A. Transamination and deamination
C. NH3 metabolism in the liver
Among the NH2 transfer reactions, trans-
In addition to urea synthesis itself
(see
aminations
(1) are particularly important.
p. 182), the precursors NH3 and aspartate are
They are catalyzed by transaminases, and oc-
also mainly formed in the liver. Amino nitro-
cur in both catabolic and anabolic amino acid
gen arising in tissue is transported to the liver
metabolism. During transamination, the
by the blood, mainly in the form of glutamine
amino group of an amino acid (amino acid
(Gln) and alanine (Ala; see p. 338). In the liver,
1) is transferred to a 2-oxoacid (oxoacid 2).
Gln is hydrolytically deaminated by glutami-
From the amino acid, this produces a 2-oxo-
nase [3] into glutamate (Glu) and NH3. The
acid (a), while from the original oxoacid, an
amino group of the alanine is transferred by
amino acid is formed (b). The NH2 group is
alanine transaminase [1] to 2-oxoglutarate (2-
temporarily taken over by enzyme-bound
OG; formerly known as α-ketoglutarate). This
pyridoxal phosphate (PLP; see p. 106), which
transamination (A) produces another gluta-
thus becomes pyridoxamine phosphate.
mate. NH3 is finally released from glutamate
If the NH2 is released as ammonia, the
by oxidative deamination (A). This reaction is
process is referred to as deamination. There
catalyzed by glutamate dehydrogenase [4], a
are different mechanisms for this (see p. 180).
typical liver enzyme. Aspartate (Asp), the sec-
A particularly important one is oxidative
ond amino group donor in the urea cycle, also
deamination (2). In this reaction, the α-amino
arises from glutamate. The aspartate
group is initially oxidized into an imino group
transaminase [2] responsible for this reaction
(2a), and the reducing equivalents are trans-
is found with a high level of activity in the
ferred to NAD+ or NADP+. In the second step,
liver, as is alanine transaminase [1].
the imino group is then cleaved by hydrolysis.
Transaminases are also found in other tis-
As in transamination, this produces a 2-oxo-
sues, from which they leak from the cells into
acid (C). Oxidative deamination mainly takes
the blood when injury occurs. Measurement
place in the liver, where glutamate is broken
of serum enzyme activity (serum enzyme di-
down in this way into 2-oxoglutarate and
agnosis; see also p. 98) is an important
ammonia, catalyzed by glutamate dehydro-
method of recognizing and monitoring the
genase. The reverse reaction initiates biosyn-
course of such injuries. Transaminase activity
thesis of the amino acids in the glutamate
in the blood is for instance important for di-
family (see p. 184).
agnosing liver disease (e. g., hepatitis) and
myocardial disease (cardiac infarction).
B. Mechanism of transamination
In the absence of substrates, the aldehyde
group of pyridoxal phosphate is covalently
bound to a lysine residue of the transaminase
(1). This type of compound is known as an
aldimine or “Schiff’s base.” During the reac-
tion, amino acid 1 (A, 1a) displaces the lysine
residue, and a new aldimine is formed (2). The
double bond is then shifted by isomerization.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Protein Metabolism
179
A. Transamination and deamination
B. Mechanism of transamination
H
Amino
Lysine residue
acid
OOC C R
O
O
O
O
C
a
C
NH
3
H3N
C H
O
C
N
H
H C
R1
R1
H
Aldimine 2
O
Amino acid 1
Oxoacid 1
OOC C R
O
Pyridoxamine
P
PLP
N
H
H3N
phosphate
H3C
N
H C
1.
O
H
O
O
O
O
O
PLP (Aldimine 1)
C
C
P
H3C
N
O C
H3N
C H
b
2.
H
Aldimine 2
R2
R2
Oxoacid 2
Amino acid 2
Rearrangement
1. Transamination
OOC
R
Imino acid
O
O
C
Ketimine
O
O
C
a
C
N
H CH2
H3N
C H
H2N
C
O
R2
R2
O
N
A
N
A
P
Amino acid
P
P
b
H2O
H3C
N
NH2
H
CH2
3.
H2O
H
HO
O
O
O
NH4
C
P
Oxoacid
O
C
H3C
N
OOC
C
R
R2
H
4.
O
2. Oxidative deamination
Oxoacid
Pyridoxamine-P
C. NH3 metabolism in the liver
Alanine trans-
1
aminase [PLP]
2.6.1.2
Aspartate trans-
2
aminase [PLP]
2.6.1.1
Glutaminase
3
3.5.1.2
Glutamate
4
dehydrogenase
1.4.1.2
Transamination
Oxidative
deamination
Hydrolytic
deamination
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
180
Metabolism
and ketogenic. This group includes phenylala-
Amino acid degradation
nine, tyrosine, tryptophan, and isoleucine.
A large number of metabolic pathways are
Degradation of acetoacetate to acetyl CoA
available for amino acid degradation, and an
takes place in two steps (not shown). First,
overview of these is presented here. Further
acetoacetate and succinyl CoA are converted
details are given on pp. 414 and 415.
into acetoacetyl CoA and succinate (enzyme:
3-oxoacid-CoA transferase 2.8.3.5). Acetoacetyl
CoA is then broken down by β-oxidation into
A. Amino acid degradation : overview
two molecules of acetyl CoA (see p. 164),
During the degradation of most amino acids,
while succinate can be further metabolized
the α-amino group is initially removed by
via the tricarboxylic acid cycle.
transamination or deamination. Various
mechanisms are available for this, and these
B. Deamination
are discussed in greater detail in B. The carbon
skeletons that are left over after deamination
There are various ways of releasing ammonia
undergo further degradation in various ways.
(NH3) from amino acids, and these are illus-
During degradation, the 20 proteinogenic
trated here using the example of the amino
amino acids produce only seven different
acids glutamine, glutamate, alanine, and ser-
degradation products
(highlighted in pink
ine.
and violet). Five of these metabolites (2-oxo-
[1] In the branched-chain amino acids (Val,
glutarate, succinyl CoA, fumarate, oxaloace-
Leu, Ile) and also tyrosine and ornithine, deg-
tate, and pyruvate) are precursors for gluco-
radation starts with a transamination. For ala-
neogenesis and can therefore be converted
nine and aspartate, this is actually the only
into glucose by the liver and kidneys (see
degradation step. The mechanism of transa-
p. 154). Amino acids whose degradation sup-
mination is discussed in detail on p. 178.
plies one of these five metabolites are there-
[2] Oxidative deamination, with the forma-
fore referred to as glucogenic amino acids.
tion of NADH+H+, only applies to glutamate in
The first four degradation products listed are
animal metabolism. The reaction mainly takes
already intermediates in the tricarboxylic acid
place in the liver and releases NH3 for urea
cycle, while pyruvate can be converted into
formation (see p. 178).
oxaloacetate by pyruvate carboxylase and thus
[3] Two amino acids—asparagine and glu-
made available for gluconeogenesis (green
tamine—contain acid-amide groups in the
arrow).
side chains, from which NH3 can be released
With two exceptions (lysine and leucine;
by hydrolysis (hydrolytic deamination). In the
see below), all of the proteinogenic amino
blood, glutamine is the most important trans-
acids are also glucogenic. Quantitatively,
port molecule for amino nitrogen. Hydrolytic
they represent the most important precursors
deamination of glutamine in the liver also
for gluconeogenesis. At the same time, they
supplies the urea cycle with NH3.
also have an anaplerotic effect—i. e., they re-
[4] Eliminating deamination takes place in
plenish the tricarboxylic acid cycle in order to
the degradation of histidine and serine. H2O is
feed the anabolic reactions that originate in it
first eliminated here, yielding an unsaturated
(see p. 138).
intermediate. In the case of serine, this inter-
Two additional degradation products (ace-
mediate is first rearranged into an imine (not
toacetate and acetyl CoA) cannot be chan-
shown), which is hydrolyzed in the second
neled into gluconeogenesis in animal metab-
step into NH3 and pyruvate, with H2O being
olism, as there is no means of converting
taken up. H2O does not therefore appear in
them into precursors of gluconeogenesis.
the reaction equation.
However, they can be used to synthesize ke-
tone bodies, fatty acids, and isoprenoids.
Amino acids that supply acetyl CoA or aceto-
acetate are therefore known as ketogenic
amino acids. Only leucine and lysine are
purely ketogenic. Several amino acids yield
degradation products that are both glucogenic
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Protein Metabolism
181
A. Amino acid degradation: overview
Several
Alanine
Threonine
pathways
Glucose
Cysteine
Pyruvate
Serine
Glycine
Asparagine
Acetyl
Aceto-
CoA
acetate
Aspartate
Oxalo-
Ketone
acetate
body
Phenyl-
alanine
Pyruvate
Tricarboxylic
Fuma-
Tyrosine
rate
acid cycle
Leucine
Lysine
Iso-
2-Oxo-
leucine
Succinyl
glutarate
Tryptophan
Alanine
CoA
Valine
Acetyl
Glutamate
Histidine
CoA
Glutamine
Proline
Propionyl
CoA
Cysteine
Pyruvate
Methionine
Serine
Arginine
Glucogenic
Transamination
Hydrolytic deamination
Glucogenic
Ketogenic
and ketogenic
Oxidative deamination
Eliminating deamination
B. Deamination
COO
NADH+H
NAD
COO
COO
O
C
N
A
N
A
H3N C H
H3N C H
NH
4
(CH2)
(CH2)
2
(CH2)
2
2
COO
COO
CONH2
2
2-OG
Glu
3
Gln
1
NH4
H2O
COO
COO
H3N C
H
H3N
C
H
Ala
Pyr
4
Ser
C
H3
H2C
OH
COO
O C
C
H3
1
Alanine transaminase [PLP] 2.6.1.2
3
Glutaminase 3.5.1.2
2
Glutamate dehydrogenase 1.4.1.2
4
Serine dehydratase [PLP] 4.2.1.13
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
182
Metabolism
[1] In the first step, carbamoyl phosphate is
Urea cycle
formed in the mitochondria from hydrogen
Amino acids are mainly broken down in the
carbonate (HCO3-) and NH4+, with two ATP
liver. Ammonia is released either directly or
molecules being consumed. In this com-
indirectly in the process (see p. 178). The deg-
pound, the carbamoyl residue (-O-CO-NH2)
radation of nucleobases also provides signifi-
is at a high chemical potential. In hepatic
cant amounts of ammonia (see p. 186).
mitochondria, enzyme [1] makes up about
Ammonia (NH3) is a relatively strong base,
20% of the matrix proteins.
and at physiological pH values it is mainly
[2] In the next step, the carbamoyl residue
present in the form of the ammonium ion
is transferred to the non-proteinogenic amino
NH4+ (see p. 30). NH3 and NH4+ are toxic, and
acid ornithine, converting it into citrulline,
at higher concentrations cause brain damage
which is also non-proteinogenic. This is
in particular. Ammonia therefore has to be
passed into the cytoplasm via a transporter.
effectively inactivated and excreted. This can
[3] The second NH2 group of the later urea
be carried out in various ways. Aquatic ani-
molecule is provided by aspartate, which
mals can excrete NH4+ directly. For example,
condenses with citrulline into argininosucci-
fish excrete NH4+ via the gills (ammonotelic
nate. ATP is cleaved into AMP and diphos-
animals). Terrestrial vertebrates, including
phate (PPi) for this endergonic reaction. To
humans, hardly excrete any NH3, and instead,
shift the equilibrium of the reaction to the
most ammonia is converted into urea before
side of the product, diphosphate is removed
excretion (ureotelic animals). Birds and rep-
from the equilibrium by hydrolysis.
tiles, by contrast, form uric acid, which is
[4] Cleavage of fumarate from argininosuc-
mainly excreted as a solid in order to save
cinate leads to the proteinogenic amino acid
water (uricotelic animals).
arginine, which is synthesized in this way in
The reasons for the neurotoxic effects of
animal metabolism.
ammonia have not yet been explained. It
[5] In the final step, isourea is released
may disturb the metabolism of glutamate
from the guanidinium group of the arginine
and its precursor glutamine in the brain (see
by hydrolysis (not shown), and is immedi-
p. 356).
ately rearranged into urea. In addition, orni-
thine is regenerated and returns via the orni-
thine transporter into the mitochondria,
A. Urea cycle
where it becomes available for the cycle
Urea (H2N-CO-NH2) is the diamide of car-
once again.
bonic acid. In contrast to ammonia, it is neu-
The fumarate produced in step [4] is con-
tral and therefore relatively non-toxic. The
verted via malate to oxaloacetate [6, 7], from
reason for the lack of basicity is the molecule’s
which aspartate is formed again by transami-
mesomeric characteristics. The free electron
nation [9]. The glutamate required for reac-
pairs of the two nitrogen atoms are delocal-
tion [9] is derived from the glutamate dehy-
ized over the whole structure, and are there-
drogenase reaction [8], which fixes the sec-
fore no longer able to bind protons. As a small,
ond NH4+ in an organic bond. Reactions [6]
uncharged molecule, urea is able to cross bio-
and [7] also occur in the tricarboxylic acid
logical membranes easily. In addition, it is
cycle. However, in urea formation they take
easily transported in the blood and excreted
place in the cytoplasm, where the appropriate
in the urine.
isoenzymes are available.
Urea is produced only in the liver, in a cyclic
The rate of urea formation is mainly con-
sequence of reactions (the urea cycle) that
trolled by reaction [1]. N-acetyl glutamate, as
starts in the mitochondria and continues in
an allosteric effector, activates carbamoyl-
the cytoplasm. The two nitrogen atoms are
phosphate synthase. In turn, the concentration
derived from NH4+ (the second has previously
of acetyl glutamate depends on arginine and
been incorporated into aspartate; see below).
ATP levels, as well as other factors.
The keto group comes from hydrogen carbo-
nate (HCO3-), or CO2 that is in equilibrium
with HCO3-.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Protein Metabolism
183
A. Urea cycle
1
Carbamoyl phosphate synthase (NH3)
3
Argininosuccinate synthase 6.3.4.5
6.3.4.16
2
Ornithine carbamoyltransferase 2.1.3.3
4
Argininosuccinate lyase 4.3.2.1
P
N-Acetyl
5
Arginase 3.5.3.1
A
glutamate
A
2
2
P P
P
P
P
6
Fumarate hydratase 4.2.1.2
7
Malate dehydrogenase 1.1.1.37
Carbamoyl
phosphate
8
Glutamate dehydrogenase 1.4.1.2
NH
1
4
P
NH2
C
9
Aspartate transaminase [PLP]
COO
O
H3N C
H
COO
HCO3
CH
H3N
C
H
2
2
CH2
Ornithine
CH2
Citrulline
CH2
CH2
H N
H
CH2
P
Transporter
H
N
NH2
Urea
H
C
O
NH2
O
COO
C
H3N
C
H
COO
H2N
CH2
H3N
C
H
5
A
CH2
CH
2
P
P
P
H2O
Arginine
CH2
Arginino-
CH2
succinate
COO
CH2
3
N
NH2
H2O
H
C
C
H
N
NH2
H
C
P
P
H2N
H
C
HN
A
4
COO
P
OOC
CH
2
C
COO
Fumarate
H
COO
CH2
H2O
COO
NH4
H
C NH3
CH
2
6
COO
CH
2
Aspartate
COO
8
C O
COO
N
A
CH2
COO
CH2
COO
N
A
2-Oxo-
CH2
H
C
OH
N
A
glutarate
CH2
H
C NH3
COO
C
O
COO
Malate
7
COO
Glutamate
9
Oxaloacetate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
184
Metabolism
have to be supplied in food. For example,
Amino acid biosynthesis
animal metabolism is no longer capable of
carrying out de-novo synthesis of the aro-
A. Symbiotic nitrogen fixation
matic amino acids (tyrosine is only non-es-
Practically unlimited quantities of elementary
sential because it can be formed from phenyl-
nitrogen (N2) are present in the atmosphere.
alanine when there is an adequate supply
However, before it can enter the natural nitro-
available). The branched-chain amino acids
gen cycle, it has to be reduced to NH3 and
(valine, leucine, isoleucine, and threonine) as
incorporated into amino acids (“fixed”). Only
well as methionine and lysine, also belong to
a few species of bacteria and bluegreen algae
the essential amino acids. Histidine and argi-
are capable of fixing atmospheric nitrogen.
nine are essential in rats; whether the same
These exist freely in the soil, or in symbiosis
applies in humans is still a matter of debate. A
with plants. The symbiosis between bacteria
supply of these amino acids in food appears to
of the genus Rhizobium and legumes
be essential at least during growth.
(Fabales)—such as clover, beans, and peas—is
The nutritional value of proteins
(see
of particular economic importance. These
p. 360) is decisively dependent on their es-
plants are high in protein and are therefore
sential amino acid content. Vegetable pro-
nutritionally valuable.
teins—e. g., those from cereals—are low in ly-
In symbiosis with Fabales, bacteria live as
sine and methionine, while animal proteins
bacteroids in root nodules inside the plant
contain all the amino acids in balanced pro-
cells. The plant supplies the bacteroids with
portions. As mentioned earlier, however,
nutrients, but it also benefits from the fixed
there are also plants that provide high-value
nitrogen that the symbionts make available.
protein. These include the soy bean, one of the
The N2-fixing enzyme used by the bacteria
plants that is supplied with NH3 by symbiotic
is nitrogenase. It consists of two components:
N2 fixers (A).
an Fe protein that contains an [Fe4S4] cluster
Non-essential amino acids are those that
as a redox system (see p. 106), accepts elec-
arise by transamination from 2-oxoacids in
trons from ferredoxin, and donates them to
the intermediary metabolism. These belong
the second component, the Fe-Mo protein.
to the glutamate family (Glu, Gln, Pro, Arg,
This molybdenum-containing protein trans-
derived from 2-oxoglutarate), the aspartate
fers the electrons to N2 and thus, via various
family (only Asp and Asn in this group, de-
intermediate steps, produces ammonia (NH3).
rived from oxaloacetate), and alanine, which
Some of the reducing equivalents are trans-
can be formed by transamination from pyru-
ferred in a side-reaction to H+. In addition to
vate. The amino acids in the serine family (Ser,
NH3, hydrogen is therefore always produced
Gly, Cys) and histidine, which arise from in-
as well.
termediates of glycolysis, can also be synthe-
sized by the human body.
B. Amino acid biosynthesis: overview
The proteinogenic amino acids (see p. 60) can
be divided into five families in relation to
their biosynthesis. The members of each fam-
ily are derived from common precursors,
which are all produced in the tricarboxylic
acid cycle or in catabolic carbohydrate metab-
olism. An overview of the biosynthetic path-
ways is shown here; further details are given
on pp. 412 and 413.
Plants and microorganisms are able to syn-
thesize all of the amino acids from scratch, but
during the course of evolution, mammals
have lost the ability to synthesize approxi-
mately half of the 20 proteinogenic amino
acids. These essential amino acids therefore
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Protein Metabolism
185
A. Symbiotic nitrogen fixation
Bacteroids
N
A
4
Ferredoxin
N
A
4
A
Fe-Protein
2
P P P
[Fe4S4]
per
Host cell
A
e
2
P P
P
Air
N
2
Root
nodule
FeMo-Protein
N2
[Fe4S4], [FeMoCo]
8 H
Iron-molyb-
denum
cofactor
H2
2 NH3
Soil
Nitrogenase
Vascular bundle
1.18.6.1
Ammonia
B. Amino acid biosynthesis: overview
Serine family
Cysteine
Glucose 6-
Ribose 5-
phosphate
phosphate
Pentose phos-
phate pathway
3-Phospho-
glycerate
Erythrose
Glycine
Serine
Glyco-
4-phosphate
lysis
Phospho-
enolpyruvate
Pyruvate family
Histidine
Valine
Alanine
Leucine
Pyruvate
Serine
Lysine
Asparagine
NH3
Trypto-
phan
Oxalo-
Tricarboxylic
Phenyl-
Tyrosine
Threo-
Aspar-
acetate
acid cycle
alanine
nine
tate
Synthesized from
Aromatic
phenylalanine
2-Oxo-
family
glutarat
Iso-
leucine
Methionine
2 NH3
Ornithine
Aspartate family
Urea cycle
Glutamine
Glutamate
Essential amino acid
NH3
Transamination
Proline
Arginine
Reductive amination
Glutamate family
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
186
Metabolism
species. The same applies to birds and many
Nucleotide degradation
reptiles. Most other animals continue purine
The nucleotides are among the most complex
degradation to reach allantoic acid or urea
metabolites. Nucleotide biosynthesis is elab-
and glyoxylate.
orate and requires a high energy input (see
Pyrimidine (right). In the degradation of
p. 188). Understandably, therefore, bases and
pyrimidine nucleotides (2), the free bases ura-
nucleotides are not completely degraded, but
cil
(Ura) and thymine (Thy) are initially re-
instead mostly recycled. This is particularly
leased as important intermediates. Both are
true of the purine bases adenine and guanine.
further metabolized in similar ways. The pyri-
In the animal organism, some 90% of these
midine ring is first reduced and then hydro-
bases are converted back into nucleoside
lytically cleaved. In the next step, E-alanine
monophosphates by linkage with phosphori-
arises by cleavage of CO2 and NH3 as the
bosyl diphosphate (PRPP) (enzymes [1] and
degradation product of uracil. When there is
[2]). The proportion of pyrimidine bases that
further degradation, E-alanine is broken
are recycled is much smaller.
down to yield acetate, CO2, and NH3. Propio-
nate, CO2, and NH3 arise in a similar way from
b-aminoisobutyrate, the degradation product
A. Degradation of nucleotides
of thymine (see p. 419).
The principles underlying the degradation of
purines (1) and pyrimidines (2) differ. In the
B. Hyperuricemia
human organism, purines are degraded into
uric acid and excreted in this form. The purine
The fact that purine degradation in humans
ring remains intact in this process. In contrast,
already stops at the uric acid stage can lead to
the ring of the pyrimidine bases (uracil, thy-
problems, since—in contrast to allantoin—uric
mine, and cytosine) is broken down into small
acid is poorly soluble in water. When large
fragments, which can be returned to the me-
amounts of uric acid are formed or uric acid
tabolism or excreted (for further details, see
processing is disturbed, excessive concentra-
p. 419).
tions of uric acid can develop in the blood
Purine (left). The purine nucleotide guano-
(hyperuricemia). This can result in the accu-
sine monophosphate (GMP, 1) is degraded in
mulation of uric acid crystals in the body.
two steps—first to the guanosine and then to
Deposition of these crystals in the joints can
guanine (Gua). Guanine is converted by de-
cause very painful attacks of gout.
amination into another purine base, xanthine.
Most cases of hyperuricemia are due to
In the most important degradative path-
disturbed uric acid excretion via the kidneys
way for adenosine monophosphate (AMP), it
(1). A high-purine diet (e. g., meat) may also
is the nucleotide that deaminated, and inosine
have unfavorable effects (2). A rare hereditary
monophosphate (IMP) arises. In the same way
disease, Lesch-Nyhan syndrome, results from
as in GMP, the purine base hypoxanthine is
a defect in hypoxanthine phosphoribosyl-
released from IMP. A single enzyme, xanthine
transferase (A, enzyme [1]). The impaired re-
oxidase [3], then both converts hypoxanthine
cycling of the purine bases caused by this
into xanthine and xanthine into uric acid. An
leads to hyperuricemia and severe neurolog-
oxo group is introduced into the substrate in
ical disorders.
each of these reaction steps. The oxo group is
Hyperuricemia can be treated with
derived from molecular oxygen; another reac-
allopurinol, a competitive inhibitor of xan-
tion product is hydrogen peroxide
(H2O2),
thine oxidase. This substrate analogue differs
which is toxic and has to be removed by
from the substrate hypoxanthine only in the
peroxidases.
arrangement of the atoms in the 5-ring.
Almost all mammals carry out further deg-
radation of uric acid with the help of uricase,
with further opening of the ring to allantoin,
which is then excreted. However, the pri-
mates, including humans, are not capable of
synthesizing allantoin. Uric acid is therefore
the form of the purines excreted in these
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nucleotide Metabolism
187
A. Degradation of nucleotides
1.
Purine nucleotides
2.
Pyrimidine nucleotides
GMP
AMP
UMP
CMP
dTMP
PRPP
P P
G
A
U
C
T
P
P
P
P
P
H2O
H2O
2
P
NH3
1
Uracil
Thymine
Free
Ade
IMP
Ura
Thy
G
I
bases
P
Adenine
N
A
N
A
P
P
P
N
A
N
A
O
Rib
P
P
P
R
P
C
PRPP
Dihydro-
HN C
Dihydro-
1
H
I
uracil
thymine
Gua
Guanine
(R = H)
C
CH
(R = CH
2
3)
O
N
Phospho-
H2O
P
H
ribosyl
H2O
H2O
diphos-
phate
PRPP
COO
NH3
Rib P
H2N
H
C
R
N-Carbamoyl
N-Carbamoyl
O
O2
O
β-amino-
H2O2
β-alanine
C
CH
H
2
isobutyrate
2O
O
N
C
C
(R = H)
N
N
(R = CH
3)
HN
C
HN
C
H
CH
3
CH
H
H2O
C
C
HC
C
2O
N
N
O
N
N
H
H
H
COO
NH3,
NH3,
CO2
CO2
Xanthine (Xan)
Hypoxanthine (Hyp)
H
C
R
O2
CH2
β-Amino-
H2O
β-Alanine
isobutyrate
3
(R = H)
NH3
(R = CH
3)
H2O2
Excreted
by primates,
B. Hyperuricemia (gout)
O
birds, and
H
reptiles
Causes:
C
2a
N
HN
C
Disturbed uric
C O
1
acid excretion
C
C
N
O
N
Uric acid
GMP
AMP
H
2
Elevated uric acid
H
formation
Excreted
2b
2b
a) Unbalanced
by most
a) nutrition
H
mammals
O
N
b) Impaired
H2N
C
Gua
Ade
IMP
C
O
b) recycling of
C
C
b) purine bases
N
2b
O
N
H
H
Allantoin
H
Hypo-
Recycling
Xanthine
xanthine
reactions
2 Urea +
Allantoic acid
glyoxylate
O
Allopurinol
H
C
C
Hypoxanthine phosphoribosyl-
HN
C
1
transferase 2.4.2.8
Uric
N
1
HC
C
Adenine phosphoribosyl-
acid
N
2
N
transferase 2.4.2.7
H
3
Xanthine oxidase [Fe, Mo, FAD] 1.1.3.22
Allopurinol
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
188
Metabolism
(1b) and then converted into dihydroorotate
Purine and pyrimidine biosynthesis
by closure of the ring (1c). In mammals, steps
The bases occurring in nucleic acids are de-
1a to 1c take place in the cytoplasm, and are
rivatives of the aromatic heterocyclic com-
catalyzed by a single multifunctional enzyme.
pounds purine and pyrimidine (see p. 80).
In the next step (1d), dihydroorotate is oxi-
The biosynthesis of these molecules is com-
dized to orotate by an FMN-dependent dehy-
plex, but is vital for almost all cells. The syn-
drogenase. Orotate is then linked with phos-
thesis of the nucleobases is illustrated here
phoribosyl diphosphate (PRPP) to form the
schematically. Complete reaction schemes
nucleotide orotidine
5 -monophosphate
are given on pp. 417 and 418.
(OMP). Finally, decarboxylation yields uridine
5 -monophosphate (UMP).
Purine biosynthesis starts with PRPP (the
A. Components of nucleobases
names of the individual intermediates are
The pyrimidine ring is made up of three com-
given on p. 417). Formation of the ring starts
ponents: the nitrogen atom N-1 and carbons
with transfer of an amino group, from which
C-4 to C-6 are derived from aspartate, carbon
the later N-9 is derived (2a). Glycine and a
C-2 comes from HCO3-, and the second nitro-
formyl group from N10-formyl-THF then sup-
gen (N-3) is taken from the amide group of
ply the remaining atoms of the five-mem-
glutamine.
bered ring (2b, 2c). Before the five-membered
The synthesis of the purine ring is more
ring is closed (in step 2f), atoms N-3 and C-6
complex. The only major component is gly-
of the later six-membered ring are attached
cine, which donates C-4 and C-5, as well as N-
(2d, 2e). Synthesis of the ring then continues
7. All of the other atoms in the ring are in-
with N-1 and C-2 (2g, 2i). In the final step (2j),
corporated individually. C-6
comes from
the six-membered ring is closed, and inosine
HCO3-. Amide groups from glutamine provide
5 -monophosphate arises. However, the IMP
the atoms N-3 and N-9. The amino group
formed does not accumulate, but is rapidly
donor for the inclusion of N-1 is aspartate,
converted into AMP and GMP. These reactions
which is converted into fumarate in the proc-
and the synthesis of the other nucleotides are
ess, in the same way as in the urea cycle (see
discussed on p. 190.
p. 182). Finally, the carbon atoms C-2 and C-8
are derived from formyl groups in N10-
Further information
formyl-tetrahydrofolate (see p. 108).
The regulation of bacterial aspartate
carbamoyltransferase by ATP and CTP has
B. Pyrimidine and purine synthesis
been particularly well studied, and is dis-
The major intermediates in the biosynthesis
cussed on p. 116. In animals, in contrast to
of nucleic acid components are the
prokaryotes, it is not ACTase but carbamoyl-
mononucleotides uridine monophosphate
phosphate synthase that is the key enzyme in
(UMP) in the pyrimidine series and inosine
pyrimidine synthesis. It is activated by ATP
monophosphate (IMP, base: hypoxanthine) in
and PRPP and inhibited by UTP.
the purines. The synthetic pathways for pyri-
The biosynthesis of the purines is also
midines and purines are fundamentally dif-
regulated by feedback inhibition. ADP and
ferent. For the pyrimidines, the pyrimidine
GDP inhibit the formation of PRRPP from ri-
ring is first constructed and then linked to
bose-5 -phosphate. Similarly, step 2a is in-
ribose 5 -phosphate to form a nucleotide. By
hibited by AMP and GMP.
contrast, synthesis of the purines starts di-
rectly from ribose 5 -phosphate. The ring is
then built up step by step on this carrier mol-
ecule.
The precursors for the synthesis of the
pyrimidine ring are carbamoyl phosphate,
which arises from glutamate and HCO3- (1a)
and the amino acid aspartate. These two com-
ponents are linked to N-carbamoyl aspartate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nucleotide Metabolism
189
A. Components of nucleobases
Glu
4
Fumarate
Glycine
HCO3
NH3
C
H2C
3
5
2g
2f
N
C
2h
COO
[NH2]
[NH2]
2b
C
C
6
2
6
7
1a
Gln
Asp
N
C
5
THF
N
1
N
C
1
THF
[CHO]
C
8
2c
1
C
C
[CHO]
OHC
2
N
HCO3
N
4
2i
N10
OOC
3
9
THF
CHO
CH
2
N10
[NH2]
[NH2]
THF
C
H3N H COO
Aspartate
Gln
Glu
Gln
Glu
2d
2a
Pyrimidine
Purine
B. Pyrimidine and purine synthesis
[NH
Carbamoyl
Phosphoribosyl
P
R
P P
2]
C
N
phosphate
diphosphate
N
C
C
C
N
C
O P
N
C
1a
C
C
N
C
N
C
C
C
C
C
C
N
C
N
C
C
C
C
C
1b
1c
1d
1e
1f
HCO3
N
C
N
C
C
P
R
P
R
C
N-Carbamoyl
Dihydro-
Orotate 5'-mono- Uridine 5'-mono-
N
C
aspartate
orotate
phosphate
phosphate (UMP)
Aspartate
5'
O
UMP
1. Pyrimidines
P
O
CH2
O
C
1'
HN
CH
Phosphoribosyl
R
H H
P
=
C
CH
diphosphate
H
H
O
N
P
R
P P
N
C
OH OH
P
O
CH2
2a
C
O
5'-Phosphoribosyl residue
Glycine
H H
H
H
N
N
N
N
C
C
C
C
C
OH OH
P
R
2b
C
2c
C
2d
C
N
N
N
N
O
P
R
P
R
P
R
IMP
C
N
HN
C
C
C
C
C
N
N
N
N
CH
C
N
C
N
C
N
C
HC
C
C
C
C
C
N
N
C
C
C
C
C
C
2e,
N
2g,
N
2i
N
2j
N
N
N
N
N
P
O
CH2
2f
2h
O
P
R
P
R
P
R
P
R
H H
Inosine 5'-mono-
H
H
phosphate (IMP)
2. Purines
OH OH
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
190
Metabolism
side diphosphates. This reduction is a com-
Nucleotide biosynthesis
plex process in which several proteins are
De novo synthesis of purines and pyrimidines
involved. The reducing equivalents needed
yields the monophosphates IMP and UMP,
come from NADPH+H+. However, they are
respectively (see p. 188). All other nucleotides
not transferred directly from the coenzyme
and deoxynucleotides are synthesized from
to the substrate, but first pass through a redox
these two precursors. An overview of the
series that has several steps (1).
pathways involved is presented here; further
In the first step, thioredoxin reductase re-
details are given on p. 417. Nucleotide syn-
duces a small redox protein, thioredoxin, via
thesis by recycling of bases (the salvage path-
enzyme-bound FAD. This involves cleavage of
way) is discussed on p. 186.
a disulfide bond in thioredoxin. The resulting
SH groups in turn reduce a catalytically active
disulfide bond in nucleoside diphosphate
A. Nucleotide synthesis: overview
reductase
(“ribonucleotide reductase”). The
The synthesis of purine nucleotides (1) starts
free SH groups formed in this way are the
from IMP. The base it contains, hypoxanthine,
actual electron donors for the reduction of
is converted in two steps each into adenine or
ribonucleotide diphosphates.
guanine. The nucleoside monophosphates
In eukaryotes, ribonucleotide reductase is a
AMP and GMP that are formed are then phos-
tetramer consisting of two R1 and two R2
phorylated by nucleoside phosphate kinases to
subunits. In addition to the disulfide bond
yield the diphosphates ADP and GDP, and
mentioned, a tyrosine radical in the enzyme
these are finally phosphorylated into the
also participates in the reaction (2). It initially
triphosphates ATP and GTP. The nucleoside
produces a substrate radical (3). This cleaves a
triphosphates serve as components for RNA,
water molecule and thereby becomes radical
or function as coenzymes (see p. 106). Con-
cation. Finally, the deoxyribose residue is pro-
version of the ribonucleotides into deoxyribo-
duced by reduction, and the tyrosine radical is
nucleotides occurs at the level of the diphos-
regenerated.
phates and is catalyzed by nucleoside diphos-
The regulation of ribonucleotide reductase
phate reductase (B).
is complex. The substrate-specificity and ac-
The biosynthetic pathways for the pyrimi-
tivity of the enzyme are controlled by two
dine nucleotides (2) are more complicated.
allosteric binding sites (a and b) in the R1
The first product, UMP, is phosphorylated first
subunits. ATP and dATP increase or reduce
to the diphosphate and then to the
the activity of the reductase by binding at
triphosphate, UTP. CTP synthase then converts
site a. Other nucleotides interact with site b,
UTP into CTP. Since pyrimidine nucleotides
and thereby alter the enzyme’s specificity.
are also reduced to deoxyribonucleotides at
the diphosphate level, CTP first has to be hy-
drolyzed by a phosphatase to yield CDP before
dCDP and dCTP can be produced.
The DNA component deoxythymidine tri-
phosphate (dTTP) is synthesized from UDP in
several steps. The base thymine, which only
occurs in DNA (see p. 80), is formed by meth-
ylation of dUMP at the nucleoside monophos-
phate level. Thymidylate synthase and its
helper enzyme dihydrofolate reductase are
important target enzymes for cytostatic drugs
(see p. 402).
B. Ribonucleotide reduction
2 -Deoxyribose, a component of DNA, is not
synthesized as a free sugar, but arises at the
diphosphate level by reduction of ribonucleo-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nucleotide Metabolism
191
A. Nucleotide synthesis: overview
1. Purine
Precursors
2. Pyrimidine
Precursors
1. nucleotides
2. nucleotides
De novo synthesis
De novo synthesis
3
AMP
IMP
GMP
UMP
dUMP
dTMP
4
4
4
4
1
1
1
1
dADP
ADP
GDP
dGDP
dCDP
CDP
UDP
dUDP
dTDP
5
5
5
5
5
5
5
2
dATP
ATP
GTP
dGTP
dCTP
CTP
UTP
dTTP
DNA
RNA
RNA
DNA
DNA
RNA
RNA
DNA
Ribonucleoside
2
CTP synthase 6.3.4.2
4
Nucleoside phosphate kinase 2.7.4.4
1
diphosphate
reductase 1.17.4.1
3
Thymidylate synthase 2.1.1.45
5
Nucleoside diphosphate kinase 2.7.4.6
B. Ribonucleotide reduction
NADPH
N
A
N
A
NADP
NTP
NTP
+H
P
P
a
a
R1
R1
6
ATP
ATP
dATP
dATP
b
b
S
SH
S S
S
S
Substrate
Thio-
Thio-
S
SH
(NDP)
redoxin
redoxin
O
O
(oxidized)
(reduced)
R2
R2
Tyrosine
radical
2. Ribonucleotide reductase
1
R
CH2
R
CH2
HS SH
S S
Base
Base
O
O
H
H
1
H
H
H
H
H
OH OH
OH OH
H2O
NDP
Radical
P P
O
P
P
O
Base
O Base
O
2
H H
H H
H
O
OH
NDP
HO OH
HO H
dNDP
H2O
Nucleoside
Deoxyribonucleoside
diphosphate
diphosphate
R
CH2
R
CH2
Base
Base
O
O
H
H
H
H
H
H
H
1
Ribonucleoside diphosphate reductase 1.17.4.1
3
OH H
OH
6
Thioredoxin reductase [FAD] 1.6.4.5
2e , H
dNDP
1. Overview
3. Reaction mechanism
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
192
Metabolism
chains have to be converted into less polar
Heme biosynthesis
groups.
Heme, an iron-containing tetrapyrrole pig-
[5] Initially, the four acetate residues (R1)
ment, is a component of O2-binding proteins
are decarboxylated into methyl groups. The
(see p. 106) and a coenzyme of various oxi-
resulting coproporphyrinogen III returns to
doreductases (see p. 32). Around 85% of heme
the mitochondria again. The subsequent steps
biosynthesis occurs in the bone marrow, and a
are catalyzed by enzymes located either on or
much smaller percentage is formed in the
inside the inner mitochondrial membrane.
liver. Both mitochondria and cytoplasm are
[6] An oxidase first converts two of the
involved in heme synthesis.
propionate groups (R2) into vinyl residues.
The formation of protoporphyrinogen IX com-
pletes the modification of the side chains.
A. Biosynthesis of heme
[7] In the next step, another oxidation pro-
Synthesis of the tetrapyrrole ring starts in the
duces the conjugated π-electron system of
mitochondria.
protoporphyrin IX.
[1] Succinyl CoA (upper left), an intermedi-
[8] Finally, a divalent iron is incorporated
ate in the tricarboxylic acid cycle, undergoes
into the ring. This step also requires a specific
condensation with glycine and subsequent
enzyme, ferrochelatase. The heme b or Fe-pro-
decarboxylation to yield
5-aminolevulinate
toporphyrin IX formed in this way is found in
(ALA). The ALA synthase responsible for this
hemoglobin and myoglobin, for example (see
step is the key enzyme of the whole pathway.
p. 280), where it is noncovalently bound, and
Synthesis of ALA synthase is repressed and
also in various oxidoreductases (see p. 106).
existing enzyme is inhibited by heme, the
end product of the pathway. This is a typical
Further information
example of end-product or feedback inhibi-
tion.
There are a large number of hereditary or
[2] 5-Aminolevulinate now leaves the mi-
acquired disturbances of porphyrin synthesis,
tochondria. In the cytoplasm, two molecules
known as porphyrias, some of which can
condense to form porphobilinogen, a com-
cause severe clinical pictures. Several of these
pound that already contains the pyrrole ring.
diseases lead to the excretion of heme pre-
Porphobilinogen synthase is inhibited by lead
cursors in feces or urine, giving them a dark
ions. This is why acute lead poisoning is asso-
red color. Accumulation of porphyrins in the
ciated with increased concentrations of ALA
skin can also occur, and exposure to light then
in the blood and urine.
causes disfiguring, poorly healing blisters.
[3] The tetrapyrrole structure characteristic
Neurological disturbances are also common
of the porphyrins is produced in the next
in the porphyrias.
steps of the synthetic pathway. Hydroxyme-
It is possible that the medieval legends
thylbilane synthase catalyzes the linkage of
about human vampires (“Dracula”) originated
four porphobilinogen molecules and cleavage
in the behavior of porphyria sufferers (avoid-
of an NH2 group to yield uroporphyrinogen III.
ance of light, behavioral disturbances, and
[4] Formation of this intermediate step re-
drinking of blood in order to obtain heme—
quires a second enzyme, uroporphyrinogen III
which markedly improves some forms of por-
synthase. If this enzyme is lacking, the
phyria).
“wrong” isomer, uroporphyrinogen I, is
formed.
The tetrapyrrole structure of uroporphyri-
nogen III is still very different from that of
heme. For example, the central iron atom is
missing, and the ring contains only eight of
the 11 double bonds. In addition, the ring
system only carries charged R side chains
(four acetate and four propionate residues).
As heme groups have to act in the apolar
interior of proteins, most of the polar side
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Porphyrin Metabolism
193
A. Heme biosynthesis
Tricarboxylic
2
acid cycle
R2
R1
Succinyl
COO
R1
R2
CoA
A
B
CH2
N
N
Heme
Fe2+
CH2
N
N
2+
D
C
S
R1
R1
C O
A
2
R3
R
3
Vinyl
H2C COO
Glycine
residue
NH3
R
Fe2
8
COO
1
CH2
CH2
S
2
A
Protoporphyrin IX
CH3
CH
CH2
2
CO2
6[H]
R1
R2
R3
7
1COO
Protoporphyrinogen IX
1
COO
CH2
5-Amino-
2
2 CO2
Mitochondrion
levulinate
CH
2
CH2
6
(ALA)
CH2
O2
O
C
5
H2C
C
5
CH2
O
Cytoplasm
NH3
NH3
Coproporphyrinogen III
COO
COO
CH2
CH2
CH3
CH2
CH2
4 CO2
5
2
R1
R2
R3
2 H2O
COO
R2
R1
OOC CH2
COO COO
R1
R2
A
B
CH
CH2
COO
CH2
CH2
2
NH HN
Pyrrole
3
4
CH2
CH2
CH2
ring
C
C
NH HN
H2C
C
CH
D
C
R1
R1
N
R1
R2
R3
4 NH4
H2O
NH3
H
R3
R3
Porphobilinogen
Uroporphyrinogen III
5-Aminolevulinate synthase
Porphobilinogen synthase
Hydroxymethylbilane synthase
1
2
3
[PLP] 2.3.1.37
4.2.1.24
4.3.1.8
Uroporphyrinogen-III synthase
4
4.2.1.75
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
194
Metabolism
Some of the bilirubin conjugates are bro-
Heme degradation
ken down further in the intestine by bacterial
b-glucuronidases. The bilirubin released is
A. Degradation of heme groups
then reduced further via intermediate steps
Heme is mainly found in the human organism
into colorless stercobilinogen, some of which
as a prosthetic group in erythrocyte hemoglo-
is oxidized again into orange to yellow-col-
bin. Around 100-200 million aged erythro-
ored stercobilin. The end products of bile pig-
cytes per hour are broken down in the human
ment metabolism in the intestine are mostly
organism. The degradation process starts in
excreted in feces, but a small proportion is
reticuloendothelial cells in the spleen, liver,
resorbed
(enterohepatic circulation; see
and bone marrow.
p. 314). When high levels of heme degrada-
[1] After the protein part (globin) has been
tion are taking place, stercobilinogen appears
removed, the tetrapyrrole ring of heme is
as urobilinogen in the urine, where oxidative
oxidatively cleaved between rings A and B
processes darken it to form urobilin.
by heme oxygenase. This reaction requires
In addition to hemoglobin, other heme pro-
molecular oxygen and NADPH+H+, and pro-
teins (myoglobin, cytochromes, catalases, and
duces green biliverdin, as well as CO (carbon
peroxidases; see p. 32) also supply heme
monoxide) and Fe2+, which remains available
groups that are degraded via the same path-
for further use (see p. 286).
way. However, these contribute only about
[2] In another redox reaction, biliverdin is
10-15% to a total of ca. 250 mg of bile pig-
reduced by biliverdin reductase to the orange-
ment formed per day.
colored bilirubin. The color change from pur-
ple to green to yellow can be easily observed
Further information
in vivo in a bruise or hematoma.
The color of heme and the other porphyrin
Hyperbilirubinemias. An elevated bilirubin
systems (see p. 106) results from their numer-
level (> 10 mg L-1) is known as hyperbiliru-
ous conjugated double bonds. Heme contains
binemia. When this is present, bilirubin dif-
a cyclic conjugation (highlighted in pink) that
fuses from the blood into peripheral tissue
is removed by reaction
[1]. Reaction
[2]
and gives it a yellow color (jaundice). The
breaks the π system down into two smaller
easiest way of observing this is in the white
separate systems (highlighted in yellow).
conjunctiva of the eyes.
For further degradation, bilirubin is trans-
Jaundice can have various causes. If in-
ported to the liver via the blood. As bilirubin is
creased erythrocyte degradation (hemolysis)
poorly soluble, it is bound to albumin for
produces more bilirubin, it causes hemolytic
transport. Some drugs that also bind to albu-
jaundice. If bilirubin conjugation in the liver is
min can lead to an increase in free bilirubin.
impaired—e. g., due to hepatitis or liver cir-
[3] The hepatocytes take up bilirubin from
rhosis—it leads to hepatocellular jaundice,
the blood and conjugate it in the endoplasmic
which is associated with an increase in un-
reticulum with the help of UDP-glucuronic
conjugated (“indirect”) bilirubin in the blood.
acid into the more easily soluble bilirubin
By contrast, if there is a disturbance of bile
monoglucuronides and diglucuronides. To do
drainage (obstructive jaundice, due to gall-
this, UDP-glucuronosyltransferase forms ester-
stones or pancreatic tumors), then conjugated
type bonds between the OH group at C-1 of
(“direct”) bilirubin in the blood increases. Neo-
glucuronic acid and the carboxyl groups in
natal jaundice (physiologic jaundice) usually
bilirubin (see p. 316). The glucuronides are
resolves after a few days by itself. In severe
then excreted by active transport into the
cases, however, unconjugated bilirubin can
bile, where they form what are known as
cross the blood-brain barrier and lead to
the bile pigments.
brain damage (kernicterus).
Glucuronide synthesis is the rate-deter-
mining step in hepatic bilirubin metabolism.
Drugs such as phenobarbital, for example, can
induce both conjugate formation and the
transport process.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Porphyrin Metabolism
195
A. Degradation of heme groups
Heme
Cleavage by
1
N
A
N
A
R2
CH3
3
3
CO
P
P
H3C
R3
A
B
N
N
2
1
Fe2
N
N
3 O2
3 H2O
Fe2
D
C
R
1
CH3
Heme
Reused
for heme
synthesis
COO
COO
Biliverdin
CH2
OOC
CH2
H2C COO
CH2
CH3
CH
CH3
CH2
CH2
CH3
CH3
CH
N
A
N
A
B
C
D
A
P
P
O N
C
N
C
N
C
N O
H
H
H
H2
H
H
H
Bilirubin
2
RES
Bilirubin
Bilirubin
3
Bilirubin
Urine
diglucuronide
U
2
P P
GlcUA
2 UDP
Slowest
Blood
step in
Albumin
hepatic
metabo-
lism
Gallbladder
Entero-
Albumin
hepatic
Bile
circulation
Bilirubin
pigments
Intestine
Binding
site for
bilirubin
Bilirubin
Bacterial
and drugs
diglucuronide
metabolism
2 GlcUA
Heme oxygenase
1
Stercobilinogen
Bilirubin
(decyclizing) 1.14.99.3
Urobilinogen
Biliverdin reductase
2
1.3.1.24
Stercobilin
Glucuronosyl
Urobilin
3
transferase 2.4.1.17
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
196
Organelles
B. Structure of an animal cell
Structure of cells
In the human body alone, there are at least
A. Comparison of prokaryotes and
200 different cell types. The illustration out-
eukaryotes
lines the basic structures of an animal cell in
an extremely simplified way. The details
Present-day living organisms can be divided
given regarding the proportion of the com-
into two large groups—the prokaryotes and
partments relative to cell volume (highlighted
eukaryotes. The prokaryotes are represented
in yellow) and their numbers per cell fre-
by bacteria (eubacteria and archaebacteria).
quency (blue) refer to mammalian hepato-
These are almost all small unicellular organ-
cytes (liver cells). The figures can vary widely
isms only a few microns (10-6 m) in size. The
from cell type to cell type.
eukaryotes include fungi, plants, and animals
The eukaryotic cell is subdivided by mem-
and comprise both unicellular and multicel-
branes. On the outside, it is enclosed by a
lular organisms. Multicellular eukaryotes are
plasma membrane. Inside the cell, there is a
made up of a wide variety of cell types that are
large space containing numerous components
specialized for different tasks. Eukaryotic cells
in solution—the cytoplasm. Additional mem-
are much larger than prokaryotic ones (vol-
branes divide the internal space into
ume ratio approximately 2000 : 1). The most
compartments
(confined reaction spaces).
important distinguishing feature of these cells
Welldefined compartments of this type are
in comparison with the prokaryotes is the fact
known as organelles.
that they have a nucleus
(karyon in
The largest organelle is the nucleus (see
Greek—hence the term).
p. 208). It is easily recognized using the light
In comparison with the prokaryotes, eu-
microscope. The endoplasmic reticulum (ER),
karyotic cells have greater specialization and
a closed network of shallow sacs and tubules
complexity in their structure and functioning.
(see pp. 226ff.), is linked with the outer mem-
Eukaryotic cells are structured into compart-
brane of the nucleus. Another membrane-
ments (see below). The metabolism and syn-
bound organelle is the Golgi apparatus (see
thesis of macromolecules are distributed
p. 228), which resembles a bundle of layered
through these reaction spaces and are sepa-
slices. The endosomes and exosomes are bub-
rately regulated. In prokaryotes, these func-
ble-shaped compartments ( vesicles) that are
tions are organized in a simpler fashion and
involved in the exchange of substances be-
are spatially closely related.
tween the cell and its surroundings. Probably
Although the storage and transfer of ge-
the most important organelles in the cell’s
netic information function according to the
metabolism are the mitochondria, which are
same principle in the prokaryotes and euka-
around the same size as bacteria
(see
ryotes, there are also differences. Eukaryotic
pp. 210ff.). The lysosomes and peroxisomes
DNA consists of very long, linear molecules
are small, globular organelles that carry out
with a total of 107 to more than 1010 base pairs
specific tasks. The whole cell is traversed by a
(bp), only a small fraction of which are used
framework of proteins known as the cytoske-
for genetic information. In eukaryotes, the
leton (see pp. 204ff.).
genes (20 000-50 000 per genome) are usu-
In addition to these organelles, plant cells
ally interrupted by non-coding regions (in-
(see p. 43) also have plastids—eg., chloro-
trons). Eukaryotic DNA is located in the nu-
plasts, in which photosynthesis takes place
cleus, where together with histones and other
(see p. 128). In their interior, there is a large,
proteins it forms the chromatin (see p. 238).
fluid-filled vacuole. Like bacteria and fungi,
In prokaryotes, by contrast, DNA is ring-
plant cells have a rigid cell wall consisting of
shaped, much shorter (up to 5
106 bp), and
polysaccharides and proteins.
located in the cytoplasm. Almost all of it is
used for information storage, and it does not
contain any introns.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Basics
197
A. Comparison of prokaryotes and eukaryotes
Prokaryotes
Eukaryotes
Organisms
Eubacteria
Fungi
1-10 µm
Archaebacteria
Plants
Animals
Form
Single-celled
Single or multi-cellular
Organelles, cytoskeleton, cell division apparatus
10-100 µm
Missing
Present, complicated, specialized
Small, circular, no
DNA
introns, plasmids
Large, in nucleus, many introns
RNA: Synthesis and maturation
Simple, in cytoplasm
Complicated, in nucleus
Protein: Synthesis and maturation
Simple, coupled with
Complicated, in the cytoplasm
RNA synthesis
and the rough endoplasmic reticulum
Metabolism
Anaerobic or aerobic
very flexible
Mostly aerobic, compartmented
Endocytosis
and Exocytosis
no
yes
B. Structure of an animal cell
Golgi complex
Plasma
membrane
6%
?
Nucleus
Lysosome
6%
1
1%
300
Rough
endoplasmic
Endosome
reticulum
1%
200
9%
1
Free
Mitochondrion
ribosomes
22%
~2000
Peroxisome
Cytoplasm
1%
400
54%
1
Number per cell
Proportion of
10-30 µm
cell volume
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
198
Organelles
Particles that are smaller and less dense
Cell fractionation
than the nuclei can be obtained by step-by-
step acceleration of the gravity on the super-
A. Isolation of cell organelles
natant left over from the first centrifugation.
To investigate the individual compartments of
However, this requires very powerful centri-
the cell (see p. 196), various procedures have
fuges (high-speed centrifuges and ultracentri-
been developed to enrich and isolate cell or-
fuges). The sequence in which the fractions
ganelles. These are mainly based on the size
are obtained is: mitochondria, membrane
and density of the various organelles.
vesicles, and ribosomes. Finally, the superna-
The isolation of cell components starts
tant from the last centrifugation contains the
with disruption of the tissue being examined
cytosol with the cell’s soluble components, in
and subsequent homogenization of it (break-
addition to the buffer.
ing down the cells) in a suitable buffer (see
The isolation steps are carried out at low
below). Homogenization using the “Potter”
temperatures on principle (usually 0-5 °C), to
(the Potter-Elvehjem homogenizer, a rotating
slow down degradation reactions—e. g., due
Teflon pestle in a glass cylinder) is particularly
to released enzymes and other influencing
suitable for animal tissue. This method is very
factors. The addition of thiols and chelating
gentle and is therefore used to isolate fragile
agents protects functional SH groups from
structures and molecules. Other cell disrup-
oxidation. Isolated cell organelles quickly
tion procedures include enzymatic lysis with
lose their biological activity despite these pre-
the help of enzymes that break down the cell
cautions. Nevertheless, it is possible by work-
wall, mechanical disruption by grinding fro-
ing carefully to isolate mitochondria that will
zen tissue, cutting or smashing with rotating
still take up substrates for a few hours in the
knives, large pressure changes, osmotic
test tube and produce ATP via oxidative phos-
shock, and repeated freezing and thawing.
phorylation.
To isolate intact organelles, it is important
for the homogenization solution to be iso-
B. Marker molecules
tonic—i. e., the osmotic value of the buffer
has to be the same as that of the interior of
During cell fractionation, it is very important
the cell. If hypotonic solutions were used, the
to analyze the purity of the fractions obtained.
organelles would take up water and burst,
Whether or not the intended organelle is
while in hypertonic solutions they would
present in a particular fraction, and whether
shrink.
or not the fraction contains other compo-
Homogenization is followed by coarse fil-
nents, can be determined by analyzing
tration through gauze to remove intact cells
characteristic marker molecules. These are
and connective-tissue fragments. The actual
molecules that occur exclusively or predom-
fractionation of cellular components is then
inantly in one type of organelle. For example,
carried out by centrifugation steps, in which
the activity of organelle-specific enzymes
the gravitational force (given as multiples of
(marker enzymes) is often assessed. The dis-
the earth’s gravity, g
= 9.81 m s-2) is gradu-
tribution of marker enzymes in the cell re-
ally increased (differential centrifugation; see
flects the compartmentation of the processes
p. 200). Due to the different shapes and den-
they catalyze. These reactions are discussed in
sities of the organelles, this leads to succes-
greater detail here under the specific organ-
sive sedimentation of each type out of the
elles.
suspension.
Nuclei already sediment at low accelera-
tions that can be achieved with bench-top
centrifuges. Decanting the residue
(the
“supernatant”) and carefully suspending the
sediment (or “pellet”) in an isotonic medium
yields a fraction that is enriched with nuclei.
However, this fraction may still contain other
cellular components as contaminants—e. g.,
fragments of the cytoskeleton.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Basics
199
A. Isolation of cell organelles
Homo-
Filter
genize
Gauze
Slice
Whole
Tissue
cells,
connective
tissue
Potter
homo-
Buffer
genizer
Centrifuge
g = 300 000
g = 100 000
g = 15 000
g = 600
120'
60'
15'
10'
Cytosol
Super-
natant
Centrifuge
tube
Pellet
Ribosomes
Plasma membrane
Mitochondria
Nucleus
Viruses
ER fragments
Lysosomes
Cytoskeleton
Macromolecules
Small vesicles
Peroxisomes
Microsomal
(Plants:
fraction
chloroplasts)
B. Marker molecules
Golgi complex
Nucleus
Ribosomes
Plasma membrane
α-Mannosidase II
DNA
rRNA
Na+/K+ ATPase
3.2.1.24
3.6.1.37
Phosphodiesterase I
3.1.4.1
Endoplasmic
reticulum
Lysosome
β-N-Acetylhexos-
Glucose 6-phospha-
tase 3.1.3.9
aminidase 3.2.1.52
RNA
β-Galactosidase
3.2.1.23
Mitochondrion
Succinate dehydro-
Endosome
genase 1.3.5.1
Uptake of
Cytochrome-c
peroxidase
oxidase
1.11.1.7
1.9.3.1
Cytosol
Peroxisome
L-Lactate
Catalase
dehydrogenase
1.11.1.6
1.1.1.27
Cell fraction
Marker enzyme
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
200
Organelles
B. Density gradient centrifugation
Centrifugation
Density gradient centrifugation is used to
A. Principles of centrifugation
separate macromolecules that differ only
slightly in size or density. Two techniques
In a solution, particles whose density is higher
are commonly used.
than that of the solvent sink (sediment), and
In zonal centrifugation, the sample being
particles that are lighter than it float to the
separated (e. g., a cell extract or cells) is placed
top. The greater the difference in density, the
on top of the centrifugation solution as a thin
faster they move. If there is no difference in
layer. During centrifugation, the particles
density (isopyknic conditions), the particles
move through the solution due to their
hover. To take advantage of even tiny differ-
greater density. The rate of movement basi-
ences in density to separate various particles
cally depends on their molecular mass (see A,
in a solution, gravity can be replaced with the
formulae). Centrifugation stops before the
much more powerful “centrifugal force” pro-
particles reach the bottom of the tube. Dril-
vided by a centrifuge.
ling a hole into the centrifugation tube and
Equipment. The acceleration achieved by
allowing the contents to drip out makes it
centrifugation is expressed as a multiple
possible to collect the different particles in
of the earth’s gravitational force
(g
=
separate fractions. During centrifugation, the
9.81 m s-2). Bench-top centrifuges can reach
solution tube is stabilized in the tube by a
acceleration values of up to 15 000 g, while
density gradient. This consists of solutions of
highspeed refrigerated centrifuges can reach
carbohydrates or colloidal silica gel, the con-
50000 g and ultracentrifuges, which operate
centration of which increases from the sur-
with refrigeration and in a vacuum, can reach
face of the tube to the bottom. Density gra-
500000 g. Two types of rotor are available in
dients prevent the formation of convection
high-powered centrifuges: fixed angle rotors
currents, which would impair the separation
and swingout rotors that have movable bucket
of the particles.
containers. The tubes or buckets used for cen-
Isopyknic centrifugation, which takes
trifugation are made of plastic and have to be
much longer, starts with a CsCl solution in
very precisely adjusted to avoid any imbalan-
which the sample material (e. g., DNA, RNA,
ces that could lead to accidents.
or viruses) is homogeneously distributed. A
Theory. The velocity (v) of particle sedi-
density gradient only forms during centrifu-
mentation during centrifugation depends on
gation, as a result of sedimentation and dif-
the angular velocity ω of the rotor, its effective
fusion processes. Each particle moves to the
radius (reff, the distance from the axis of rota-
region corresponding to its own buoyant den-
tion), and the particle’s sedimentation prop-
sity. Centrifugation stops once equilibrium
erties. These properties are expressed as the
has been reached. The samples are obtained
sedimentation coef•cient S
(1 Svedberg,
by fractionation, and their concentration is
= 10-13 s). The sedimentation coef cient de-
measured using the appropriate methods.
pends on the mass M of the particle, its shape
(expressed as the coef cient of friction, f), and
its density
(expressed as the reciprocal
density v_, “partial specific volume”).
At the top right, the diagram shows the
densities and sedimentation coef cients for
biomolecules, cell organelles, and viruses.
Proteins and protein-rich structures have
densities of around 1.3 g cm-3, while nucleic
acids show densities of up to 2 g cm-3. Equi-
librium sedimentation of nucleic acids there-
fore requires high-density media—e. g., con-
centrated solutions of cesium chloride (CsCl).
To allow comparison of S values measured in
different media, they are usually corrected to
values for water at 20 °C (“S20W” ).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Basics
201
A. Principles of centrifugation
Centrifuge
bucket
Mitochondria
1.2
Microsomes
Soluble
Nuclei
1.4
proteins
Viruses
reff
Fixed angle rotor
1.6
Glycogen
DNA
Swing-out
centrifuge
bucket
Ribosomes, Polysomes
1.8
RNA
2.0
Axis of rotation
1
10
102
103
104
105
106
107
Swing-out bucket rotor
Sedimentation coefficient
(Svedberg, S)
g:
Gravitational acceleration
g = ω2 · reff
s:
Sedimentation coefficient
(S = 10-13 s)
v:
Sedimentation velocity
M:
Molecular mass
(cm · s-1)
v = ω2 · reff · s
v :
Partial specific particle volume
(cm3 · g-1)
ω:
Angular velocity
r :
Density of the solution
(rad · s-1)
M · (1 - v · r)
(g · cm3)
s =
reff:
Effective radius (cm)
f
f :
Coefficient of friction
B. Density gradient centrifugation
Particles migrate
Particles distributed
Probe
according to S-value
according to density
Centrifugation
Centrifugation
Sucrose
CsCl
density
density
gradient
gradient
After t =
0
0.5
1
2 h
0
2
4
8 h
Zonal centrifugation
Isopyknic centrifugation
Sucrose or
CsCl gradient
1
2
3
4
5
6
7
8
9
10
11
0
1
2
3
4
5
6
7
8
9
Fraction
Fraction
Fractionation
Detection
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
202
Organelles
tors,
30 aminoacyl-tRNA synthases, 340
Cell components and cytoplasm
tRNA molecules, 2-3 mRNAs (each of which
is 10 times the length of the section shown),
The Gram-negative bacterium Escherichia coli
and six molecules of RNA polymerase.
(E. coli) is a usually harmless symbiont in the
• About 330 other enzyme molecules, includ-
intestine of mammals. The structure and char-
ing 130 glycolytic enzymes and 100 en-
acteristics of this organism have been partic-
zymes from the tricarboxylic acid cycle.
ularly well characterized. E. coli is also fre-
• 30 000
small organic molecules with
quently used in genetic engineering
(see
masses of 100-1000 Da—e. g., metabolites
p. 258).
of the intermediary metabolism and coen-
zymes. These are shown at a magnification
10 times higher in the bottom right corner.
A. Components of a bacterial cell
• And finally, 50 000 inorganic ions. The rest
consists of water.
A single E. coli cell has a volume of about
0.88 µm3. One-sixth of this consists of mem-
The illustration shows that the cytoplasm of
branes and one-sixth is DNA (known as the
cells is a compartment densely packed with
“nucleoid”). The rest of the internal space of
macromolecules and smaller organic mole-
the cell is known as cytoplasm (not “cytosol”;
cules. The distances between organic mole-
see p. 198).
cules are small. They are only separated by a
The main component of E. coli—as in all
few water molecules.
cells—is water (70%). The other components
All of the molecules are in motion. Due to
are macromolecules (proteins, nucleic acids,
constant collisions, however, they do not ad-
polysaccharides), small organic molecules,
vance in a straight path but move in zigzags.
and inorganic ions. The majority of the macro-
Due to their large mass, proteins are particu-
molecules are proteins, which represent ca.
larly slow. However, they do cover an average
55% of the dry mass of the cell. When a num-
of 5 nm in 1 ms—a distance approximately
ber of assumptions are made about the dis-
equal to their own length. Statistically, a pro-
tribution and size (average mass 40 kDa) of
tein is capable of reaching any point in a
proteins, it can be estimated that there are
bacterial cell in less than a second.
approximately 250 000 protein molecules in
the cytoplasm of an E. coli cell. In eukaryotic
C. Biochemical functions of the cytoplasm
cells, which are about a thousand times larger,
it is estimated that the number of protein
In eukaryotes, the cytoplasm, representing
molecules is in the order of several billion.
slightly more than 50% of the cell volume, is
the most important cellular compartment. It
is the central reaction space of the cell. This is
B. Looking inside a bacterial cell
where many important pathways of the inter-
The illustration shows a schematic view in-
mediary metabolism take place—e. g., glycol-
side the cytoplasm of E. coli, magnified ap-
ysis, the pentose phosphate pathway, the ma-
proximately one million times. At this magni-
jority of gluconeogenesis, and fatty acid syn-
fication, a single carbon atom would be the
thesis. Protein biosynthesis (translation; see
size of a grain of salt, and an ATP molecule
p. 250) also takes place in the cytoplasm. By
would be as large as a grain of rice. The detail
contrast, fatty acid degradation, the tricarbox-
shown is
100 nm long, corresponding to
ylic acid cycle, and oxidative phosphorylation
about 1/600th of the volume of a cell in E.
are located in the mitochondria (see p. 210).
coli. To make the macromolecules clearer,
small molecules such as water, cofactors,
and metabolites have all been omitted from
the illustration. The section of the cytoplasm
shown contains:
• Several hundred macromolecules, which
are needed for protein biosynthesis—i. e.,
30 ribosomes, more than 100 protein fac-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Basics
203
A. Components of a bacterial cell
Proteins
RNA (8%)
DNA (1%)
(17%, 3 000)
Sugars (1%, 250)
Inorganic
ions (1%, 20)
Lipids (1%, 50)
Water
(70%, 1)
Nucleotides
(0.4%, 100)
Amino acids
Number of
(0.4%, 100)
different mole-
Other small
cule types
organic
molecules
(0.2%, 300)
B. View into a bacterial cell
Ribosome
DNA
mRNA
tRNA
Proteins
RNA-Polymerase
Protein
Water
Carbohydrate Amino acid
C. Biochemical functions of the cytoplasm
Schematic
Gluconeogenesis
Pentose phos-
net of the
reactions
phate cycle
in the cytoplasm
Fatty acid
Glycolysis
biosynthesis
Protein
biosynthesis
And many other
reactions
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
204
Organelles
to other cell components (villin, D-actinin,
Cytoskeleton: components
spectrin), or disrupt the helical structure of F
The cytoplasm of eukaryotic cells is traversed
actin (gelsolin). The activity of these proteins
by three-dimensional scaffolding structures
is regulated by protein kinases via Ca2+ and
consisting of filaments (long protein fibers),
other second messengers (see p. 386).
which together form the cytoskeleton. These
filaments are divided into three groups, based
B. Intermediate filaments
on their diameters: microfilaments (6-8 nm),
intermediate filaments (ca. 10 nm), and mi-
The components of the intermediate fila-
crotubules (ca. 25 nm). All of these filaments
ments belong to five related protein families.
are polymers assembled from protein compo-
They are specific for particular cell types. Typ-
nents.
ical representatives include the cytokeratins,
desmin, vimentin, glial fibrillary acidic protein
(GFAP), and neurofilament. These proteins all
A. Actin
have a rod-shaped basic structure in the cen-
Actin, the most abundant protein in eukary-
ter, which is known as a superhelix (“coiled
otic cells, is the protein component of the
coil”; see keratin, p. 70). The dimers are ar-
microfilaments (actin filaments). Actin occurs
ranged in an antiparallel fashion to form tet-
in two forms—a monomolecular form (G actin,
ramers. A staggered head-to-head arrange-
globular actin) and a polymer (F actin, fila-
ment produces protofilaments. Eight protofi-
mentous actin). G actin is an asymmetrical
laments ultimately form an intermediary fil-
molecule with a mass of 42 kDa, consisting
ament.
of two domains. As the ionic strength in-
Free protein monomers of intermediate fil-
creases, G actin aggregates reversibly to
aments rarely occur in the cytoplasm, in con-
form F actin, a helical homopolymer. G actin
trast to microfilaments and microtubules.
carries a firmly bound ATP molecule that is
Their polymerization leads to stable polymers
slowly hydrolyzed in F actin to form ADP.
that have no polarity.
Actin therefore also has enzyme properties
(ATPase activity).
C. Tubulins
As individual G actin molecules are always
oriented in the same direction relative to one
The basic components of the tube-shaped mi-
another, F actin consequently has polarity. It
crotubules are α- and β-tubulin (53
and
has two different ends, at which polymeriza-
55 kDa). These form α,β-heterodimers, which
tion takes place at different rates. If the ends
in turn polymerize to form linear protofila-
are not stabilized by special proteins (as in
ments. Thirteen protofilaments form a ring-
muscle cells), then at a critical concentration
shaped complex, which then grows into a
of G actin the (+) end of F actin will constantly
long tube as a result of further polymeriza-
grow, while the (-) end simultaneously de-
tion.
cays. These partial processes can be blocked
Like microfilaments, microtubules are dy-
by fungal toxins experimentally. Phalloidin, a
namic structures with (+) and (-) ends. The
toxin contained in the Amanita phalloides
(-) end is usually stabilized by bonding to the
mushroom, inhibits decay by binding to the
centrosome. The
(+) end shows dynamic
(-) end. By contrast, cytochalasins, mold tox-
instability. It can either grow slowly or
ins with cytostatic effects, block polymeriza-
shorten rapidly. GTP, which is bound by the
tion by binding to the (+) end.
microtubules and gradually hydrolyzed into
Actin-associated proteins. The cytoplasm
GDP, plays a role in this. Various proteins can
contains more than
50 different proteins
also be associated with microtubules.
that bind specifically to G actin and F actin.
Their actin uptake has various different func-
tions. This type of bonding can serve to regu-
late the G actin pool (example: profilin), influ-
ence the polymerization rate of G actin (vil-
lin), stabilize the chain ends of F actin (fragin,
b-actinin), attach filaments to one another or
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Cytoskeleton
205
A. Actin
Associates
Dissociates
more easily
more easily
ATP
ADP
Polymerization
Pi
Depolymerization
F-actin
ATP
ADP
helical polymer
G-Actin
Slow
Pi
monomers
microfilament (detail)
hydrolysis
42 kDa
Phalloidin
Cytochalasin
(-) End:
dissociation
(+) End:
polymerization
favored
favored
B. Intermediate filaments
IF Proteins:
Cytokeratins
Dimer
Desmin
Vimentin
Glial fibrillary
Superhelical structure
acidic protein
Neurofilaments
Lamins
Tetramer
Protofilament
Intermediate
filament
C. Tubulins
binds GTP and
25 nm
slowly hydrolyzes it
αβ
Protofilament
Tubulin
Microtubule
53 and 55 kDa
(cylindrical polymer)
heterodimer
Plant
alkaloids:
Vinblastine,
vincristine,
colchicine
Taxol
(-) End: stabilized by binding
(+) End: grows
to the centromere
or shortens
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
206
Organelles
B. Microtubules
Structure and functions
Only the cell’s microtubules are shown here.
The cytoskeleton carries out three major
They radiate out in all directions from a center
tasks:
near the nucleus, the centrosome. The tube-
• It represents the cell’s mechanical scaffold-
shaped microtubules are constantly being
ing, which gives it its typical shape and
synthesized and broken down at their (+)
connects membranes and organelles to
ends. In the centriole, the (-) end is blocked
each other. This scaffolding has dynamic
by associated proteins (see p. 204). The (+)
properties; it is constantly being synthe-
end can also be stabilized by associated pro-
sized and broken down to meet the cell’s
teins—e. g., when the microtubules have
requirements and changing conditions.
reached the cytoplasmic membrane.
• It acts as the motor for movement of animal
The microtubules are involved in defining
cells. Not only muscle cells (see p. 332), but
the shape of the cell and also serve as guiding
also cells of noncontractile tissues contain
tracks for the transport of organelles. To-
many different motor proteins, which they
gether with associated proteins (dynein, kine-
use to achieve coordinated and directed
sin), microtubules are able to carry out me-
movement. Cell movement, shape changes
chanical work—e. g., during the transport of
during growth, cytoplasmic streaming, and
mitochondria, the movement of cilia (hair-
cell division are all made possible by com-
like cell protrusions in the lungs, intestinal
ponents of the cytoskeleton.
epithelium, and oviduct) and the beating of
• It serves as a transport track within the cell.
the flagella of sperm. Microtubules also play a
Organelles and other large protein com-
special role in the mitotic period of cell divi-
plexes can move along the filaments with
sion (see p. 394).
the help of the motor proteins.
C. Architecture
A. Microfilaments and intermediate
The complex structure and net-like density of
filaments
the cytoskeleton is illustrated here using
The illustration schematically shows a detail
three examples in which the cytoskeletal
of the microvilli of an intestinal epithelial cell
components are visualized with the help of
as an example of the structure and function of
antibodies.
the components of the cytoskeleton (see also
1. The border of an intestinal epithelial cell
C1).
is seen here (see also B). There are micro-
Microfilaments of F actin traverse the mi-
filaments (a) passing from the interior of the
crovilli in ordered bundles. The microfila-
cell out into the microvilli. The filaments are
ments are attached to each other by actin-as-
firmly held together by spectrin (b), an asso-
sociated proteins, particularly fimbrin and vil-
ciated protein, and they are anchored to in-
lin. Calmodulin and a myosin-like ATPase con-
termediate filaments (c).
nect the microfilaments laterally to the
2. Only microtubules are seen in this fibro-
plasma membrane. Fodrin, another microfila-
blast cell. They originate from the microtubule
ment-associated protein, anchors the actin
organizing center (centrosome) and radiate
fibers to each other at the base, as well as
out as far as the plasma membrane.
attaching them to the cytoplasmic membrane
3. Keratin filaments are visible here in an
and to a network of intermediate filaments. In
epithelial cell. Keratin fibers belong to the
this example, the microfilaments have a
group of intermediate filaments (see pp. 70,
mainly static function. In other cases, actin is
204; d = nucleus).
also involved in dynamic processes. These in-
clude muscle contraction (see p. 332), cell
movement, phagocytosis by immune cells,
the formation of microspikes and lamellipo-
dia (cellular extensions), and the acrosomal
process during the fusion of sperm with the
egg cell.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Cytoskeleton
207
A. Microfilaments and intermediate filaments
End of
Microvillus
F-Actin
Actin
filament
(micro-
filament)
Villin
Mechanical
Fimbrin
framework
Calmodulin
Plasma
Fodrin
membrane
Intermediate
filament
B. Microtubules
2Microtubule -i
associated protein stabilizes end
Secretory
vesicle
Nucleus
Motor for movement
Stable
End grows or
ends
becomes shorter
Nucleus
Centrosome
A mitochondrion
migrates along
Rail for transport
a microtubule
C. Architecture
a
d
b
c
d
1. Microfilaments
2. Microtubules
3. Intermediate filaments
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
208
Organelles
proteins can enter the nucleus without dif -
Nucleus
culty. By contrast, larger proteins
(over
40 kDa) can only pass through the nuclear
A. Nucleus
pores if they carry a nuclear localization se-
The nucleus is the largest organelle in the eu-
quence consisting of four successive basic
karyotic cell. With a diameter of about 10 µm,
amino acids inside their peptide chains (see
it is easily recognizable with the light micro-
p. 228). mRNAs and rRNAs formed in the nu-
scope. This is the location for storage, replica-
cleus cross the pores into the cytoplasm as
tion, and expression of genetic information.
complexes with proteins (see below).
The nucleus is separated from the cyto-
plasm by the nuclear envelope, which consists
C. Relationships between the nucleus and
of the outer and inner nuclear membranes.
cytoplasm
Each of the two nuclear membranes has two
layers, and the membranes are separated
Almost all of the RNA in the cell is synthesized
from each other by the perinuclear space.
in the nucleus. In this process, known as
The outer nuclear membrane is continuous
transcription, the information stored in DNA
with the rough endoplasmic reticulum and
is transcribed into RNA (see p. 242). As men-
is covered with ribosomes. The inner side of
tioned above, ribosomal RNA (rRNA) is mainly
the membrane is covered with a protein layer
produced in the nucleolus, while messenger
(the nuclear lamina), in which the nuclear
and transfer RNA
(mRNA and tRNA) are
structures are anchored.
formed in the region of the euchromatin. En-
The nucleus contains almost all of the cell’s
zymatic duplication of DNA—replication—also
DNA (around 1 % of which is mitochondrial
only takes place in the nucleus (see p. 240).
DNA). Together with histones and structural
The nucleotide components required for
proteins, the nuclear DNA forms the chroma-
transcription and replication have to be im-
tin (see p. 238). It is only during cell division
ported into the nucleus from the cytoplasm.
that chromatin condenses into chromosomes,
Incorporation of these components into RNA
which are also visible with the light micro-
leads to primary products, which are then
scope. During this phase, the nuclear mem-
altered by cleavage, excision of introns, and
brane temporarily disintegrates.
the addition of extra nucleotides (RNA matu-
During the phase between cell divisions,
ration; see p. 242). It is only once these pro-
the interphase, it is possible to distinguish
cess have been completed that the RNA mol-
between the more densely packed hetero-
ecules formed in the nucleus can be exported
chromatin and loose euchromatin using an
into the cytoplasm for protein synthesis
electron microscope. Active transcription of
(translation; see p. 250).
DNA into mRNA takes place in the region of
The nucleus is not capable of synthesizing
the euchromatin. A particularly electron-
proteins. All of the nuclear proteins therefore
dense region is noticeable in many
have to be imported—the histones with which
nuclei—the nucleolus
(several nucleoli are
DNA is associated in chromatin, and also the
sometimes present). The DNA in the nucleolus
so-called non-histone proteins
(DNA poly-
contains numerous copies of the genes for
merases and RNA polymerases, auxiliary and
rRNAs (see p. 242). They are constantly under-
structural proteins, transcription factors, and
going transcription, leading to a high local
ribosomal proteins). Ribosomal RNA (rRNA)
concentration of RNA.
already associates with proteins in the nucle-
olus to form ribosome precursors.
A special metabolic task carried out by the
B. Nuclear pores
nucleus is biosynthesis of NAD+. The immedi-
The exchange of substances between the nu-
ate precursor of this coenzyme, nicotinamide
cleus and the cytoplasm is mediated by pore
mononucleotide (NMN+), arises in the cyto-
complexes with complicated structures,
plasm and is then transported into the nucle-
which traverse the nuclear membrane. The
olus, where it is enzymatically converted into
nuclear pores consist of numerous proteins
the dinucleotide NAD+. Finally, NAD+ then re-
that form several connected rings of varying
turns to the cytoplasm.
diameter. Low-molecular structures and small
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nucleus
209
A. Nucleus
B. Nuclear pores
10 µm
Cytoplasmic fibrils
Outer
Outer
nuclear
ring
Outer
Inner
membrane
nuclear
nuclear
membrane
membrane
Lumen
Inner
Inner
nuclear
ring
Nuclear pores
Nucleolus
membrane
Basket
Euchromatin
DNA, RNA, histones,
Heterochromatin non-histone proteins
Distal ring
C. Interactions between nucleus and cytoplasm
Nucleoplasm
Cytoplasm
Nucleolus
DNA
Replication
DNA
Transcription
N
N
A
45S-RNA
hnRNAs
pre-tRNAs
Processing
NAD
synthesis
tRNAs
rRNAs
mRNAs
Ribosomal
subunits
Polysome
Translation
N
N
A
Ribosomal
NMN
NAD
proteins,
Histones,
Non-histone proteins
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
210
Organelles
Both mitochondrial membranes are very
Structure and functions
rich in proteins. Porins (see p. 214) in the
outer membrane allow small molecules
A. Mitochondrial structure
(< 10 kDa) to be exchanged between the cy-
Mitochondria are bacteria-sized organelles
toplasm and the intermembrane space. By
(about 1 × 2 µm in size), which are found in
contrast, the inner mitochondrial membrane
large numbers in almost all eukaryotic cells.
is completely impermeable even to small
Typically, there are about 2000 mitochondria
molecules (with the exception of O2, CO2,
per cell, representing around 25% of the cell
and H2O). Numerous transporters in the inner
volume. Mitochondria are enclosed by two
membrane ensure the import and export of
membranes—a smooth outer membrane and
important metabolites (see p. 212). The inner
a markedly folded or tubular inner mitochon-
membrane also transports respiratory chain
drial membrane, which has a large surface
complexes, ATP synthase, and other enzymes.
and encloses the matrix space. The folds of
The matrix is also rich in enzymes (see B).
the inner membrane are known as cristae,
and tube-like protrusions are called tubules.
B. Metabolic functions
The intermembrane space is located between
the inner and the outer membranes.
Mitochondria are also described as being the
The number and shape of the mitochon-
cell’s biochemical powerhouse, since—through
dria, as well as the numbers of cristae they
oxidative phosphorylation (see p. 112)—they
have, can differ widely from cell type to cell
produce the majority of cellular ATP. Pyruvate
type. Tissues with intensive oxidative meta-
dehydrogenase (PDH), the tricarboxylic acid
bolism—e. g., heart muscle—have mitochon-
cycle, β-oxidation of fatty acids, and parts of
dria with particularly large numbers of cris-
the urea cycle are located in the matrix. The
tae. Even within one type of tissue, the shape
respiratory chain, ATP synthesis, and enzymes
of the mitochondria can vary depending on
involved in heme biosynthesis (see p. 192) are
their functional status. Mitochondria are mo-
associated with the inner membrane.
bile, plastic organelles.
The inner membrane itself plays an impor-
Mitochondria probably developed during
tant part in oxidative phosphorylation. As it is
an early phase of evolution from aerobic bac-
impermeable to protons, the respiratory
teria that entered into symbiosis with pri-
chain—which pumps protons from the matrix
meval anaerobic eukaryotes. This endosym-
into the intermembrane space via complexes
biont theory is supported by many findings.
I, III, and IV—establishes a proton gradient
For example, mitochondria have a ring-
across the inner membrane, in which the
shaped DNA (four molecules per mitochon-
chemical energy released during NADH oxi-
drion) and have their own ribosomes. The
dation is conserved (see p. 126). ATP synthase
mitochondrial genome became smaller and
then uses the energy stored in the gradient to
smaller during the course of evolution. In hu-
form ATP from ADP and inorganic phosphate.
mans, it still contains
16 569 base pairs,
Several of the transport systems are also de-
which code for two rRNAs, 22 tRNAs, and 13
pendent on the H+ gradient.
proteins. Only these 13 proteins (mostly sub-
In addition to the endoplasmic reticulum,
units of respiratory chain complexes) are pro-
the mitochondria also function as an
duced in the mitochondrion. All of the other
intracellular calcium reservoir. The mitochon-
mitochondrial proteins are coded by the nu-
dria also play an important role in
“pro-
clear genome and have to be imported into
grammed cell death”—apoptosis (see p. 396).
the mitochondria after translation in the cy-
toplasm (see p. 228). The mitochondrial en-
velope consisting of two membranes also
supports the endosymbiont theory. The inner
membrane, derived from the former sym-
biont, has a structure reminiscent of proka-
ryotes. It contains the unusual lipid cardioli-
pin (see p. 50), but hardly any cholesterol (see
p. 216).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Mitochondria
211
A. Mitochondrial structure
2 µm
Outer membrane
DNA
Inner membrane
ATP
Creatine
Enzymes
synthase
kinase
of
lipid
metabo-
lism
Transporter
Enzymes
Nucleotide
Porin
of
kinases
oxidative
Cristae
metabo-
lism
Matrix space
Respi-
Intermembrane space
ratory
chain
Inter-
Matrix
Inner
membrane
Outer
space
membrane
space
membrane
B. Metabolic functions
Inter-
Pyruvate
Acyl CoA
Carnitine shuttle
membrane
space
Cytoplasmic
calcium ions
Pyruvate
Acyl CoA
Urea cycle
Urea
Ca2
NH3
β-Oxidation
Ca2
-
Acetyl CoA
HCO3
A
P P
ADP
Tri-
CO2
CO2
carboxylic
acid cycle
ETF
H+
H2O
A
A
A
P P P
N
A
P P
P P P
ATP
Respiratory chain
P
Inner membrane
ATP synthesis
H+
H+
H+
H+
Outer membrane
O2
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
212
Organelles
B. Malate and glycerophosphate shuttles
Transport systems
Two systems known as “shuttles” are avail-
Mitochondria are surrounded by an inner and
able to allow the import of reducing equiva-
an outer membrane (see p. 210). The outer
lents that arise from glycolysis in the cyto-
membrane contains porins, which allow
plasm in the form of NADH+H+. There is no
smaller molecules up to 10 kDa in size to
transporter in the inner membrane for
pass. By contrast, the inner membrane is also
NADH+H+ itself.
impermeable to small molecules (with the
In the malate shuttle (left)—which operates
exception of water and the gases O2, CO2,
in the heart, liver, and kidneys, for exam-
and NH3). All of the other substrates of mito-
ple—oxaloacetic acid is reduced to malate by
chondrial metabolism, as well as its products,
malate dehydrogenase (MDH, [2a]) with the
therefore have to be moved through the inner
help of NADH+H+. In antiport for 2-oxogluta-
membrane with the help of special transport-
rate, malate is transferred to the matrix,
ers.
where the mitochondrial isoenzyme for
MDH [2b] regenerates oxaloacetic acid and
A. Transport systems
NADH+H+. The latter is reoxidized by complex
I of the respiratory chain, while oxaloacetic
The transport systems of the inner mitochon-
acid, for which a transporter is not available in
drial membrane use various mechanisms.
the inner membrane, is first transaminated to
Metabolites or ions can be transported alone
aspartate by aspartate aminotransferase (AST,
(uniport, U), together with a second substance
[3a]). Aspartate leaves the matrix again, and
(symport, S), or in exchange for another mol-
in the cytoplasm once again supplies oxalo-
ecule
(antiport, A). Active transport—i. e.,
acetate for step [2a] and glutamate for return
transport coupled to ATP hydrolysis—does
transport into the matrix [3b]. On balance,
not play an important role in mitochondria.
only NADH+H+ is moved from the cytoplasm
The driving force is usually the proton gra-
into the matrix; ATP is not needed for this.
dient across the inner membrane (blue star)
The glycerophosphate shuttle (right) was
or the general membrane potential (red star;
discovered in insect muscle, but is also active
see p. 126).
in the skeletal musculature and brain in
The pyruvate (left) formed by glycolysis in
higher animals. In this shuttle, NADH+H+
the cytoplasm is imported into the matrix in
formed in the cytoplasm is used to reduce
antiport with OH-. The OH- ions react in the
glycerone 3-phosphate, an intermediate of
intermembrane space with the H+ ions abun-
glycolysis
(see p. 150) to glycerol
3-phos-
dantly present there to form H2O. This main-
phate. Via porins, this enters the intermem-
tains a concentration gradient of OH-. The
brane space and is oxidized again there on the
import of phosphate (H2PO–) is driven in a
exterior side of the inner membrane by the
similar way. The exchange of the ATP formed
flavin enzyme glycerol 3-phosphate dehydro-
in the mitochondrion for ADP via an adenine
genase back into glycerone 3-phosphate. The
nucleotide translocase (center) is also depen-
reducing equivalents are passed on to the
dent on the H+ gradient. ATP with a quadruple
respiratory chain via ubiquinone (coenzyme
negative charge is exchanged for ADP with a
Q).
triple negative charge, so that overall one
The carnitine shuttle for transporting acyl
negative charge is transported into the H+-
residues into the mitochondrial matrix is dis-
rich intermembrane space. The import of ma-
cussed on p. 164.
late by the tricarboxylate transporter, which is
important for the malate shuttle (see B) is
coupled to the export of citrate, with a net
export of one negative charge to the exterior
again. In the opposite direction, malate can
leave the matrix in antiport for phosphate.
When Ca2+ is imported (right), the metal cat-
ion follows the membrane potential. An elec-
troneutral antiport for two H+ or two Na+
serves for Ca2+ export.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Mitochondria
213
A. Transport systems
Driving force:
ATP4
2-Oxoglutarate/citrate
Membrane
potential
ADP3
Proton
2-oxo-
Malate-
gradient
glutarate/
citrate
A
A
H2PO4
ATP4
H2O
HPO42
Malate-
ADP3
A
Citrate export
H2PO4
Malate shuttle
OH
2
A
HPO4
H+
U
OH
ATP synthesis
Ca2+
Pyruvate
A
Ca2+
2+ storage
Ca
+
H
Breakdown
Ca2+
release
Pyruvate
Conversion
(Na + )
A
Inner mitochondrial membrane
B. Malate and glycerophosphate shuttle
1
Glyceraldehyde
2a
2b
Malate dehydrogenase 1.1.1.37
4
Glycerol 3-phosphate dehydrogenase 1.1.1.8
3-phosphate
dehydrogenase
3a
3b
5
Aspartate transaminase 2.6.1.1
Glyceral 3-phosphate DH (FAD) 1.1.99.5
1.2.1.12
Glucose 6-
NADH + H
+
phosphate
1,3-Bisphos-
Glyco-
Cyto-
Cytoplasma
phoglycerate
lysis
plasm
Glu
Oxalo-
N
A
Fructose 1,6-
acetate
1
bisphosphate
3b
2a
Glyceral 3-
Glycerone 3-
N
A
phosphate
phosphate
Asp
2-Oxo-
glutarate
Malate
4
Outer
Glycerophos-
Glycerol 3-
phate shuttle
phosphate
Mitochon-
Malate
drial
membrane
shuttle
Porin
Complex I
Ubiquinone
Inner
I
2-Oxo-
Malate
Q
Asp
glutarate
N
A
5
3a
2b
Complex III
Oxalo-
Glu
N
A
acetate
Complex IV
Matrix
NADH + H
+
2H+ , O
Mitochondrion
H2O
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
214
Organelles
B. Membrane lipids
Structure and components
The illustration shows a model of a small
A. Structure of the plasma membrane
section of a membrane. The phospholipids
are the most important group of membrane
All biological membranes are constructed ac-
lipids. They include phosphatidylcholine
cording to a standard pattern. They consist of
(lecithin), phosphatidylethanolamine, phos-
a continuous bilayer of amphipathic lipids ap-
phatidylserine,
phosphatidylinositol,
and
proximately 5 nm thick, into which proteins
sphingomyelin
(for their structures, see
are embedded. In addition, some membranes
p. 50). In addition, membranes in animal cells
also carry carbohydrates (mono- and oligo-
also contain cholesterol (with the exception
saccharides) on their exterior, which are
of inner mitochondrial membranes). Glycoli-
bound to lipids and proteins. The proportions
pids (a ganglioside is shown here) are mainly
of lipids, proteins, and carbohydrates differ
found on the outside of the plasma mem-
markedly depending on the type of cell and
brane. Together with the glycoproteins, they
membrane (see p. 216).
form the exterior coating of the cell (the gly-
Membrane lipids are strongly amphipathic
cocalyx).
molecules with a polar hydrophilic
“head
group” and an apolar hydrophobic “tail.” In
membranes, they are primarily held together
C. Membrane proteins
by the hydrophobic effect (see p. 28) and
Proteins can be anchored in or on membranes
weak Van der Waals forces, and are therefore
in various ways. Integral membrane proteins
mobile relative to each other. This gives mem-
cross right through the lipid bilayer. The sec-
branes a more or less fluid quality.
tions of the peptide chains that lie within the
The fluidity of membranes primarily de-
bilayer usually consist of 20 to 25 mainly
pends on their lipid composition and on tem-
hydrophobic amino acid residues that form a
perature. At a specific transition temperature,
right-handed α-helix.
membranes pass from a semicrystalline state
Type I and II membrane proteins only
to a more fluid state. The double bonds in the
contain one transmembrane helix of this
alkyl chains of unsaturated acyl residues in
type, while type III proteins contain several.
the membrane lipids disturb the semicrystal-
Rarely, type I and II polypeptides can aggre-
line state. The higher the proportion of unsa-
gate to form a type IV transmembrane pro-
turated lipids present, therefore, the lower the
tein. Several groups of integral membrane
transition temperature. The cholesterol con-
proteins—e. g., the porins (see p. 212)—pene-
tent also influences membrane fluidity. While
trate the membrane with antiparallel β-sheet
cholesterol increases the fluidity of semicrys-
structures. Due to its shape, this tertiary
talline, closely-packed membranes, it stabil-
structure is known as a “β-barrel.”
izes fluid membranes that contain a high pro-
Type V and VI proteins carry lipid anchors.
portion of unsaturated lipids.
These are fatty acids (palmitic acid, myristic
Like lipids, proteins are also mobile within
acid), isoprenoids (e. g., farnesol), or glycoli-
the membrane. If they are not fixed in place
pids such as glycosyl phosphatidylinositol
by special mechanisms, they float within the
(GPI) that are covalently bound to the peptide
lipid layer as if in a two-dimensional liquid;
chain.
biological membranes are therefore also de-
Peripheral membrane proteins are associ-
scribed as being a “fluid mosaic.”
ated with the head groups of phospholipids
Lipids and proteins can shift easily within
or with another integral membrane protein
one layer of a membrane, but switching be-
(not shown).
tween the two layers (“flip/flop”) is not possi-
ble for proteins and is only possible with dif-
ficulty for lipids (with the exception of cho-
lesterol). To move to the other side, phospho-
lipids require special auxiliary proteins
(translocators, “flipases”).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Biological membranes
215
A. Structure of the plasma membrane
Extracellular
side
Glycoprotein
Oligosaccharide
Phospholipid
Lipid
Glycolipid
bilayer
Peripheral
membrane
Integral
Cytoplasmic
protein
membrane proteins
side
B. Membrane lipids
a
b
c
d
a
Phosphatidyl-
inositol
b
Sphingomyelin
Polar
c
Ganglioside
d
Phosphatidyl-
Outer
ethanolamine
leaflet
e
Cerebroside
Apolar
f
Phosphatidyl-
choline
Inner
g
Cholesterol
leaflet
h
Phosphatidyl-
serine
Phospholipids
Polar
Glycolipids
Cholesterol
e
f
g
h
C. Membrane proteins
Type I Type II
Type III
Type IV
Type V
N
C
TypeVI
C
N
Lipid
anchor
1. α-Helical
2. β-Barrel
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
216
Organelles
p. 140) and photosynthesis (see p. 128)—also
Functions and composition
occur in membranes.
The most important membranes in animal
5. Interactions with other cells for the pur-
cells are the plasma membrane, the inner
poses of cell fusion and tissue formation, as
and outer nuclear membranes, the mem-
well as communication with the extracellular
branes of the endoplasmic reticulum (ER) and
matrix.
the Golgi apparatus, and the inner and outer
6. Anchoring of the cytoskeleton
(see
mitochondrial membranes. Lysosomes, peroxi-
p. 204) to maintain the shape of cells and
somes, and various vesicles are also separated
organelles and to provide the basis for move-
from the cytoplasm by membranes. In plants,
ment processes.
additional membranes are seen in the plastids
and vacuoles. All membranes show pola-
B. Composition of membranes
rity—i. e., there is a difference in the composi-
tion of the inner layer (facing toward the
Biological membranes consist of lipids, pro-
cytoplasm) and the outer layer (facing away
teins, and carbohydrates (see p. 214). These
from it).
components occur in varying proportions
(left). Proteins usually account for the largest
proportion, at around half. By contrast, carbo-
A. Functions of membranes
hydrates, which are only found on the side
Membranes and their components have the
facing away from the cytoplasm, make up
following functions:
only a few percent. An extreme composition
is seen in myelin, the insulating material in
1. Enclosure and insulation of cells and or-
nerve cells, three-quarters of which consists
ganelles. The enclosure provided by the
of lipids. By contrast, the inner mitochondrial
plasma membrane protects cells from their
membrane is characterized by a very low pro-
environment both mechanically and chemi-
portion of lipids and a particularly high pro-
cally. The plasma membrane is essential for
portion of proteins.
maintaining differences in the concentration
When the individual proportions of lipids
of many substances between the intracellular
in membranes are examined more closely
and extracellular compartments.
(right part of the illustration), typical patterns
2. Regulated transport of substances,
for particular cells and tissues are also found.
which determines the internal milieu and is
The illustration shows the diversity of the
a precondition for homeostasis—i.e., the
membrane lipids and their approximate
maintenance of constant concentrations of
quantitative composition. Phospholipids are
substances and physiological parameters.
predominant in membrane lipids in compar-
Regulated and selective transport of substan-
ison with glycolipids and cholesterol. Triacyl-
ces through pores, channels, and transporters
glycerols (neutral fats) are not found in mem-
(see p. 218) is necessary because the cells and
branes.
organelles are enclosed by membrane sys-
Cholesterol is found almost exclusively in
tems.
eukaryotic cells. Animal membranes contain
3. Reception of extracellular signals and
substantially more cholesterol than plant
transfer of these signals to the inside of the
membranes, in which cholesterol is usually
cell (see pp. 384 ff.), as well as the production
replaced by other sterols. There is no choles-
of signals.
terol at all in prokaryotes (with a few excep-
4. Enzymatic catalysis of reactions. Impor-
tions). The inner mitochondrial membrane of
tant enzymes are located in membranes at the
eukaryotes is also low in cholesterol, while it
interface between the lipid and aqueous
is the only membrane that contains large
phases. This is where reactions with apolar
amounts of cardiolipin. These facts both sup-
substrates occur. Examples include lipid
port the endosymbiotic theory of the devel-
biosynthesis (see p. 170) and the metabolism
opment of mitochondria (see p. 210).
of apolar xenobiotics (see p. 316). The most
important reactions in energy conver-
sion—i. e., oxidative phosphorylation
(see
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Biological membranes
217
A. Functions of membranes
S
A B
3
2
4
1
5
A+B
A B
C+D
6
Boundary
Controlled
Signal re-
Enzymatic
Contact
Anchor
metabolite
ception and
reactions
with
for cyto-
transport
transmission
other cells
skeleton
B. Composition of membranes
Membrane components
Relative proportion of lipids
Nerve cell: Plasma membrane
Erythrocyte: Plasma membrane
*
*
*
*
*
*
Cardiolipin
Liver cell: Plasma membrane
Mitochondrion
Inner membrane
Both membranes
Phosphatidyl-
Lipids
choline
*
Phosphatidyl-
Glycolipids
Carbohydrates
serine
*
Phospholipids
Phosphatidyl-
Cholesterol
Proteins
ethanolamine
*
Sphingomyelin
Other lipids
*
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
218
Organelles
potential also plays a role; the processes are
Transport processes
summed up by the term “electrochemical
gradient” (see p. 126). These processes there-
A. Permeability
fore involve passive transport, which runs
Only small, uncharged molecules such as
“downhill” on the slope of a gradient.
gases, water, ammonia, glycerol, or urea are
By contrast, active transport can also run
able to pass through biological membranes by
“uphill”—i. e., against a concentration or
free diffusion. With increasing size, even com-
charge gradient. It therefore requires an input
pounds of this type are no longer able to pass
of energy, which is usually supplied by the
through. Membranes are impermeable to glu-
hydrolysis of ATP (see p. 124). The transporter
cose and other sugars, for example.
first binds its “cargo” on one side of the mem-
The polarity of a molecule is also impor-
brane. ATP-dependent phosphorylation then
tant. Apolar substances, such as benzene,
causes a conformation change that releases
ethanol, diethyl ether, and many narcotic
the cargo on the other side of the membrane
agents are able to enter biological membranes
(see p. 220). A non-spontaneous transport
easily. By contrast, membranes are imperme-
process can also take place through coupling
able to strongly polar compounds, particu-
to another active transport process (known as
larly those that are electrically charged. To
secondary active transport; see p. 220).
be able to take up or release molecules of
Using the transport systems in the mem-
this type, cells have specialized channels and
branes, cells regulate their volume, internal
transporters in their membranes (see below).
pH value, and ionic environment. They con-
centrate metabolites that are important for
energy metabolism and biosynthesis, and ex-
B. Passive and active transport
clude toxic substances. Transport systems
Free diffusion is the simplest form of mem-
also serve to establish ion gradients, which
brane transport. When it is supported by in-
are required for oxidative phosphorylation
tegral membrane proteins, it is known as fa-
and stimulation of muscle and nerve cells,
cilitated diffusion (or facilitated transport).
for example (see p. 350).
1. Channel proteins have a polar pore
through which ions and other hydrophilic
C. Transport processes
compounds can pass. For example, there are
Another classification of transport processes
channels that allow selected ions to pass (ion
is based on the number of particles trans-
channels; see p. 222) and porins that allow
ported and the direction in which they
molecules below a specific size to pass in a
move. When a single molecule or ion passes
more or less nonspecific fashion (see p. 212).
through the membrane with the help of a
2. Transporters recognize and bind the
channel or transporter, the process is de-
molecules to be transported and help them
scribed as a uniport (example: the transport
to pass through the membrane as a result of a
of glucose into liver cells). Simultaneous
conformational change. These proteins (per-
transport of two different particles can take
meases) are thus comparable with enzy-
place either as a symport (example: the trans-
mes—although with the difference that they
port of amino acids or glucose together with
“catalyze” vectorial transport rather than an
Na+ ions into intestinal epithelial cells) or as
enzymatic reaction. Like enzymes, they show
an antiport. Ions are often transported in an
a certain affinity for each molecule trans-
antiport in exchange for another similarly
ported
(expressed as the dissociation
charged ion. This process is electroneutral
constant, Kd in mol L-1) and a maximum
and therefore more energetically favorable
transport capacity (V).
(example: the exchange of HCO3- for Cl- at
Free diffusion and transport processes fa-
the erythrocyte membrane).
cilitated by ion channels and transport pro-
teins always follow a concentration gradient—
i. e., the direction of transport is from the
site of higher concentration to the site of
lower concentration. In ions, the membrane
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Biological membranes
219
A. Permeability of membranes
B. Passive and active transport
Electrochemical gradient
Small molecules
Charge gradient
Apolar
O2, N2
Benzene
Polar, uncharged
Free
diffusion
High
Low
H2O
concentration
concentration
Urea
Glycerol
CO2 , NH3
Free
diffusion
Channel
protein
Large molecules
Pore
Polar, uncharged
e.g. glucose
Passive
transport
Ions
No
membrane
H , Na , K , Mg2
passage
Transporter
Facilitated
Ca2 , NH4
diffusion
HCO3 , Cl , H2PO4
Amino acids
Nucleotides
Confor-
mational
change
C. Transport processes
A
Glucose
P P P
Uniport
U
A
P P
P
Glucose
P
Transport
ATPase
Na
H2O
S
Symport
Confor-
mational
Active
change
transport
P
Cl
Cl
HCO
3
HCO
3
A
Antiport
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
220
Organelles
C. Aquaporin-1
Transport proteins
Aquaporins help water to pass through bio-
Illustrations B-D show transporters whose
logical membranes. They form hydrophilic
structure has been determined experimen-
pores that allow H2O molecules, but not hy-
tally or established on analogy with other
drated ions or larger molecules, to pass
known structures. They all belong to group
through. Aquaporins are particularly impor-
III of the α-helical transmembrane proteins
tant in the kidney, where they promote the
(see p. 214).
reuptake of water (see p. 328). Aquaporin-2
in the renal collecting ducts is regulated by
A. Transport mechanisms
antidiuretic hormone
(ADH, vasopressin),
which via cAMP leads to shifting of the chan-
Some cells couple the “pure” transport forms
nels from the ER into the plasma membrane.
discussed on p. 218—i. e., passive transport (1)
Aquaporin-1, shown here, occurs in the
and active transport (2)—and use this mech-
proximal tubule and in Henle’s loop. It con-
anism to take up metabolites. In secondary
tains eight transmembrane helices with dif-
active transport (3), which is used for exam-
ferent lengths and orientations. The yellow-
ple by epithelial cells in the small intestine
colored residues form a narrowing that only
and kidney to take up glucose and amino
H2O molecules can overcome.
acids, there is a symport (S) located on the
luminal side of the membrane, which takes up
the metabolite M together with an Na+ ion.
D. Sarcoplasmic Ca2+ pump
An ATP-dependent Na+ transporter (Na+/K+
Transport ATPases transport cations—they are
ATPase; see p. 350) on the other side keeps
“ion pumps.” ATPases of the F type—e. g., mito-
the intracellular Na+ concentration low and
chondrial ATP synthase (see p. 142)—use H+
thus indirectly drives the uptake of M. Finally,
transport for ATP synthesis. Enzymes of the V
a uniport (U) releases M into the blood.
type, using up ATP, “pump” protons into lyso-
somes and other acidic cell compartments (see
B. Glucose transporter Glut-1
p. 234). P type transport ATPases are particu-
larly numerous. These are ATP-driven cation
The glucose transporters (Glut) are a family of
transporters that undergo covalent phosphor-
related membrane proteins with varying dis-
ylation during the transport cycle.
tribution in the organs. Glut-1 and Glut-3
The Ca2+ ATPase shown also belongs to the
have a relatively high af nity for glucose (Kd
P type. In muscle, its task is to pump the Ca2+
= 1 mM). They occur in nearly all cells, and
released into the cytoplasm to trigger muscle
ensure continuous glucose uptake. Glut-2 is
contraction back into the sarcoplasmic retic-
found in the liver and pancreas. This form has
ulum (SR; see p. 334). The molecule (1) con-
a lower af nity (Kd = 15-20 mM). The rate of
sists of a single peptide chain that is folded into
glucose uptake by Glut-2 is therefore strongly
various domains. In the transmembrane part,
dependent on the blood glucose level (nor-
which is formed by numerous α-helices, there
mally 4-8 mM). Transport by Glut-4 (Kd =
are binding sites for two Ca2+ ions (blue) ATP
5 mM), which is mainly expressed in muscle
is bound to the cytoplasmic N domain (green).
and fat cells, is controlled by insulin, which
Four different stages can be distinguished
increases the number of transporters on the
in the enzyme’s catalytic cycle (2). First, bind-
cell surface (see p. 388). Glut-5 mediates sec-
ing of ATP to the N domain leads to the uptake
ondary active resorption of glucose in the in-
of two Ca2+ into the transmembrane part (a).
testines and kidney (see A).
Phosphorylation of an aspartate residue in the
Glut-1 consists of a single peptide chain
P domain (b) and dissociation of ADP then
that spans the membrane with 12 α-helices
causes a conformation change that releases
of different lengths. The glucose is bound by
the Ca2+ ions into the SR (c). Finally, dephos-
the peptide loops that project on each side of
phorylation of the aspartate residue restores
the membrane.
the initial conditions (d).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Biological membranes
221
A. Transport mechanisms
Epithelial cells
N
A
P P P
Na
E
Na
S
A
P P
M
M
+
P
A
+
P
P P
A
M
P P P
M
M
U
1. Facilitated diffusion
2. Active transport
3. Secondary active transport
B. Glucose transporter Glut-1
D. Sarcoplasmic Ca2
pump
Binding
N
ATP
site for
ATP
P
β
N
C
Glucose
Blood
“Stalk”
Cyto-
plasm
Plasma
membrane
Binding
Trans-
sites
membrane
for Ca2
domain
Cytoplasm
SR
Glucose
1. Structure
A
C. Aquaporin-1
P
P
P
Ca2
A
P
P
P
A
P P
P
a)
N
C
Tubular
lumen
H2O
d)
P
b)
c)
Plasma
membrane
A
P
P P
Tubule cell
2. Catalytic cycle
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
222
Organelles
tion, the S4 helices are thought to snap up-
Ion channels
wards like springs and thus open the central
Ion channels facilitate the diffusion of ions
pore (b).
through biological membranes. Some ion
channels open and close depending on the
B. Nicotinic acetylcholine receptor
membrane potential (voltage-gated channels,
A) or in response to specific ligands (ligand-
Many receptors for neurotransmitters func-
gated channels, B). Other channels operate
tion as ligand-gated channels for Na+, K+
passively. In these cases, transport depends
and/or Ca2+ ions (see p. 354). The ones that
only on the concentration gradient (C).
have been studied in the greatest detail are
the nicotinic receptors for acetylcholine (see
p. 352). These consist of five separate but
A. Voltage-gated Na+ channel
structurally closely related subunits. Each
Voltage-gated Na+ channels play a decisive
forms four transmembrane helices, the sec-
part in the conduction of electrical impulses
ond of which is involved in the central pore in
in the nervous system (see p. 350). These
each case. The type of monomer and its ar-
channels open when the membrane potential
rangement in the complex is not identical in
in their environment reverses. Due to the high
all receptors of this type. In the neuromuscu-
equilibrium potential for Na+ (see p. 126), an
lar junction
(see p. 334), the arrangement
inflow of Na+ ions takes place, resulting in
αβγαδ is found (1).
local depolarization of the membrane, which
In the interior of the structure, acetylcho-
propagates by activation of neighboring volt-
line binds to the two α-subunits and thus
age-dependent Na+ channels. A spreading de-
opens the pore for a short time (1-2 ms).
polarization wave of this type is known as an
Negatively charged residues are arranged in
action potential (see p. 350). Externally di-
three groups in a ring shape inside it. They are
rected K+ channels are involved in the repola-
responsible for the receptor’s ion specificity. It
rization of the membrane. In their function-
is thought that binding of the neurotransmit-
ing, these resemble the much more simply
ter changes the position of the subunits in
structured K+ channels shown in C. The Ca2+
such a way that the pore expands (3). The
channels that trigger exocytosis of vesicles
bound acetylcholine dissociates again and is
(see p. 228) are also controlled by the action
hydrolytically inactivated
(see p. 356). The
potential.
receptor is thus able to function again.
The voltage-gated Na+ channels in higher
animals are large complexes made up of sev-
C. K+ channel in Streptomyces lividans
eral subunits. The α-subunit shown here me-
diates Na+ transport. It consists of a very long
The only detailed structures of ion channels
peptide chain (around 2000 amino acid resi-
established so far are those of potassium
dues), which is folded into four domains, each
channels like that of an outwardly directed
with six transmembrane helices (left). The S6
K+ channel in the bacterium Streptomyces liv-
helices of all the domains
(blue) together
idans. It consists of four identical subunits
form a centrally located hydrophilic pore
(blue, yellow, green, and red), each of which
which can be made narrow or wide depend-
contains two long α-helices and one shorter
ing on the channel’s functional status. The six
one. In the interior of the cell (bottom), the K+
S4 helices (green) function as voltage sensors.
ions
(violet) enter the structure’s central
The current conception of the way in which
channel. Before they are released to the out-
the opening and closing mechanism functions
side, they have to pass through what is known
is shown in a highly simplified form on the
as a “selectivity filter.” In this part of the
right. For the sake of clarity, only one of the
channel, several C=O groups in the peptide
four domains (domain IV) is shown. The S4
chain form a precisely defined opening that
helices contain several positively charged res-
is only permeable to non-hydrated K+ ions.
idues. When the membrane is polarized (a),
the surplus negative charges on the inner side
keep the helix in the membrane. If this attrac-
tion is removed as a result of local depolariza-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Biological membranes
223
A. Voltage-gated Na+ channel
I
II
III
IV
2
3
1
4
1
2
3
4
5
6
6
5
N
C
a
Membrane
Narrow pore
polarized,
helix 4
Na
in position 1
Pore
α-Subunit
4
3
2
Voltage-
Outside
5
1
sensitive
6
helix
(helix 4)
Inside
Membrane
b
depolarized,
Wide pore
helix 4
in position 2
1. Structure
2. Mechanism
Na
B. Nicotinic acetylcholine receptor
N α C
δ
β
α
γ
δ β
αL
α
H
Acetyl-
choline
Subunits
Na
Helix 2
1. Structure
2. Pore
3. Mechanism
C. K + channel in Streptomyces lividans
A
B
C
D
Outside
Plasma-
membrane
Subunits
Outside
Inside
Plasma
membrane
Inside
K
Homotetramer
From above
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
224
Organelles
B. Insulin receptor
Membrane receptors
The receptor for the hormone insulin (see
To receive and pass on chemical or physical
p. 76) belongs to the family of 1-helix recep-
signals, cells are equipped with receptor pro-
tors.
teins. Many of these are integral membrane
These molecules span the membrane with
proteins in the plasma membrane, where
only one α-helix. The subunits of the dimeric
they receive signals from their surroundings.
receptor (red and blue) each consist of two
Other receptor proteins are located in inter-
polypeptides (α and β) bound by disulfide
cellular membranes. The receptors for lipo-
bonds. The α-chains together bind the insulin,
philic hormones are among the few that func-
while the β-chains contain the transmem-
tion in a soluble form. They regulate gene
brane helix and, at the C-terminus, domains
transcription in the nucleus (see p. 378).
with tyrosine kinase activity. In the activated
state, the kinase domains phosphorylate
A. Principle of receptor action
themselves and also mediator proteins (re-
ceptor substrates) that set in motion cascades
Membrane-located receptors can be divided
of further phosphorylations (see pp. 120 and
into three parts, which have different tasks.
388).
The receptor domain reacts specifically to a
given signal. Signals of this type can be of a
purely physical nature. For example, many
C. 7-helix receptors
organisms react to light. In this way, plants
A large group of receptors span the mem-
adapt growth and photosynthesis to light
brane with α-helices seven times. These are
conditions, while animals need light recep-
known as 7-helix receptors. Via their effector
tors for visual processing (C; see p. 358). Me-
domains, they bind and activate trimeric pro-
chanoreceptors are involved in hearing and in
teins, which in turn bind and hydrolyze GTP
pressure regulation, among other things.
and are therefore called G proteins. Most G
Channels that react to action potentials (see
proteins, in turn, activate or inhibit enzymes
p. 350) can be regarded as receptors for elec-
that create secondary signaling molecules
trical impulses.
(second messengers; see p. 386). Other G pro-
However, most receptors do not react to
teins regulate ion channels. The illustration
physical stimuli, but rather to signal mole-
shows the complex of the light receptor rho-
cules. Receptors for these chemical signals
dopsin, with the associated G protein trans-
contain binding sites in the receptor domain
ducin (see p. 358). The GTP-binding α-subunit
that are complementary to each ligand. In this
(green) and the γ-subunit (violet) of transdu-
respect, they resemble enzymes (see p. 94).
cin are anchored in the membrane via lipids
As the effector domain of the receptor is usu-
(see p. 214). The β-subunit is shown in detail
ally separated by a membrane, a mechanism
on p. 72.
for signal transfer between the domains is
needed. Little is yet known regarding this. It
is thought that conformation changes in the
D. T-cell receptor
receptor protein play a decisive part. Some
The cells of the immune system communicate
receptors dimerize after binding of the ligand,
with each other particularly intensively. The
thereby bringing the effector domains of two
T-cell receptor plays a central role in the acti-
molecules into contact (see p. 392).
vation of T lymphocytes (see p. 296). The cell
The way in which the effector works differs
at the top has been infected with a virus, and
from case to case. By binding or interconver-
it indicates this by presenting a viral peptide
sion, many receptors activate special media-
(violet) with the help of a class I MHC protein
tor proteins, which then trigger a signal cas-
(yellow and green). The combination of the
cade (signal transduction; see p. 384). Other
two molecules is recognized by the dimeric
receptors function as ion channels. This is
T-cell receptor (blue) and converted into a
particularly widespread in receptors for neu-
signal that activates the T cell (bottom) and
rotransmitters (see p. 354).
thereby enhances the immune response to
the virus.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Biological membranes
225
A. Principle of receptor action
Signal
Chemical signals
Physical signals
Metabolites
Light
Recipient
Hormones
Electrical impulses
Mechanical stimuli
Neurotransmitters
Mediators
Transmitter
Odorants
Tastants
Activator
Other molecules
Protein kinase
Inorganic ions
Effector
Ion channel
Cellular response
Mediator
Cellular response
Protein activated
G-protein
by ions
receptor substrate
B. Insulin receptor
D. T-cell receptor
α2
α1
Antigen-presenting cell
Insulin
Tyrosine
β2
β1
kinase
domain
α2
α1
MHC
β2
P
P
β1
protein
(class I)
P
P
β2-
Adaptor
Adaptor
Micro-
protein
protein
globulin
Receptor
Protein kinase
substrate
Protein kinase
C. 7-Helix receptors
Viral
peptide
Lipid
Receptor
anchor
α-Sub-
unit
GTP
G protein
T-cell
β-Sub-
receptor
unit
γ-Sub-
unit
Signaling
G protein
Effector
substance
enzyme
GDP
Activated
Second
receptor
GDP
GTP
Precursor
messenger
T-helper cell
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
226
Organelles
will take and whether its transport will be
ER: structure and function
constitutive or regulated depends on the sig-
The endoplasmic reticulum (ER) is an exten-
nal sequences or signal structures that pro-
sive closed membrane system consisting of
teins carry with them like address labels (see
tubular and saccular structures. In the area
p. 228). In addition to proteins, the Golgi ap-
of the nucleus, the ER turns into the external
paratus also transports membrane lipids to
nuclear membrane. Morphologically, a dis-
their targets.
tinction is made between the rough ER (rER)
and the smooth ER (sER). Large numbers of
B. Smooth endoplasmic reticulum
ribosomes are found on the membranes of the
rER, which are lacking on the sER. On the
Regions of the ER that have no bound ribo-
other hand, the sER is rich in membrane-
somes are known as the smooth endoplasmic
bound enzymes, which catalyze partial reac-
reticulum (sER). In most cells, the proportion
tions in the lipid metabolism as well as bio-
represented by the sER is small. A marked sER
transformations.
is seen in cells that have an active lipid me-
tabolism, such as hepatocytes and Leydig
cells. The sER is usually made up of branching,
A. Rough endoplasmic reticulum and the
closed tubules.
Golgi apparatus
Membrane-located enzymes in the sER
The rER
(1) is a site of active protein
catalyze lipid synthesis. Phospholipid synthe-
biosynthesis. This is where proteins destined
sis (see p. 170) is located in the sER, for exam-
for membranes, lysosomes, and export from
ple, and several steps in cholesterol biosyn-
the cell are synthesized. The remaining pro-
thesis (see p. 172) also take place there. In
teins are produced in the cytoplasm on ribo-
endocrine cells that form steroid hormones, a
somes that are not bound to membranes.
large proportion of the reaction steps in-
Proteins synthesized at the rER (1) are
volved also take place in the sER (see p. 376).
folded and modified after translation (protein
In the liver’s hepatocytes, the proportion
maturation; see p. 230). They remain either in
represented by the sER is particularly high.
the rER as membrane proteins, or pass with
It contains enzymes that catalyze so-called
the help of transport vesicles (2) to the Golgi
biotransformations. These are reactions in
apparatus (3). Transport vesicles are formed
which apolar foreign substances, as well as
by budding from existing membranes, and
endogenous substances—e. g., steroid hormo-
they disappear again by fusing with them
nes—are chemically altered in order to inacti-
(see p. 228).
vate them and/or prepare them for conjuga-
The Golgi apparatus (3) is a complex net-
tion with polar substances (phase I reactions;
work, also enclosed, consisting of flattened
see p. 316). Numerous cytochrome P450
membrane saccules (“cisterns”), which are
enzymes are involved in these conversions
stacked on top of each other in layers. Pro-
(see p. 318) and can therefore be regarded
teins mature here and are sorted and packed.
as the major molecules of the sER.
A distinction is made between the cis, medial,
The sER also functions as an intracellular
and trans Golgi regions, as well as a trans
calcium store, which normally keeps the Ca2+
Golgi network (tGN). The post-translational
level in the cytoplasm low. This function is
modification of proteins, which starts in the
particularly marked in the sarcoplasmic retic-
ER, continues in these sections.
ulum, a specialized form of the sER in muscle
From the Golgi apparatus, the proteins are
cells (see p. 334). For release and uptake of
transported by vesicles to various targets in
Ca2+, the membranes of the sER contain sig-
the cells—e. g., to lysosomes (4), the plasma
nal-controlled Ca2+ channels and energy-de-
membrane (6), and secretory vesicles (5) that
pendent Ca2+ ATPases (see p. 220). In the lu-
release their contents into the extracellular
men of the sER, the high Ca2+ concentration is
space by fusion with the plasma membrane
buffered by Ca2+-binding proteins.
(exocytosis; see p. 228). Protein transport can
either proceed continuously (constitutive), or
it can be regulated by chemical signals. The
decision regarding which pathway a protein
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Endoplasmic reticulum and Golgi apparatus
227
A. Rough endoplasmic reticulum and Golgi apparatus
Morphological
Biochemical
structures
Nucleus
processes
1. Transport
2. Protein synthesis
2. and folding
2. Signal peptide
1. rER
2. cleavage
2. Formation of
2. Transport
2. disulfide bonds
2. vesicle
2. Oligomerization
2. N-Glycosylation
cis
folding
medial
3. Golgi complex
3. cis:
trans
3. Phosphorylation
3. medial:
Membrane
trans-Golgi network
3. Sugar cleavage
protein
3. Addition of GlcNAc
4. Lysosome
3. Addition of
3. Gal and NeuAc
5. Secretory
3. trans-Golgi network:
5. vesicle
3. Sorting
4. Hydrolysis of
4. macromolecules
5. Proteolysis
6. Cytoplasmic
6. membrane
6. Exocytosis
B. Smooth endoplasmic reticulum
Steroid
Phospholipids
Cholesterol
hormones
Outside
Calcium store
Plasma
membrane
Xeno-
Metabolites
Inside
biotics
Ca2+
Enzyme systems
of the lipid metabolism
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
228
Organelles
organelles (selective protein transfer). Struc-
Protein sorting
tural signals can also activate enzymes that
modify the proteins and thereby determine
A. Protein sorting
their subsequent fate. Examples include lyso-
The biosynthesis of all proteins starts on free
somal proteins (see p. 234) and membrane
ribosomes (top). However, the paths that the
proteins with lipid anchors (see p. 214).
proteins follow soon diverge, depending on
After they have been used, signal peptides
which target they are destined for. Proteins
at the N terminus are cleaved off by specific
that carry a signal peptide for the ER (1) follow
hydrolases (symbol: scissors). In proteins that
the secretory pathway (right). Proteins that do
contain several successive signal sequences,
not have this signal follow the cytoplasmic
this process can expose the subsequent sig-
pathway (left).
nals. By contrast, signal peptides that have to
Secretory pathway. Ribosomes that syn-
be read several times are not cleaved.
thesize a protein with a signal peptide for
the ER settle on the ER (see p. 228). The pep-
C. Exocytosis
tide chain is transferred into the lumen of the
rER. The presence or absence of other signal
Exocytosis is a term referring to processes
sequences and signal regions determines the
that allow cells to expel substances (e. g., hor-
subsequent transport pathway.
mones or neurotransmitters) quickly and in
Proteins that have stop-transfer sequences
large quantities. Using a complex protein ma-
(4) remain as integral membrane proteins in
chinery, secretory vesicles fuse completely or
the ER membrane. They then pass into other
partially with the plasma membrane and re-
membranes via vesicular transport
(see
lease their contents. Exocytosis is usually
p. 226). From the rER, their pathway then
regulated by chemical or electrical signals. As
leads to the Golgi apparatus and then on to
an example, the mechanism by which neuro-
the plasma membrane. Proteins destined to
transmitters are released from synapses (see
remain in the rER—e. g., enzymes—find their
p. 348) is shown here, although only the most
way back from the Golgi apparatus to the rER
important proteins are indicated.
with the help of a retention signal (2). Other
The decisive element in exocytosis is the
proteins move from the Golgi apparatus to
interaction between proteins known as
the lysosomes (3; see p. 234), to the cell
SNAREs that are located on the vesicular
membrane (integral membrane proteins or
membrane (v-SNAREs) and on the plasma
constitutive exocytosis), or are transported
membrane (t-SNAREs). In the resting state
out of the cell (9; signal-regulated exocytosis)
(1), the v-SNARE synaptobrevin is blocked by
by secretory vesicles (8).
the vesicular protein synaptotagmin. When an
Cytoplasmic pathway. Proteins that do not
action potential reaches the presynaptic
have a signal peptide for the ER are synthe-
membrane, voltage-gated Ca2+ channels
sized in the cytoplasm on free ribosomes, and
open (see p. 348). Ca2+ flows in and triggers
remain in that compartment. Special signals
the machinery by conformational changes in
mediate further transport into the mitochon-
proteins. Contact takes place between synap-
dria (5; see p. 232), the nucleus (6; see p. 208)
tobrevin and the t-SNARE synaptotaxin (2).
or peroxisomes (7).
Additional proteins known as SNAPs bind to
the SNARE complex and allow fusion between
the vesicle and the plasma membrane (3). The
B. Translocation signals
process is supported by the hydrolysis of GTP
Signal peptides are short sections at the N or C
by the auxiliary protein Rab.
terminus, or within the peptide chain. Areas
The toxin of the bacterium Clostridium bot-
on the protein surface that are formed by
ulinum, one of the most poisonous substances
various sections of the chain or by various
there is, destroys components of the exocyto-
chains are known as signal regions. Signal
sis machinery in synapses through enzymatic
peptides and signal regions are structural sig-
hydrolysis, and in this way blocks neurotrans-
nals that are usually recognized by receptors
mission.
on organelles (see A). They move the proteins,
with the help of additional proteins, into the
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Endoplasmic reticulum and Golgi apparatus
229
A. Protein sorting
Cytoplasmic pathway
Secretory pathway
Ribosomes
4
2
-KDEL
+
++
Protein
H
3N
*
Rough ER
1
*
Retention
Cytoplasm
5
*
Retention
-SKL
H2N
Golgi complex
7
6
Receptor
R
R
8
Peroxisomes
Mitochondria
3
Secretory
R
*
vesicle
Lysosomes
2 +
Nucleus
Ca
9
Cell membrane
*Standard pathway (without signal)
B. Translocation signals
C. Exocytosis
++
+
α-Helix
Transmitter
Vesicle
Rab · GTP
H2N
Signal peptide
1
(secretory pathway)
+
OOC
Synapto-
GTP
brevin
Signal sequence
(v-SNARE)
Synapto-
2
(ER proteins)
tagmin
...LEDK
Syntaxin
(t-SNARE)
Mannose 6-
Signal group
phosphate
3
P
Ca2
(lysosomal proteins)
Synaptic cleft
Channel
1.
Apolar
sequence
GTP
Stop-transfer sequence
4
(membrane proteins)
Synaptotagmin
+++
releases
Action
Signal peptide
Ca2
v-SNAREs
potential
5
(mitochondrial
proteins)
Ca2
...KKKRK...
2.
Signal sequence
6
Rab · GDP
(nuclear proteins)
GDP
Signal sequence
7
Membrane
P
(peroxisomes)
...FKS...
fusion
SNAP
+NSF
Signal region
8
(secretory vesicle)
3.
SNARE complex
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
230
Organelles
all plasma proteins with the exception of al-
Protein synthesis and maturation
bumin are glycosylated. Together with glyco-
lipids, numerous glycoproteins on the cell
A. Protein synthesis in the rER
surface form the glycocalyx. Inside the ER,
With all proteins, protein biosynthesis
the carbohydrate parts of the glycoproteins
(Translation; for details, see p. 250) starts on
are cotranslationally transferred to the grow-
free ribosomes in the cytoplasm (1). Proteins
ing chain, and are then converted into their
that are exported out of the cell or into lyso-
final form while passing through the ER and
somes, and membrane proteins of the ER and
Golgi apparatus.
the plasma membrane, carry a signal peptide
N-bound oligosaccharides (see p. 44) are
for the ER at their N-terminus. This is a section
always bound to the acid-amide group of as-
of 15-60 amino acids in which one or two
paragine residues. If a glycosylation sequence
strongly basic residues (Lys, Arg) near the N-
(-Asn-X-Ser(Thr)-, where X can be any
terminus are followed by a strongly hydro-
amino acid) appears in the growing peptide
phobic sequence of
10-15
residues
(see
chain, then a transglycosylase in the ER mem-
p. 228).
brane [1] transfers a previously produced core
As soon as the signal peptide (red) appears
oligosaccharide consisting of
14
hexose
on the surface of the ribosome (2), an RNA-
residues en-bloc from the carrier molecule
containing signal recognition particle
(SRP,
dolichol diphosphate to the peptide.
green) binds to the sequence and initially
Dolichol is a long-chain isoprenoid (see
interrupts translation (3). The SRP then binds
p. 52) consisting of
10-20 isoprene units,
to an SRP receptor in the rER membrane, and
which is embedded in the ER membrane. A
in this way attaches the ribosome to the ER
hydroxyl group at the end of the molecule is
(4). After this, the SRP dissociates from the
bound to diphosphate, on which the nuclear
signal peptide and from the SRP receptor
oligosaccharide is built up in an extended
and is available again for step 3. This ender-
reaction sequence (not shown here in detail).
gonic process is driven by GTP hydrolysis (5).
The core structure consists of two residues of
Translation now resumes. The remainder of
N-acetylglucosamine
(GlcNAc), a branched
the protein, still unfolded, is gradually intro-
group of nine mannose residues (Man) and
duced into a channel (the translocon) in the
three terminal glucose resides (Glc).
lumen of the rER (6), where a signal peptidase
As the proprotein passes through the ER,
located in the inner ER membrane cleaves the
glycosidases [2] remove the glucose residues
signal peptide while translation is still taking
completely and the mannoses partially
place (7). This converts the preprotein into a
(“trimming”), thereby producing the man-
proprotein, from which the mature protein
nose-rich type of oligosaccharide residues.
finally arises after additional post-transla-
Subsequently, various glycosyltransferases [3]
tional modifications (8) in the ER and in the
transfer additional monosaccharides
(e. g.,
Golgi apparatus.
GlcNAc, galactose, fucose, and N-acetylneura-
If the growing polypeptide contains a stop-
minic acid; see p. 38) to the mannose-rich
transfer signal (see p. 228), then this hydro-
intermediate and thereby produce the com-
phobic section of the chain remains stuck in
plex type of oligosaccharide. The structure of
the membrane outside the translocon, and an
the final oligosaccharide depends on the type
integral membrane protein arises. In the
and activity of the glycosyltransferases pre-
course of translation, an additional signal se-
sent in the ER of the cell concerned, and is
quence can re-start the transfer of the chain
therefore genetically determined (although
through the translocon. Several repetitions of
indirectly).
this process produce integral membrane pro-
teins with several transmembrane helices
(see p. 214).
B. Protein glycosylation
Most extracellular proteins contain covalently
bound oligosaccharide residues. For example,
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Endoplasmic reticulum and Golgi apparatus
231
A. Protein synthesis in the rough endoplasmic reticulum
1
2
3
4
mRNA
SRP
Signal
peptide
Free
ribosome
ER membrane
ER-lumen
SRP receptor
Translocon
5
6
7
8
GDP+Pi
GTP
Post-transl.
Signal peptidase
modification
B. Protein glycosylation
Protein
1
glycosyltransferase
2.4.1.119
Ribosome
2
O-glycosidases 3.2.1.n
Dolichol (Dol)
Translocon
Glycosyltransferases
3
2.4.1.n
P
Glucose
1
P
N-acetylglucosamine
Asn
Mannose
Asn
Galactose
N-acetylneuraminic
acid
H2O
UDP
GDP
Asn
2
Dol
Precursors
Dol
Core structure
Asn
3
P
Mannose-rich
P
type
Dolichol diphosphate (9-23 isoprene units)
Complex
type
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
232
Organelles
tible to proteinases. To protect partly folded
Protein maturation
proteins, there are auxiliary proteins called
After translation, proteins destined for the
chaperones because they guard immature
secretory pathway (see p. 228) first have to
proteins against damaging contacts. Chaper-
fold into their native conformation within the
ones are formed increasingly during tempera-
rER (see p. 230). During this process they are
ture stress and are therefore also known as
supported by various auxiliary proteins.
heat-shock proteins (hsp). Several classes of
hsp are distinguished. Chaperones of the
hsp70 type (Dna K in bacteria) are common,
A. Protein folding in the rER
as are type hsp60 chaperonins (GroEL/ES in
To prevent incorrect folding of the growing
bacteria). Class hsp90 chaperones have spe-
protein during protein biosynthesis, chaper-
cial tasks (see p. 378).
ones (see B) in the lumen of the rER bind to
While small proteins can often reach their
the peptide chain and stabilize it until trans-
native conformation without any help (1),
lation has been completed. Binding protein
larger molecules require hsp70 proteins for
(BiP) is an important chaperone in the ER.
protection against aggregation which bind as
Many secretory proteins—e. g., pancreatic
monomers and can dissociate again, depend-
ribonuclease (RNAse; see p. 74)—contain sev-
ent on ATP (3). By contrast, type hsp60 chap-
eral disulfide bonds that are only formed ox-
eronins form large, barrel-shaped complexes
idatively from SH groups after translation.
with 14 subunits in which proteins can fold
The eight cysteine residues of the RNAse can
independently while shielded from their en-
in principle form 105 different pairings, but
vironment (4). The function of hsp60 has been
only the combination of the four disulfide
investigated in detail in the bacterial
bonds shown on p. 75 provides active en-
chaperonin GroEL (right). The barrel has two
zyme. Incorrect pairings can block further
chambers, which are closed with a lid (GroES)
folding or lead to unstable or insoluble con-
during folding of the guest protein. Driven by
formations. The enzyme protein disulfide iso-
ATP hydrolysis, the chambers open and close
merase [1] accelerates the equilibration be-
alternately—i. e., the release of the fully folded
tween paired and unpaired cysteine residues,
protein from one chamber is coupled to the
so that incorrect pairs can be quickly split
uptake of an unfolded peptide in the second
before the protein finds its final conformation.
chamber.
Most peptide bonds in proteins take on the
trans conformation (see p. 66). Only bonds
C. Protein import in mitochondria
with proline residues
(-X-Pro-) can be
present in both cis and trans forms.
Class hsp70 chaperones are also needed for
In the protein’s native conformation, every
translocation of nuclear-coded proteins from
X-Pro bond has to have the correct conforma-
the cytoplasm into the mitochondria (see
tion (cis or trans). As the uncatalyzed transi-
p. 228). As two membranes have to be
tion between the two forms is very slow,
crossed to reach the matrix, there are two
there is a proline cis-trans isomerase [2] in
translocator complexes: TOM
(“transport
the ER that accelerates the conversion.
outer membrane”) and TIM (“transport inner
membrane”). For transport, proteins are un-
folded in the cytoplasm and protected by
B. Chaperones and chaperonins
hsp70. TOM recognizes the positively charged
Most proteins fold spontaneously into their
signal sequence at the protein’s N terminus
native conformation, even in the test tube.
(see p. 228) and with the help of the mem-
In the cell, where there are very high concen-
brane potential threads the chains through
trations of proteins (around 350 g L-1), this
the central pores of the two complexes. Inside
is more dif cult. In the unfolded state, the
TIM, further hsp70 molecules bind and pull
apolar regions of the peptide chain (yellow)
the chain completely into the matrix. As with
tend to aggregate—due to the hydrophobic
import into the ER, the signal peptide is pro-
effect
(see p. 28)—with other proteins or
teolytically removed by a signal peptidase
with each other to form insoluble products
during translocation.
(2). In addition, unfolded proteins are suscep-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Endoplasmic reticulum and Golgi apparatus
233
A. Protein folding in the rER
1
Protein disulfide isomerase 5.3.4.1
Cyto-
2
Peptidyl proline cis-trans-isomerase 5.2.1.8
plasm
gER
SH
1
S
2
S
S S
S S
S S
S
S
SH
S
SH
S
BiP (Chaperonin)
Pro (cis)
Pro (trans)
B. Chaperones and chaperonins
GroES
1
Folded
Small
protein
protein
1
7 ATP
7 ADP
GroEL
Aggregation
hsp 60
4
2
2
3
(Chape-
7
P
7
P
ronin)
4
7 ADP
7 ATP
hsp 70
(chaperone)
Insoluble
precipitate
or degradation
3
Folded protein
Folded protein
C. Protein import in mitochondria
Signal peptide
Nuclear-coded
Cytoplasm
for mitochondrial
mitochondrial protein
hsp 70
import
TOM
Outer
mitochondrial
complex
membrane
TIM
Inner
complex
mitochondrial
membrane
Mitochondrial
matrix
1.
Peptidase
2.
3.
hsp 70
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
234
Organelles
C. Synthesis and transport of lysosomal
Lysosomes
proteins
A. Structure and contents
Primary lysosomes arise in the region of the
Golgi apparatus. Lysosomal proteins are syn-
Animal lysosomes are organelles with a diam-
thesized in the rER and are glycosylated there
eter of 0.2-2.0 µm with various shapes that
as usual (1; see p. 228). The next steps are
are surrounded by a single membrane. There
specific for lysosomal proteins (right part of
are usually several hundred lysosomes per
the illustration). In a two-step reaction, ter-
cell. ATP-driven V-type proton pumps are ac-
minal mannose residues (Man) are phos-
tive in their membranes (see p. 220). As these
phorylated at the C-6 position of the man-
accumulate H+ in the lysosomes, the content
nose. First, N-acetylglucosamine 1-phosphate
of lysosomes with pH values of 4.5-5 is much
is transferred to the OH group at C-6 in a
more acidic than the cytoplasm (pH 7-7.3).
terminal mannose residue, and N-acetylglu-
The lysosomes are the cell’s “stomach,”
cosamine is then cleaved again. Lysosomal
serving to break down various cell compo-
proteins therefore carry a terminal mannose
nents. For this purpose, they contain some
6-phosphate (Man6-P; 2).
40 different types of hydrolases, which are
The membranes of the Golgi apparatus
capable of breaking down every type of mac-
contain receptor molecules that bind Man
romolecule. The marker enzyme of lysosomes
6-P. They recognize lysosomal proproteins
is acid phosphatase. The pH optimum of lyso-
by this residue and bind them (3). With the
somal enzymes is adjusted to the acid pH value
help of clathrin, the receptors are concen-
and is also in the range of pH 5. At neutral pH,
trated locally. This allows the appropriate
as in the cytoplasm, lysosomal enzymes only
membrane sections to be pinched off and
have low levels of activity. This appears to be a
transported to the endolysosomes with the
mechanism for protecting the cells from di-
help of transport vesicles (4), from which pri-
gesting themselves in case lysosomal enzymes
mary lysosomes arise through maturation (5).
enter the cytoplasm at any time. In plants and
Finally, the phosphate groups are removed
fungi, the cell vacuoles (see p. 43) have the
from Man 6-P (6).
function of lysosomes.
The Man 6-P receptors are reused. The fall
in the pH value in the endolysosomes releases
B. Functions
the receptors from the bound proteins (7)
which are then transported back to the Golgi
Lysosomes serve for enzymatic degradation of
apparatus with the help of transport vesicles.
macromolecules and cell organelles, which
are supplied in various ways. The example
shows the degradation of an overaged mito-
Further information
chondrion by autophagy. To accomplish this,
Many hereditary diseases are due to genetic
the lysosome encloses the organelle (1). Dur-
defects in lysosomal enzymes. The metabo-
ing this process, the primary lysosome con-
lism of glycogen ( glycogenoses), lipids (
verts into a secondary lysosome, in which the
lipidoses), and proteoglycans ( mucopoly-
hydrolytic degradation takes place (2). Finally,
saccharidoses) is particularly affected. As the
residual bodies contain the indigestible resi-
lysosomal enzymes are indispensable for the
dues of the lysosomal degradation process.
intracellular breakdown of macromolecules,
Lysosomes are also responsible for the degra-
unmetabolized macromolecules or degrada-
dation of macromolecules and particles taken
tion products accumulate in the lysosomes
up by cells via endocytosis and phagocyto-
in these diseases and lead to irreversible cell
sis—e. g., lipoproteins, proteohormones, and
damage over time. In the longer term, en-
bacteria (heterophagy). In the process, lyso-
largement takes place, and in severe cases
somes fuse with the endosomes (3) in which
there may be failure of the organ affected—
the endocytosed substances are supplied.
e. g., the liver.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lysosomes
235
A. Structure and contents
B. Functions
Overaged
Enzymatic
Approx. 40 different hydrolases
mitochondrion
degradation
with acidic pH optima
Secondary
lysosome
Lysosome
1
pH = 4.5 - 5.0
Acid
2
hydrolases
Nucleases
Proteases
Glycosidases
Primary
3
Lipases
lysosome
Phosphatases
Residual
Sulfatases
body
Phospholipases
Endosome
Secondary
lysosome
H
Endocytotic
vesicle
Macro-
Cytoplasm
H -transporting
molecules,
pH = 7.0 - 7.3
ATPase
particles
Cell
3.6.1.35
Endocytosis
A
A
H
or phagocytosis
P P P
P P P
C. Synthesis and transport of lysosomal proteins
Terminal mannose
1
rER
Protein
CH2OH
biosynthesis
H
O
N-Glycosy-
H
OH
HO
O R
lation
HO
H
H
cis-Golgi
Glyco-
U
2
P
P
-GlcNAc
apparatus
protein
Phosphory-
1
R
lation
U
P
CH2OH
trans-Golgi
No Man-6-
P
H
O
O
network
H
3
no transport
Binding to
OH H
O P
O
Receptor
Man-6- P
HO
O
receptor
H
NH
Sorting
CH2
Sealing
O
C
H
O
Vesicular
CH3
O
R
transport
OH
H HO
HO
4
H
H
7
H2O
Endo-
lysosome
2
Acidification
8
Dissociation
GlcNAc
of the receptor-
protein
5
P
O
CH2
complex
O
Low pH triggers
OHHO
conformational
change
Primary
HO
O
R
lysosome
Phosphate
1
GlcNAc-Phosphotransferase 2.7.8.17
6
cleavage
2
GlcNAc-Phosphoglycosidase 3.1.4.45
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
236
Molecular genetics
mine for uracil—it is identical to that of the
Molecular genetics: overview
sense strand. In this way, the DNA triplet TTC
Nucleic acids (DNA and various RNAs) are of
gives rise in hnRNA to the RNA codon UUC.
central importance in the storage, transmis-
RNA maturation. In eukaryotes, the hnRNA
sion, and expression of genetic information.
initially formed is modified several times be-
The decisive factor involved is their ability to
fore it can leave the nucleus as messenger
enter into specific base pairings with each
RNA (mRNA, 4). During RNA maturation,
other (see p. 84). The individual processes in-
superfluous
(“intervening”) sequences
(in-
volved, which are summed up in an overview
trons) are removed from the molecule, and
here, are discussed in more detail on the fol-
both ends of the transcript are protected by
lowing pages.
the addition of further nucleotides
(see
p. 246).
Translation. Mature mRNA enters the cyto-
A. Expression and transmission of genetic
plasm, where it binds to ribosomes, which
information
convert the RNA information into a peptide
Storage. The genetic information of all cells is
sequence. The ribosomes (see p. 250) consist
stored in the base sequence of their DNA (RNA
of more than 100 proteins and several RNA
only occurs as a genetic material in viruses;
molecules (rRNA; see p. 82). rRNA plays a role
see p. 404). Functional sections of DNA that
as a ribosomal structural element and is also
code for inheritable structures or functions
involved in the binding of mRNA to the ribo-
are referred to as genes. The 30 000-40 000
some and the formation of the peptide bond.
human genes represent only a few percent of
The actual information transfer is based on
the genome, which consists of approximately
the interaction between the mRNA codons
5
109 base pairs (bp). Most genes code for
and another type of RNA, transfer RNA
proteins—i. e., they contain the information
(tRNA; see p. 82). tRNAs, of which there are
for the sequence of amino acid residues of a
numerous types, always provide the correct
protein (its sequence). Every amino acid res-
amino acid to the ribosome according to the
idue is represented in DNA by a code word (a
sequence information in the mRNA. tRNAs are
codon) consisting of a sequence of three base
loaded with an amino acid residue at the 3
pairs (a triplet). At the level of DNA, codons
end. Approximately in the middle, they
are defined as sequences of the sense strand
present the triplet that is complementary to
read in the 5
3 direction (see p. 84). A DNA
each mRNA codon, known as the anticodon
codon for the amino acid phenylalanine, for
(GAA in the example shown). If the codon UUC
example, is thus TTC (2).
appears on the mRNA, the anticodon binds a
Replication. During cell division, all of the
molecule of Phe-t-RNAPhe to the mRNA (5)
genetic information has to be passed on to the
and thus brings the phenylalanine residue at
daughter cells. To achieve this, the whole of
the other end of the molecule into a position
the DNA is copied during the S phase of the
in which it can take over the growing poly-
cell cycle (see p. 394). In this process, each
peptide chain from the neighboring tRNA (6).
strand serves as a matrix for the synthesis
Amino acid activation. Before binding to
of a complementary second strand (1; see
the ribosomes, tRNAs are loaded with the
p. 240).
correct amino acids by specific ligases (7;
Transcription. For expression of a gene—i. e.,
see p. 248). It is the amino acid tRNA ligases
synthesis of the coded protein—the DNA se-
that carry out the transfer (translation) of the
quence information has to be converted into a
genetic information from the nucleic acid
protein sequence. As DNA itself is not in-
level to the protein level.
volved in protein synthesis, the information
is transferred from the nucleus to the site of
synthesis in the cytoplasm. To achieve this,
the template strand in the relevant part of
the gene is transcribed into an RNA (hnRNA).
The sequence of this RNA is thus complemen-
tary to that of the template strand (3), but—
with the exception of the exchange of thy-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Overview
237
A. Expression and transmission of genetic information
5'
3'
N
N
T
T
C
N
N
N
N
A A
G
N
N
3'
5'
5'
3'
N
N
T
T
C
N
N
N
N
A A
G
N
N
3'
5'
1
Replication
Sense strand
DNA
5'
3'
N
N
T
T
C
N
N
N
N
A A
G
N
N
3'
5'
2
Template strand
Transcription
hnRNA
5'
3'
N
N
U
U
C
N
N
N
N
A A
G
N
N
3'
5'
3
Template strand
RNA maturation
DNA
mRNA
Poly(A) tail
5'
3'
N
N
U U C
N
N
4
Cap
7
Phe
tRNA
Translation
mRNA
Amino acid
activation
5'
3'
N
N
U
U
C
N
N
N
N
A A
G
N
N
A
P
5'
Ribosome
6
Phe-tRNA
Protein
5
Phe
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
238
Molecular genetics
The non-histone proteins are very hetero-
Genome
geneous. This group includes structural pro-
teins of the nucleus, as well as many enzymes
A. Chromatin
and transcription factors (see p. 118), which
In the nuclei of eukaryotes (see p. 196), DNA is
selectively bind to specific segments of DNA
closely associated with proteins and RNA.
and regulate gene expression and other pro-
These nucleoprotein complexes, with a DNA
cesses.
proportion of approximately one-third, are
known as chromatin. It is only during cell
B. Histones
division
(see p. 394) that chromatin con-
denses into chromosomes that are visible
The histones are remarkable in several ways.
under light microscopy. During interphase,
With their high proportions of lysine and ar-
most of the chromatin is loose, and in these
ginine (blue shading), they are strongly basic,
conditions a morphological distinction can be
as mentioned above. In addition, their amino
made between tightly packed heterochroma-
acid sequence has hardly changed at all in the
tin and the less dense euchromatin. Euchro-
course of evolution. This becomes clear when
matin is the site of active transcription.
one compares the histone sequences in mam-
The proteins contained in chromatin are
mals, plants, and fungi (yeasts are single-
classified as either histone or non-histone
celled fungi; see p. 148). For example, the
proteins. Histones (B) are small, strongly basic
H4 histones in humans and wheat differ
proteins that are directly associated with
only in a single amino acid residue, and there
DNA. They contribute to the structural organ-
are only a few changes between humans and
ization of chromatin, and their basic amino
yeast. In addition, all of these changes are
acids also neutralize the negatively charged
“conservative”—i. e., the size and polarity
phosphate groups, allowing the dense pack-
barely differ. It can be concluded from this
ing of DNA in the nucleus. This makes it pos-
that the histones were already “optimized”
sible for the 46 DNA molecules of the diploid
when the last common predecessor of ani-
human genome, with their 5
109 base pairs
mals, plants, and fungi was alive on Earth
(bp) and a total length of about 2 m, to be
(more than 700 million years ago). Although
accommodated in a nucleus with a diameter
countless mutations in histone genes have
of only 10 µm. Histones also play a central
taken place since, almost all of these evidently
role in regulating transcription (see p. 244).
led to the extinction of the organisms con-
Two histone molecules each of types H2A
cerned.
(blue), H2B (green), H3 (yellow), and H4 (red)
The histones in the octamer carry N-termi-
form an octameric complex, around which
nal mobile “tails” consisting of some 20 amino
146 bp of DNA are wound in 1.8 turns. These
acid residues that project out of the nucleo-
particles, with a diameter of 7 nm, are re-
somes and are important in the regulation of
ferred to as nucleosomes. Another histone
chromatin structure and in controlling gene
(H1) binds to DNA segments that are not di-
expression (see A2; only two of the eight tails
rectly in contact with the histone octamers
are shown in full length). For example, the
(“linker” DNA). It covers about 20 bp and sup-
condensation of chromatin into chromo-
ports the formation of spirally wound super-
somes is associated with phosphorylation (P)
structures with diameters of 30 nm, known as
of the histones, while the transcription of
solenoids. When chromatin condenses into
genes is initiated by acetylation (A) of lysine
chromosomes, the solenoids form loops about
residues in the N-terminal region (see p. 244).
200 nm long, which already contain about
80 000 bp. The loops are bound to a protein
framework (the nuclear scaffolding), which in
turn organizes some 20 loops to form mini-
bands. A large number of stacked minibands
finally produces a chromosome. In the chro-
mosome, the DNA is so densely packed that
the smallest human chromosome already
contains more than 50 million bp.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Replication
239
A. Chromatin
Solenoid
Nuclear scaffold
200 nm
Loops
30 nm
DNA double helix
Nucleosome
10 nm
Chromosome
2 nm
700 nm
Histone
1. Organization of chromatin
molecules
DNA
N-terminal
N-terminal
tail (H2B)
tail (H3)
H2A
H2B
H3
Histone-Octamer
H4
(H2A · H2B)2(H3 · H4)2
2. Nucleosome
B. Histone
A
Acetylation
P
Phosphorylation
M
Methylation
A
P
A
A
A
A
M
10
Animals
S
G R G
K
G
G
K
G
L
G
K
G G A
K R H
R
K
V L R D N I
Plants
S
G R G
K
G
G
K
G
L
G
K
G G A
K R H
R
K
V L R D N I
Yeast
S
G R G
K
G
G
K
G
L
G
K
G G A
K R H
R
K
I
L R D N I
30
40
50
Q G I
T
K
P
A
I
R
R
L
A
R R
G G V
K
R
I
S G L I Y E
Animals
Plants
Q G I
T
K
P
A
I
R
R
L
A
R R
G G V
K
R
I
S G L I Y E
Yeast
Q G I
T
K
P
A
I
R
R
L
A
R R
G G V
K
R
I
S G L I Y E
60
70
E T
R
G
V
L
K
V
F
L
E
N
V I
R
D A V
T
Y T E H A
K R
Animals
Plants
E T
R
G
V
L
K
I
F
L
E
N
V I
R
D A V
T
Y T E H A
R
R
Yeast
E
V
R
A
V
L
K
S
F
L
E
S
V I
R
D
S
V
T
Y T E H A
K R
80
90
100
K
T V T
A
M
D
V
V
Y
A
L
K R
Q G
R
T
L
Y G F G G
Animals
Plants
K
T V T
A
M
D
V
V
Y
A
L
K R
Q G
R
T
L
Y G F G G
Yeast
K
T V T
S
L
D
V
V
Y
A
L
K R
Q G
R
T
L
Y G F G G
Amino acid sequence of histone H4
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
240
Molecular genetics
the processes taking place in this type of fork,
Replication
and only the most important are shown here.
For genetic information to be passed on dur-
The two strands of the initial DNA (1) are
ing cell division, a complete copy of the ge-
shown in blue and violet, while the newly
nome has to be produced before each mitosis.
formed strands are pink and orange.
This process is known as DNA replication.
Each fork (2) contains two molecules of
DNA polymerase III and a number of helper
proteins. The latter include DNA topoisomer-
A. Mechanism of DNA polymerases
ases and single-strand-binding proteins.
Replication is catalyzed by DNA-dependent
Topoisomerases are enzymes that unwind
DNA polymerases. These enzymes require a
the superhelical DNA double strand (gyrase,
single strand of DNA, known as the tem-
topoisomerase II) and then separate it into the
plate. Beginning at a short starting sequence
two individual strands (helicase, topoisomer-
of RNA (the primer), they synthesize a second
ase I). Since the template strand is always read
complementary strand on the basis of this
from 3
to 5
(see above), only one of the
template, and thus create a complete DNA
strands (known as the leading strand; violet/
double helix again. The substrates of the
pink) can undergo continuous replication. For
DNA polymerases are the four deoxynucleo-
the lagging strand (light blue), the reading
side triphosphates dATP, dGTP, dCTP, and
direction is the opposite of the direction of
dTTP. In each step, base pairing first binds
movement of the fork. In this matrix, the
the nucleotide that is complementary to the
new strand is first synthesized in individual
current base in the template strand. The
pieces, which are known as Okazaki frag-
α-phosphate residue of the newly bound nu-
ments after their discoverer (green/orange).
cleoside triphosphate is then subjected to nu-
Each fragment starts with a short RNA pri-
cleophilic attack by the 3 -OH group of the
mer (green), which is necessary for the func-
nucleotide incorporated immediately previ-
tioning of the DNA polymerase and is synthe-
ously. This is followed by the elimination of
sized by a special RNA polymerase (“primase,”
diphosphate and the formation of a new
not shown). The primer is then extended by
phosphoric acid diester bond. These steps
DNA polymerase III (orange). After 1000-2000
are repeated again for each nucleotide. The
nucleotides have been included, synthesis of
mechanism described means that the matrix
the fragment is interrupted and a new one is
can only be read in the 3
5
direction. In
begun, starting with another RNA primer that
other words, the newly synthesized strand
has been synthesized in the interim. The in-
always grows in the 5
3
direction. The
dividual Okazaki fragments are initially not
same mechanism is also used in transcription
bound to one another and still have RNA at
by DNA-dependent RNA polymerases
(see
the 5 end (3). At some distance from the fork,
p. 242). Most DNA and RNA polymerases con-
DNA polymerase I therefore starts to remove
sist of more than 10 subunits, the role of
the RNA primer and replace it with DNA com-
which is still unclear to some extent.
ponents. Finally, the gaps still remaining are
closed by a DNA ligase. In DNA double helices
formed in this way, only one of the strands has
B. Replication in E. coli
been newly synthesized—i. e., replication is
Although replication in prokaryotes is now
semiconservative.
well understood, many details in eukaryotes
In bacteria, some 1000 nucleotides are re-
are still unclear. However, it is certain that the
plicated per second. In eukaryotes, replication
process is in principle similar. A simplified
takes place more slowly (about 50 nucleotides
scheme of replication in the bacterium
s-1) and the genome is larger. Thousands of
Escherichia coli is shown here.
replication forks are therefore active simulta-
In bacteria, replication starts at a specific
neously in eukaryotes.
point in the circular DNA—the origin of repli-
cation—and proceeds in both directions. This
results in two diverging replication forks, in
which the two strands are replicated simulta-
neously. Numerous proteins are involved in
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Replication
241
A. Mechanism of DNA polymerases
3'
5'
A
C
T
C
Template
G G
O
Template
strand
strand
H H
T
G
5'
Newly synthe-
A
H
H
sized strand
HO
Cyt
HO
O H
P
P
P
5'
Gua
O
P
O H2C
O
P
P
1'
O
H H
3'
5'
H
H
A
C
T
C
G G
3'
HO H
T
G
A
5'
G
Thy
DNA poly-
O
O
O
HO
merase
HO
2.7.7.7
5'
Ade
P
P
P
O
P
O
P
O
P
O H2C
O
O
O
O
H H
P
P
3'
H
H
HO H
B. Replication in E. coli
Origin of
Replication fork
Old DNA strands
replication
New DNA strands
RNA primer
3'
1.
5'
DNA polymerase III
Auxiliary proteins
2.7.7.7
DNA helicase 5.99.1.2
2.
3'
5'
DNA gyrase 5.99.1.3
5'
Okazaki
3'
5'
fragments
OF2
3'
Single
strand
Movement of the
OF1
binding
replication fork
5'
proteins
3'
5'
3.
3'
5'
DNA polymerase I
2.7.7.7
OF3
DNA ligase
OF1
6.5.1.1
OF2
5'
3'
3'
4.
5'
Replication is semiconservative
5'
3'
OF3
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
242
Molecular genetics
3 end of the promoter (“transcription start”)
Transcription
and continues until the polyadenylation se-
For the genetic information stored in DNA to
quence (see below) is reached. The primary
become effective, it has to be rewritten
transcript (hnRNA) still has a length of about
(transcribed) into RNA. DNA only serves as a
6.2 kbp. During RNA maturation, the non-
template and is not altered in any way by the
coding sequences corresponding to the in-
transcription process. Transcribable segments
trons are removed, and the two ends of the
of DNA that code for a defined product are
hnRNA are modified. The translatable mRNA
called genes. It is estimated that the mamma-
still has half the length of the hnRNA and is
lian genome contains 30 000-40 000 genes,
modified at both ends (see p. 246).
which together account for less than 5 % of the
In many eukaryotic genes, the proportion
DNA.
of introns is even higher. For example, the
gene for dihydrofolate reductase (see p. 402)
is over 30 kbp long. The information is dis-
A. Transcription and maturation of RNA:
tributed over six exons, which together have a
overview
length of only about 6 kbp.
Transcription is catalyzed by DNA-dependent
RNA polymerases. These act in a similar way to
C. Transcription process
DNA polymerases (see p. 240), except that
they incorporate ribonucleotides instead of
As mentioned above, RNA polymerase II
deoxyribonucleotides into the newly synthe-
(green) binds to the 3
end of the promoter
sized strand; also, they do not require a pri-
region. A sequence that is important for this
mer. Eukaryotic cells contain at least three
binding is known as the TATA box—a short A-
different types of RNA polymerase. RNA poly-
and T-rich sequence that varies slightly from
merase I synthesizes an RNA with a sedimen-
gene to gene. A typical base sequence (“con-
tation coef cient (see p. 200) of 45 S, which
sensus sequence”) is ...TATAAA... Numerous
serves as precursor for three ribosomal RNAs.
proteins known as basal transcription factors
The products of RNA polymerase II are
are necessary for the interaction of the poly-
hnRNAs, from which mRNAs later develop,
merase with this region. Additional factors
as well as precursors for snRNAs. Finally,
can promote or inhibit the process (transcrip-
RNA polymerase III transcribes genes that
tional control; see p. 244). Together with the
code for tRNAs, 5S rRNA, and certain snRNAs.
polymerase, they form the basal transcription
These precursors give rise to functional RNA
complex.
molecules by a process called RNA maturation
At the end of initiation (2), the polymerase
(see p. 246). Polymerases II and III are inhib-
is repeatedly phosphorylated, frees itself from
ited by D-amanitin, a toxin in the Amanita
the basal complex, and starts moving along
phalloides mushroom.
the DNA in the 3 direction. The enzyme sep-
arates a short stretch of the DNA double helix
into two single strands. The complementary
B. Organization of the PEP-CK gene
nucleoside triphosphates are bound by base
The way in which a typical eukaryotic gene is
pairing in the template strand and are linked
organized is illustrated here using a gene that
step by step to the hnRNA as it grows in the
codes for a key enzyme in gluconeogenesis
5
3
direction (3). Shortly after the begin-
(see p. 154)—the phosphoenolpyruvate car-
ning of elongation, the 5 end of the transcript
boxykinase (PEP-CK).
is protected by a “cap” (see p. 246). Once the
In the rat, the PEP-CK gene is nearly 7 kbp
polyadenylation sequence has been reached
(kilobase pairs) long. Only 1863 bp, distrib-
(typical sequence: .. .AATAA...), the transcript
uted over 10 coding segments (exons, dark
is released (4). Shortly after this, the RNA
blue) carry the information for the protein’s
polymerase stops transcribing and dissociates
621 amino acids. The remainder is allotted to
from the DNA.
the promoter (pink) and intervening sequen-
ces (introns, light blue). The gene’s promoter
region (approximately 1 kbp) serves for reg-
ulation (see p. 188). Transcription starts at the
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Transcription
243
A. Transcription and maturation of RNA: overview
ATP, GTP, CTP, UTP
Transcription
Maturation
28S rRNA
Polymerase I
45S rRNA
18S rRNA
5.8S rRNA
hnRNA
mRNA
Polymerase II
snRNA-
snRNA
precursors
α-Amanitin
5S rRNA
Polymerase III
Precursors
tRNA
snRNA
DNA
DNA-directed RNA
polymerase 2.7.7.6
B. Organization of the PEP-CK gene
Translation
Transcription
Translation
Exon
Intron
end (TAA)
start
start (ATG)
Transcription
100 Bases
end
5'
3'
DNA
Promoter
region
Transcription
5'
3'
hn RNA
RNA maturation
5'
3'
Poly-A sequence
mRNA
Cap
AUG
UAA
C. Process of transcription
TATA-Box
Transcription start
Polyadenylation
...TATAAA..
..AATAAA..
sequence
3'
5'
1.
Basal transcription complex
3'
5'
2.
RNA Polymerase II
Template strand
P
Elongation
P
P
3'
5'
3.
Cap
P
P
P
Termination
3'
5'
4.
hnRNA
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
244
Molecular genetics
ent proteins are involved in the basal com-
Transcriptional control
plex. This alone, however, is still not suf cient
Although all cells contain the complete ge-
for transcription to start. In addition, positive
nome, they only use a fraction of the informa-
signals have to be emitted by more distant
tion in it. The genes known as “housekeeping
trans-active factors, integrated by the coacti-
genes,” which code for structural molecules
vator/mediator complex, and passed on to the
and enzymes of intermediate metabolism, are
basal complex (see B).
the only ones that undergo constant tran-
The actual signal for starting elongation
scription. The majority of genes are only ac-
consists of the multiple phosphorylation of a
tive in certain cell types, in specific metabolic
domain in the C-terminal region of the poly-
conditions, or during differentiation. Which
merase. In phosphorylated form, it releases
genes are transcribed and which are not is
itself from the basal complex along with a
regulated by transcriptional control (see also
few TFs and starts to synthesize hnRNA.
p. 118). This involves control elements (cis-
active elements) in the gene’s promoter re-
B. Regulation of PEP-CK transcription
gion and gene-specific regulatory proteins
(transcription factors, trans -active factors),
Phosphoenolpyruvate carboxykinase (PEP-CK),
which bind to the control elements and
a key enzyme in gluconeogenesis, is regulated
thereby activate or inhibit transcription.
by several hormones, all of which affect the
transcription of the PEP-CK gene. Cortisol,
glucagon, and thyroxin induce PEP-CK, while
A. Initiation of transcription
insulin inhibits its induction (see p. 158).
In the higher organisms, DNA is blocked by
More than ten control elements (dark red),
histones (see p. 238) and is therefore not ca-
distributed over approximately 1 kbp, have so
pable of being transcribed without special
far been identified in the promoter of the PEP-
positive regulation. In eukaryotes, it is there-
CK gene (top). These include response ele-
fore histones that play the role of repressors
ments for the glucocorticoid receptor (GRE;
(see p. 118). For transcription to be set in mo-
see p. 378), for the thyroxin receptor (TRE),
tion at all, the chromatin first has to be re-
and for the steroid-like retinoic acid (AF-1).
structured.
Additional control elements (CRE, cAMP-re-
In the resting state, the lysine residues in
sponsive element) bind the transcription fac-
the N-terminal “tail” of the histones
(see
tor C/EBP, which is activated by cAMP-de-
p. 238) are not acetylated. In this state, which
pendent protein kinase A through phosphor-
can be produced by histone deacetylases [1],
ylation. This is the way in which glucagon,
the nucleosomes are stable. It is only the in-
which raises the cAMP level (see p. 158),
teraction of activator and regulator proteins
works. Control element P1 binds the hor-
with their control elements that allows the
mone-independent factor NF-1 (nuclear fac-
binding of coactivator complexes that have
tor-1). All proteins that bind to the control
histone acetylase activity [2]. They acetylate
elements mentioned above are in contact
the histone tails and thereby loosen the nu-
with a coactivator/mediator complex (CBP/
cleosome structure suf ciently for the basal
p300), which integrates their input like a
transcription complex to form.
computer and transmits the result in the
This consists of DNA-dependent RNA poly-
form of stronger or weaker signals to the basal
merase II and basal transcription factors
transcription complex. Inhibition of PEP-CK
(TFIIX, X = A - H). First, the basal factor TFIID
transcription by insulin is mediated by an
binds to the promoter. TFIID, a large complex
insulin-responsive element (IRE) in the vicin-
of numerous proteins, contains TATA box-
ity of the GRE. Binding of an as yet unknown
binding protein
(TBP) and so-called TAFs
factor takes place here, inhibiting the binding
(TBP-associated factors). The polymerase is
of the glucocorticoid receptor to the GRE.
attached to this core with the help of TFIIB.
Before transcription starts, additional TFs
have to bind, including TFIIH, which has heli-
case activity and separates the two strands of
DNA during elongation. In all, some 35 differ-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Transcriptional control
245
A. Initiation of transcription
Enhancer
Histone deacetylase
Activator
Regulator Coactivator complex
(HRE)
protein
protein
(histone acetylase)
1
2
TATA-Box
TATA box
Histone “tails”
Nucleosome Upstream promoter element
Coactivator complex
1
Histone deacetylase n.n.n.n
Mediator complex
2
TAFs
2
Histone acetyltransferase 2.3.1.48
RNA polymerase II
3
RNA polymerase 2.7.7.6
4
DNA topoisomerase I 5.99.1.2
D F
3
HRE - hormone response element
A
4
TAFs - TBP-associated factors
TBP BE
H
TBP - TATA box-binding protein
A-H - basal transcription factors
Basal transcription complex
B. Regulation of PEP-CK transcription
Control
AF-1 IRE GRE
TRE
P4
CRE
P1 CRE
TATA-Box
element
Transcription
RAR
? GR
TR
AP-1
C/EBP HNF-1
NF-1 C/EBP
TBP
Tran-
factor
(Fos/Jun)
CREB
scription
start
Receptors for retinoate
cortisol thyroxine
Insulin
PI3K
Coactivator
Mediator
(CBP/p300)
GRE
Cortisol
GR
NF-1
Thyroxin
TR
TRE
C/EBP
P1
CRE
TATA box
cAMP
Basal transcription complex
Glucagon
PK-A
C/EBP
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
246
Molecular genetics
are called snRNPs (small nuclear ribonucleo-
RNA maturation
protein particles, pronounced
“snurps”).
Before the hnRNA produced by RNA polymer-
SnRNPs occur in five different forms (U1, U2,
ase II (see p. 242) can leave the nucleus in
U4, U5, and U6). They consist of numerous
order to serve as a template for protein syn-
proteins and one molecule of snRNA each
thesis in the cytoplasm, it has to undergo
(see p. 82).
several modifications first. Even during tran-
To ensure that the RNA message is not
scription, the two ends of the transcript have
destroyed, splicing has to take place in a
additional nucleotides added (A). The sections
very precise fashion. The start and end of
that correspond to the intervening gene se-
the hnRNA introns are recognized by a char-
quences in the DNA (introns) are then cut out
acteristic sequence (...AGGT.. . at the 5
end or
(splicing; see B). Other transcripts—e. g., the
...[C,U]AGG... at the 3
end). Another impor-
45 S precursor of rRNA formed by polymerase
tant structure is the so-called branching point
I (see p. 242)—are broken down into smaller
inside the intron. Its sequence is less con-
fragments by nucleases before export into the
served than the terminal splicing sites, but it
cytoplasm.
always contains one adenosine residue (A).
During splicing, the 2 -OH group of this resi-
due—supported by the spliceosome
(see
A. 5
and 3 modification of mRNA
C)—attacks the phosphoric acid diester bond
Shortly after transcription begins in eukary-
at the 5
end of the intron and cleaves it (b).
otes, the end of the growing RNA is blocked in
Simultaneously, an unusual 2
5
bond is
several reaction steps by a structure known as
formed inside the intron, which thereby takes
a “cap.” In hnRNAs, this consists of a GTP
on a lasso shape (c; see formula). In the second
residue that is methylated at N-7 of the gua-
step of the splicing process, the free 3 -OH
nine ring. The β-phosphate residue of the cap
group at the end of the 5
terminal exon at-
is linked to the free 5 -OH group of the ter-
tacks the A-G bond at the 3 end of the intron.
minal ribose via an ester bond. After the
As a result, the two exons are linked and the
“polyadenylation signal” has been reached
intron is released, still in a lasso shape (d).
(typical sequence: ...AAUAAA...; see p. 242),
a polyadenylate “tail” consisting of up to 200
C. Spliceosome
AMP nucleotides is also added at the free 3
end of the transcript. This reaction is cata-
As described above, it is residues of the
lyzed by a special polyadenylate polymerase.
hnRNA that carry out bond cleavage and
It is only at this point that the mRNA leaves
bond formation during the splicing process.
the nucleus as a complex with RNA-binding
It is therefore not a protein enzyme that acts
proteins.
as a catalyst here, but rather an RNA. Catalytic
Both the cap and the poly-A tail play a vital
RNAs of this type are called ribozymes (see
part in initiating eukaryotic translation (see
also p. 88). The task of the spliceosomes is to
p. 250). They help position the ribosome cor-
fix and orientate the reacting groups by es-
rectly on the mRNA near to the starting co-
tablishing base pairings between snRNAs and
don. The protection which the additional nu-
segments of the hnRNA. The probable situa-
cleotides provide against premature enzy-
tion before the adenosine attack at the
matic degradation appears to be of lesser im-
branching point on the 5
splicing site (see
portance.
B, Fig. b) is shown schematically on the right
side of the illustration. In this phase, the U1
snRNA fixes the 5
splicing site, U2 fixes the
B. Splicing of hnRNA
branching site, and U5 fixes the ends of the
Immediately after transcription, the hnRNA
two exons.
introns are removed and the exons are linked
to form a continuous coding sequence. This
process, known as splicing, is supported by
complicated RNA-protein complexes in the
nucleus, the so-called spliceosomes. The com-
ponents of these macromolecular machines
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
RNA maturation
247
A. 5' and 3' modification
B. Splicing of hnRNA: mechanism
of mRNA
5' splicing
Branching
3' splicing
O
CH3
site
point
site
C
N
HN
C
7
C
H
O
O
C
C
5'
a)
5'
...AG
GU
...G
AC...
...CAGG...
3'
H2N
N
N
H2C O P O
P O
O
5' exon
Intron
3' exon
O
O
H
H
H
H
O
P O
HO OH
O
5'
CH2
N
7-methyl guanosine
O
H
H
H
H
O OH
b)
5'
...A
..CA
GG...
3'
m7Gppp N...
Cap
Lasso
5' end
structure
..CA
GG...
3'
c)
5'
...AG
mRNA
d)
5'
...AG
GG...
3'
...CA
CH2
Ade
O
H
H
O
H
H
H
H
O
OH
OH O
H
H
O
O
P
O
CH2
Ade
H
O
O
O
Gua CH2
O P
O
H H
H H
O
H
H
O
O
O
OH
Gua
CH2
O P O
O
P
O
CH2
H
H
O
Ade
O
O
H
H
H H
O
Poly-A sequence
H
H
O
O OH
Ade CH2
O
P
O
O
To the 3' end
C. Spliceosome
Intron
Proteins
snRNA
snRNP
U1
U2
+
U1
U1
hnRNA
+
U2
U
U2
G
A
snRNA
G
+
U4
A
U4
+
U5
U5
U 4/6
CA
U5
5' exon
GG
+
U6
3' exon
U6
5'
3'
Spliceosome (schematic)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
248
Molecular genetics
tortions of the erythrocytes and disturbances
Amino acid activation
of O2 transport (sickle-cell anemia).
A. The genetic code
B. Amino acid activation
Most of the genetic information stored in the
genome codes for the amino acid sequences
Some 20 different amino acid tRNA ligases in
of proteins. For these proteins to be ex-
the cytoplasm each bind one type of tRNA
pressed, a text in
“nucleic acid language”
(see p. 82) with the corresponding amino
therefore has to be translated into “protein
acid. This reaction, known as amino acid acti-
language.” This is the origin of the use of the
vation, is endergonic and is therefore coupled
term translation to describe protein biosyn-
to ATP cleavage in two steps.
thesis. The dictionary used for the translation
First, the amino acid is bound by the en-
is the genetic code.
zyme and reacts there with ATP to form di-
As there are 20 proteinogenic amino acids
phosphate and an “energy-rich” mixed acid
(see p. 60), the nucleic acid language has to
anhydride (aminoacyl adenylate). In the sec-
contain at least as many words (codons).
ond step, the 3 -OH group (in other ligases it
However, there are only four letters in the
is the 2 -OH group) of the terminal ribose
nucleic acid alphabet (A, G, C, and U or T). To
residue of the tRNA takes over the amino
obtain 20 different words from these, each
acid residue from the aminoacyl adenylate.
word has to be at least three letters long
In aminoacyl tRNAs, the carboxyl group of
(with two letters, there would only be
the amino acid is therefore esterified with
42 = 16 possibilities). And in fact the codons
the ribose residue of the terminal adenosine
do consist of three sequential bases (triplets).
of the sequence .. .CCA-3 .
Figure 1 shows the standard code in “DNA
The accuracy of translation primarily de-
language” (i. e., as a sequence of triplets in the
pends on the specificity of the amino acid
sense strand of DNA, read in the 5
3 direc-
tRNA ligases, as incorrectly incorporated
tion; see p. 84), represented as a circular dia-
amino acid residues are not recognized by
gram. The scheme is read from the inside to
the ribosome later. A “proofreading mecha-
the outside. For example, the triplet CAT codes
nism” in the active center of the ligase there-
for the amino acid histidine. With the excep-
fore ensures that incorrectly incorporated
tion of the exchange of U for T, the DNA co-
amino acid residues are immediately re-
dons are identical to those of mRNA.
moved again. On average, an error only occurs
As the genetic code provides 43 = 64 co-
once every 1300 amino acid residues. This is a
dons for the 20 amino acids, there are several
surprisingly low rate considering how similar
synonymous codons for most amino acids—
some amino acids are—e. g., leucine and iso-
the code is degenerate. Three triplets do not
leucine.
code for amino acids, but instead signal the
end of translation
(stop codons). Another
C. Asp-tRNA ligase (dimer)
special codon, the start codon, marks the start
of translation. The code shown here is almost
The illustration shows the ligase responsible
universally applicable; only the mitochondria
for the activation of aspartate. Each subunit of
(see p. 210) and a few microorganisms devi-
the dimeric enzyme (protein parts shown in
ate from it slightly.
orange) binds one molecule of tRNAAsp (blue).
As an example of the way in which the
The active centers can be located by the
code is read, Fig. 2
shows small sections
bound ATP (green). They are associated with
from the normal and a mutated form of the
the 3 end of the tRNA. Another domain in the
β-globin gene (see p. 280), as well as the cor-
protein (upper left) is responsible for “recog-
responding mRNA and protein sequences. The
nition” of the tRNA anticodon.
point mutation shown, which is relatively fre-
quent, leads to replacement of a glutamate
residue in position 6 of the β-chain by valine
(GAG GTG). As a consequence, the mutated
hemoglobin tends to aggregate in the deoxy-
genated form. This leads to sickle-shaped dis-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Genetic code
249
A. The genetic code
1.
2. Normal globin
Sickle-cell globin
Gly
Phe
Leu
gene (β-chain)
gene (β-chain)
Glu
Ser
Asp
A
G
T
C A
G
5'
C C T
GAG
G AG
5'
C C T
G
T
G
G AG
TCA
Tyr
3'
G G A
C T C
C TC
3'
G G A
C
A
C
C TC
Ala
TCAGTC
G
G
T
GTC
Stop
A
A
C
A
C
Transcription
Transcription
T
G
C
A
T
Cys
G
C
mRNA
Val
A
Stop
A
C
T
G
T
G
5'
C C UG AG G AG
5'
C C UG UG G AG
G
Trp
T
CUC
C AC
G
T
Glu-
Val-
Arg
A
G
T
C
tRNA
tRNA
A
C
A
Leu
C
Ser
T
G
Glu
Val
G
A
C
T
A
C
Lys
C
C
A
A
T
G
Pro
G
T
G
T
Translation
Translation
Asn
A
C
C
A
T
G
G
T
His
A
C
T
G
A
C
Thr
Gln
Pro Glu Glu
Pro
Val
Glu
Met
Ile
Arg
Start
A
T
=
His
C
Mutation
Example
B. Amino acid activation
C. Asp-tRNA-Ligase (Dimer)
Amino acid
P
Anticodon recognition domain
H3N
O
P
5'
R C C O
P O CH2
tRNAAsp
O Ade
H
ATP
H H
3' 2'
H
H
1
P
P
HO OH
Aminoacyl adenylate
H3N
O
5'
R
C
C
P O CH2
O Ade
H
1'
H H
ATP
ATP
HO OH
3'
2'
H
H
H
3'
2'
HO OH
4'
H
H
1'
O
2
Active
P
O
CH2 Ade
AMP
center
5'
Active
O OH
center
tRNA
H3N
O
H
H
3'
2'
H H
R
C
C
O OH
4'
1'
Protein
O
H
H
3' 2'
Cyt
4'
H
H
1'
O
Amino acid-
P
O
CH2 Ade
5'
tRNA ligase
O OH
6.1.1.n
Amino
H
H
acyl
Asp
3' 2'
tRNA
4'
H H
1'
tRNA
O
Cyt
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
250
Molecular genetics
ing a single mRNA molecule simultaneously.
Translation I: initiation
The ribosome first binds near the start codon
Like amino acid activation (see p. 248), pro-
(AUG; see p. 248) at the 5 end of the mRNA
tein biosynthesis (translation) takes place in
(top). During translation, the ribosome moves
the cytoplasm. It is catalyzed by complex
in the direction of the 3 end until it reaches a
nucleoprotein particles, the ribosomes, and
stop codon (UAA, UAG, or UGA). At this point,
mainly requires GTP to cover its energy re-
the newly synthesized chain is released, and
quirements.
the ribosome dissociates again into its two
subunits.
A. Structure of the eukaryotic ribosome
C. Initiation of translation in E. coli
Ribosomes consist of two subunits of different
size, made up of ribosomal RNA (rRNA) and
Protein synthesis in prokaryotes is in princi-
nearly 80 proteins (the number of proteins
ple the same as in eukaryotes. However, as the
applies to rat liver ribosomes). It is customary
process is simpler and has been better studied
to give the sedimentation coef cients (see
in prokaryotes, the details involved in trans-
p. 200) of ribosomes and their components
lation are discussed here and on p. 252 using
instead of their masses. For example, the eu-
the example of the bacterium Escherichia coli.
karyotic ribosome has a sedimentation coef-
The first phase of translation, initiation, in-
ficient of 80 Svedberg units (80 S), while the
volves several steps. First, two proteins, ini-
sedimentation coef cients of its subunits are
tiation factors IF-1 and IF-3, bind to the 30 S
40 S and 60 S (S values are not additive).
subunit (1). Another factor, IF-2, binds as a
The smaller 40 S subunit consists of one
complex with GTP (2). This allows the subunit
molecule of 18 S rRNA and 33 protein mole-
to associate with the mRNA and makes it
cules. The larger 60 S subunit contains three
possible for a special tRNA to bind to the start
types of rRNA with sedimentation coef cients
codon (3). In prokaryotes, this starter tRNA
of 5 S, 5.8 S, and 28 S and 47 proteins. In the
carries the substituted amino acid N-
presence of mRNA, the subunits assemble to
formylmethionine (fMet). In eukaryotes, it car-
form the complete ribosome, with a mass
ries an unsubstituted methionine. Finally, the
about 650 times larger than that of a hemo-
50 S subunit binds to the above complex (4).
globin molecule.
During steps 3 and 4, the initiation factors are
The arrangement of the individual compo-
released again, and the GTP bound to IF-2 is
nents of a ribosome has now been deter-
hydrolyzed to GDP and Pi.
mined for prokaryotic ribosomes. It is known
In the 70 S initiation complex, formylme-
that filamentous mRNA passes through a cleft
thionine tRNA is initially located at a binding
between the two subunits near the charac-
site known as the peptidyl site (P). A second
teristic “horn” on the small subunit. tRNAs
binding site, the acceptor site (A), is not yet
also bind near this site. The illustration shows
occupied during this phase of translation.
the size of a tRNA molecule for comparison.
Sometimes, a third tRNA binding site is de-
Prokaryotic ribosomes have a similar struc-
fined as an exit site (E), from which uncharged
ture, but are somewhat smaller than those of
tRNAs leave the ribosome again (see p. 252;
eukaryotes
(sedimentation coef cient
70 S
not shown).
for the complete ribosome, 30 S and 50 S for
the subunits). Mitochondrial and chloroplast
ribosomes are comparable to prokaryotic
ones.
B. Polysomes
In cells that are carrying out intensive protein
synthesis, ribosomes are often found in a lin-
ear arrangement like a string of pearls; these
are known as polysomes. This arrangement
arises because several ribosomes are translat-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Protein biosynthesis
251
A. Structure of eukaryotic ribosomes
5S-RNA
5.8S-RNA
Large
Ribosome (80S)
(120 nt)
(160 nt)
subunit (60S)
4.2 · 106 Da
2.8 · 106 Da
mRNA
49 Proteins
28S-RNA (4718 nt)
tRNA
Small
subunit (40S)
1.4 · 106 Da
33 Proteins
18S-RNA (1874 nt)
mRNA
Hemoglobin
at the same scale
B. Polysome
C. Initiation of translation in E. coli
GTP
Start
IF-2
IF-1
IF-1
codon
GTP
(AUG)
IF-2
2
IF-3
IF-3
mRNA
N
mRNA
IF-3
IF-1
50S
3
1
Direction of
subunit
tRNA-
f-Met
IF-3
ribosomal
N
movement
30S
subunit
GTP
IF-1
IF-2
f-Met
Growing
peptide
chain
Protein
N
5
tRNA
Termination
4
Elongation
70S-
Protein
Initiation
mRNA
complex
rRNA
C
Stop codon
N
(16S)
(UAA,UAG
or
UGA)
IF-1
f-Met
IF-2
Formyl-
60S
3'
methionine
P
GDP
40S
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
252
Molecular genetics
[3] After the transfer of the growing pep-
Translation II: elongation and
tide to the A site, the free tRNA at the P site
termination
dissociates and another GTP-containing elon-
gation factor (EF-G GTP) binds to the ribo-
After translation has been initiated
(see
some. Hydrolysis of the GTP in this factor
p. 250), the peptide chain is extended by the
provides the energy for translocation of the
addition of further amino acid residues
ribosome. During this process, the ribosome
(elongation) until the ribosome reaches a
moves three bases along the mRNA in the
stop codon on the mRNA and the process is
direction of the 3
end. The tRNA carrying
interrupted (termination).
the peptide chain is stationary relative to
the mRNA and reaches the ribosome’s P site
A. Elongation and termination of protein
during translocation, while the next mRNA
biosynthesis in E. coli
codon (in this case GUG) appears at the A site.
[4] The uncharged Val-tRNA then dissoci-
Elongation can be divided into four phases:
ates from the E site. The ribosome is now
[1] Binding of aminoacyl tRNA. First, the
ready for the next elongation cycle.
peptidyl site (P) of the ribosome is occupied
When one of the three stop codons (UAA,
by a tRNA that carries at its 3
end the com-
UAG, or UGA) appears at the A site, termi-
plete peptide chain formed up to this point
nation starts.
(top left). A second tRNA, loaded with the
[5] There are no complementary tRNAs for
next amino acid (Val-tRNAVal in the example
the stop codons. Instead, two releasing factors
shown), then binds via its complementary
bind to the ribosome. One of these factors
anticodon (see p. 82) to the mRNA codon ex-
(RF-1) catalyzes hydrolytic cleavage of the
posed at the acceptor site (in this case GUG).
ester bond between the tRNA and the C-ter-
The tRNA binds as a complex with a GTP-
minus of the peptide chain, thereby releasing
containing protein, the elongation factor Tu
the protein.
(EF-Tu) (1a). It is only after the bound GTP
[6] Hydrolysis of GTP by factor RF-3 sup-
has been hydrolyzed to GDP and phosphate
plies the energy for the dissociation of the
that EF-Tu dissociates again (1b). As the bind-
whole complex into its component parts.
ing of the tRNA to the mRNA is still loose
Energy requirements in protein synthesis
before this, GTP hydrolysis acts as a delaying
are high. Four energy-rich phosphoric acid
factor, making it possible to check whether
anhydride bonds are hydrolyzed for each
the correct tRNA has been bound. A further
amino acid residue. Amino acid activation
protein, the elongation factor Ts (EF-Ts), later
uses up two energy-rich bonds per amino
catalyzes the exchange of GDP for GTP and in
acid (ATP AMP + PP; see p. 248), and two
this way regenerates the EF-Tu GTP com-
GTPs are consumed per elongation cycle. In
plex. EF-Tu is related to the G proteins in-
addition, initiation and termination each re-
volved in signal transduction (see p. 384).
quire one GTP per chain.
[2] Synthesis of the peptide bond takes
place in the next step. Ribosomal peptidyl-
transferase catalyzes (without consumption
Further information
of ATP or GTP) the transfer of the peptide
In eukaryotic cells, the number of initiation
chain from the tRNA at the P site to the NH2
factors is larger and initiation is therefore
group of the amino acid residue of the tRNA at
more complex than in prokaryotes. The cap
the A site. The ribosome’s peptidyltransferase
at the 5 end of mRNA and the polyA tail (see
activity is not located in one of the ribosomal
p. 246) play important parts in initiation.
proteins, but in the 28 S rRNA. Catalytically
However, the elongation and termination
active RNAs of this type are known as ribo-
processes are similar in all organisms. The
zymes (cf. p. 246). It is thought that the few
individual steps of bacterial translation can
surviving ribozymes are remnants of the “RNA
be inhibited by antibiotics (see p. 254).
world”—an early phase of evolution in which
proteins were not as important as they are
today.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Protein biosynthesis
253
A. Elongation and termination of protein biosynthesis in E. coli
Elongation
Peptidyl transferase
3'
1
2.3.2.12
3'
1a
1b
Valine codon
Val
GTP
Pro
GDP
Pro
EF-Tu
P
EF-Tu
Val
Val-tRNA
NH3
5'
5'
NH
3
2
GDP
EF-Ts
EF-Ts
GTP
4
3'
3'
GDP
P
GTP
EF-G
EF-G
Val
Pro
Val
Pro
NH3
5'
3
NH3
5'
tRNA
If stop codon
in A site
tRNA
5
P
GDP
50S
subunit
RF-3
GTP
Termination
30S
Termination
subunit
RF-3
factors
Stop
codon
RF-1
6
Val
Pro
mRNA
Pro
Val
NH3
COO
NH3
RF-1
Protein
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
254
Molecular genetics
synthesized by fungi and have a reactive β-
Antibiotics
lactam ring. They are mainly used against
Gram-positive pathogens, in which they in-
A. Antibiotics: overview
hibit cell wall synthesis (C).
Antibiotics are substances which, even at low
The first synthetic antibiotics were the
concentrations, inhibit the growth and repro-
sulfonamides (right). As analogues of p-ami-
duction of bacteria and fungi. The treatment
nobenzoic acid, these affect the synthesis of
of infectious diseases would be inconceivable
folic acid, an essential precursor of the coen-
today without antibiotics. Substances that
zyme THF (see p. 108). Transport antibiotics
only restrict the reproduction of bacteria are
(top center) have the properties of ion chan-
described as having bacteriostatic effects (or
nels (see p. 222). When they are deposited in
fungistatic for fungi). If the target cells are
the plasma membrane, it leads to a loss of ions
killed, then the term bactericidal (or fungici-
that damages the bacterial cells.
dal) is used. Almost all antibiotics are pro-
duced by microorganisms—mainly bacteria
B. Intercalators
of the genus Streptomyces and certain fungi.
However, there are also synthetic antibacte-
The effects of intercalators (see also p. 262)
rial substances, such as sulfonamides and gy-
are illustrated here using the example of the
rase inhibitors.
daunomycin-DNA complex, in which two
A constantly increasing problem in antibi-
daunomycin molecules (red) are inserted in
otic treatment is the development of resistant
the double helix (blue). The antibiotic’s ring
pathogens that no longer respond to the
system inserts itself between G/C base pairs
drugs available. The illustration shows a few
(bottom), while the sugar moiety occupies the
of the therapeutically important antibiotics
minor groove in the DNA (above). This leads
and their sites of action in the bacterial me-
to a localized change of the DNA conforma-
tabolism.
tion that prevents replication and transcrip-
Substances known as intercalators, such as
tion.
rifamycin and actinomycin D (bottom) are de-
posited in the DNA double helix and thereby
C. Penicillin as a “suicide substrate”
interfere with replication and transcription
(B). As DNA is the same in all cells, intercalat-
The site of action in the β-lactam antibiotics is
ing antibiotics are also toxic for eukaryotes,
muramoylpentapeptide carboxypeptidase, an
however. They are therefore only used as cy-
enzyme that is essential for cross-linking of
tostatic agents (see p. 402). Synthetic inhib-
bacterial cell walls. The antibiotic resembles
itors of DNA topoisomerase II (see p. 240),
the substrate of this enzyme (a peptide with
known as gyrase inhibitors (center), restrict
the C-terminal sequence D-Ala-D-Ala) and is
replication and thus bacterial reproduction.
therefore reversibly bound in the active cen-
A large group of antibiotics attack bacterial
ter. This brings the β-lactam ring into prox-
ribosomes. These inhibitors of translation
imity with an essential serine residue of the
(left) include the tetracyclines—broad-spec-
enzyme. Nucleophilic substitution then re-
trum antibiotics that are effective against a
sults in the formation of a stable covalent
large number of different pathogens. The ami-
bond between the enzyme and the inhibitor,
noglycosides, of which streptomycin is the
blocking the active center (see p. 96). In divid-
best-known, affect all phases of translation.
ing bacteria, the loss of activity of the enzyme
Erythromycin impairs the normal functioning
leads to the formation of unstable cell walls
of the large ribosomal subunit. Chlorampheni-
and eventually death.
col, one of the few natural nitro compounds,
inhibits ribosomal peptidyltransferase. Fi-
nally, puromycin mimics an aminoacyl tRNA
and therefore leads to premature interruption
of elongation.
The E-lactam antibiotics (bottom right) are
also frequently used. The members of this
group, the penicillins and cephalosporins, are
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Antibiotics
255
A. Antibiotics: overview
H3C
CH3
O
6
Transport antibiotics
S
N
HO
CH3
H
H2N
S
NH
OH
N
O
NH2
Sulfathiazole
C
Ion channel
OH
HO O OH O
O
DNA
Sulfonamides
Tetracycline
Proteins
Folate
Gyrase
2
synthesis
inhibitors
Aminoglycosides
3
Tetracyclines
Replication
Erythromycin
Precursors
Puromycin
THF
Translation
Chloramphenicol
Cell wall
synthesis
Transcription
Penicillins
4
Cephalosporins
OH
mRNA
DNA
O
H
HO
C
Rifamycin
1
N
CHCl
2
Actinomycin D
NH3
H
H
Daunomycin
H
C
N
S
C
CH3
Outer
Inner
Nitro
H
membrane
Murein
membrane
group
O
C N
CH3
O
NO2
H COO
Chloramphenicol
Ampicillin
B. Intercalators
C. Penicillin as “suicide substrate”
R''
O OH
O
C
R''-D-Ala-
CH3
D-Ala
OH
(substrate)
H O
H3CO O OH
H
H
O
Daunomycin
CH3
R'
H H
HO
H
H3N
H
Penicillin
(inhibitor)
R
R
S
S
CH3
CH3
C N
CH3
C HN
CH3
O
C/G pair
O
O
HO
H COO
H COO
G/C pair
Serine residue
Enzyme-inhibitor
Covalent acyl enzyme
Daunomycin-DNA complex
complex
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
256
Molecular genetics
B. Effects
Mutation and repair
Nitrous acid causes point mutations (1). For
Genetic information is set down in the base
example, C is converted to U, which in the
sequence of DNA. Changes in the DNA bases
next replication pairs with A instead of G.
or their sequence therefore have mutagenic
The alteration thus becomes permanent. Mu-
effects. Mutagens often also damage growth
tations in which a number of nucleotides not
regulation in cells, and they are then also
divisible by three are inserted or removed
carcinogenic
(see p. 400). Gene alterations
lead to reading errors in whole segments of
(mutations) are one of the decisive positive
DNA, as they move the reading frame (frame-
factors in biological evolution. On the other
shift mutations). This is shown in Fig. 2 using
hand, an excessive mutation frequency would
a simple example. From the inserted C on-
threaten the survival of individual organisms
wards, the resulting mRNA is interpreted dif-
or entire species. For this reason, every cell
ferently during translation, producing a com-
has repair mechanisms that eliminate most of
pletely new protein sequence.
the DNA changes arising from mutations (C).
C. Repair mechanisms
A. Mutagenic agents
An important mechanism for the removal of
Mutations can arise as a result of physical or
DNA damage is excision repair (1). In this
chemical effects, or they can be due to acci-
process, a specific excision endonuclease re-
dental errors in DNA replication and recombi-
moves a complete segment of DNA on both
nation.
sides of the error site. Using the sequence of
The principal physical mutagen is ionizing
the opposite strand, the missing segment is
radiation (α, β, and γ radiation, X-rays). In
then replaced by a DNA polymerase. Finally, a
cells, it produces free radicals
(molecules
DNA ligase closes the gaps again.
with unpaired electrons), which are ex-
Thymine dimers can be removed by
tremely reactive and can damage DNA.
photoreactivation (2). A specific photolyase
Short-wavelength ultraviolet light (UV light)
binds at the defect and, when illuminated,
also has mutagenic effects, mainly in skin cells
cleaves the dimer to yield two single bases
(sunburn). The most common chemical
again.
change due to UV exposure is the formation
A third mechanism is recombination repair
of thymine dimers, in which two neighboring
(3, shown in simplified form). In this process,
thymine bases become covalently linked to
the defect is omitted during replication. The
one another (2). This results in errors when the
gap is closed by shifting the corresponding
DNA is read during replication and transcription.
sequence from the correctly replicated second
Only a few examples of the group of chem-
strand. The new gap that results is then filled
ical mutagens are shown here. Nitrous acid
by polymerases and ligases. Finally, the orig-
(HNO2; salt: nitrite) and hydroxylamine
inal defect is corrected by excision repair as in
(NH2OH) both deaminate bases; they convert
Fig. 1 (not shown).
cytosine to uracil and adenine to inosine.
Alkylating compounds carry reactive
groups that can form covalent bonds with
DNA bases (see also p. 402). Methylnitro-
samines (3) release the reactive methyl cation
(CH3+), which methylates OH and NH2 groups
in DNA. The dangerous carcinogen benzo
[a]pyrene is an aromatic hydrocarbon that is
only converted into the active form in the
organism (4; see p. 316). Multiple hydroxyla-
tion of one of the rings produces a reactive
epoxide that can react with NH2 groups in
guanine residues, for example. Free radicals
of benzo[a ]pyrene also contribute to its tox-
icity.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Mutation and repair
257
A. Mutagenic agents
1.
Ionizing
Alkylating
Reactive
radiation
compounds
methyl
group
CH3
O
NH
UV
HNO2
α, β, γ
[CH
O
O
3 ]
C
C
radi-
N
X
ation
R
NO
N
CH3
R
CHC
3.
Methylnitrosamine
O
NH
O
C
C
N
Desami-
Free
R
CHC
CH3
Epoxide
nation
radicals
2. Thymine dimer
O
Deletions
CH
or
HC
insertions
C H
due to
HO
H
C
faulty
OH
recombination
Formation Base
Spon-
Chemical
4.
Mutagenic
of
exchange
taneous
modificaton
derivative of
pyrimidine
C U
loss of
of bases
benzo(a)pyrene
dimers
A I
bases
B. Effects
C. Repair mechanisms
Thymine dimer
·
GT
A
C
C
T
·
·
C
A
T
G
G
A
·
HNO2
Base
a
a
exchange
Excision
endonuclease
Photo-
·
GTAUC
T
·
lyase
·
CAT
G
GA
·
4.1.99.3
Replication
b
Light
·
GTA
C
C
T
·
·
G
TA
U
C
T
·
b
·
CAT
G
GA
·
·
C
AT
A
GA
·
Defect-
binding
Normal
Replication
protein
c
·
GTAT
C
T
·
2. Photoreactivation
·
CATAGA
·
c
Single-strand
Permanent
binding
DNA poly-
proteins
a
change
Replication
merase
1. Point mutation
b
Insertion
d
·
G
CTA
C
C
T
·
·
C
CATGG
A
·
DNA ligase
·
G
TA
C
CT
·
·
G
C
TA
C
C
T
·
·
C
AT
G
G
A
·
·
CG
AT
G
G
A
·
e
Repaired DNA
·· Val-Pro-
·· -Ala-Thr-X
··
c
2. Frameshift mutation
1. Excision repair
3. Recombinational repair
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
258
Molecular genetics
B. DNA cloning
DNA cloning
Most DNA segments—e. g., genes—occur in
The growth of molecular genetics since 1970
very small quantities in the cell. To be able
has mainly been based on the development
to work with them experimentally, a large
and refinement of methods of analyzing and
number of identical copies (“clones”) first
manipulating DNA. Genetic engineering has
have to be produced. The classic procedure
practical applications in many fields. For ex-
for cloning DNA takes advantage of the ability
ample, it has provided new methods of diag-
of bacteria to take up and replicate short,
nosing and treating diseases, and it is now
circular DNA fragments known as plasmids.
also possible to create targeted changes in
The segment to be cloned is first cut out of
specific characteristics of organisms. Since bi-
the original DNA using restriction endonu-
ological risks cannot be completely ruled out
cleases (see above; for the sake of simplicity,
with these procedures, it is particularly im-
cleavage using EcoRI alone is shown here, but
portant to act responsibly when dealing with
in practice two different enzymes are usually
genetic engineering. A short overview of im-
used). As a vehicle (“vector”), a plasmid is
portant methods involved in genetic engi-
needed that has only one EcoRI cleavage site.
neering is provided here and on the following
The plasmid rings are first opened by cleavage
pages.
with EcoRI and then mixed with the isolated
DNA fragments. Since the fragment and the
A. Restriction endonucleases
vector have the same overhanging ends, some
of the molecules will hybridize in such a way
In many genetic engineering procedures, de-
that the fragment is incorporated into the
fined DNA fragments have to be isolated and
vector DNA. When the cleavage sites are
then newly combined with other DNA seg-
now closed again using DNA ligase, a newly
ments. For this purpose, enzymes are used
combined (“recombinant”) plasmid arises.
that can cut DNA and join it together again
By pretreating a large number of host cells,
inside the cell. Of particular importance are
one can cause some of them to take up the
restriction endonucleases—a group of bacterial
plasmid (a process known as transformation)
enzymes that cleave the DNA double strand in
and replicate it along with their own genome
a sequence-specific way. The numerous re-
when reproducing. To ensure that only host
striction enzymes known are named using
bacteria that contain the plasmid replicate,
abbreviations based on the organism from
plasmids are used that give the host resistance
which they originate. The example used
to a particular antibiotic. When the bacteria
here is EcoRI, a nuclease isolated from the
are incubated in the presence of this antibi-
bacterium Escherichia coli.
otic, only the cells containing the plasmid will
Like many other restriction endonucleases,
replicate. The plasmid is then isolated from
EcoRI cleaves DNA at the site of a palin-
these cells, cleaved with EcoRI again, and the
drome—i. e., a short segment of DNA in which
fragments are separated using agarose gel
both the strand and counter-strand have the
electrophoresis (see p. 262). The desired frag-
same sequence (each read in the 5
3 direc-
ment can be identified using its size and then
tion). In this case, the sequence is 5 -GAATTC-
extracted from the gel and used for further
3 . EcoRI, a homodimer, cleaves the phos-
experiments.
phoric acid diester bonds in both strands be-
tween G and A. This results in the formation of
complementary overhanging or “sticky” ends
(AATT), which are held together by base pair-
ing. However, they are easily separated—e. g.
by heating. When the fragments are cooled,
the overhanging ends hybridize again in the
correct arrangement. The cleavage sites can
then be sealed again by a DNA ligase.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Genetic engineering
259
A. Restriction endonucleases
H
Palindrome
H
H
5'
3'
3'
G
G
A
A
T
T
C
O H
C
T
T
A
A
G
3'
5'
O
P
O H2C5'
EcoRI
DNA-Ligase
O
H
A
3.1.21.4
6.5.1.1
H
5'
3'
G
A
A
T
T
C
C
T
T
A
A
G
H
3'
5'
3'
G
OH H
O
A
A
T
T
C
3'
5'
G
O
P
O
H2C5'
G
5'
C
T
T
A A
A
3'
O
EcoRI
+
DNA
Overhanging ends
B. DNA cloning
DNA
G
A
A
T T C
G
A
A
T T
C
C
T
T
A A
G
C
T
T
A A
G
EcoRI
EcoRI
Purification
Vector DNA
(Plasmid)
Isolated
DNA ligase
DNA fragment
Host cell
Gene for
(bacterium)
antibiotic
resistance
Recombinant
plasmid
Transformation
Plasmid
Plasmid
isolation
Cleavage
Bacterial
with
genome
EcoRI
Bacterial culture in the
Cloned
presence of antibiotic
DNA fragment
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
260
Molecular genetics
B. Sequencing of DNA
DNA sequencing
The nucleotide sequence of DNA is nowadays
A. Gene libraries
usually determined using the so-called chain
termination method. In single-strand se-
It is often necessary in genetic engineering to
quencing, the DNA fragment (a) is cloned
isolate a DNA segment when its details are not
into the DNA of phage M13 (see p. 404),
fully known—e. g., in order to determine its
from which the coded single strand can be
nucleotide sequence. In this case, one can use
easily isolated. This is hybridized with a pri-
what are known as DNA libraries. A DNA li-
mer—a short, synthetically produced DNA
brary consists of a large number of vector DNA
fragment that binds to 3
end of the intro-
molecules containing different fragments of
duced DNA segment (b).
foreign DNA. For example, it is possible to take
Based on this hybrid, the missing second
all of the mRNA molecules present in a cell and
strand can now be generated in the test tube
transcribe them into DNA. These DNA frag-
by adding the four deoxyribonucleoside tri-
ments (known as copy DNA or cDNA) are then
phosphates (dNTP) and a suitable DNA poly-
randomly introduced into vector molecules.
merase (c). The trick lies in also adding small
A library of genomic DNA can be estab-
amounts of dideoxynucleoside triphosphates
lished by cleaving the total DNA from a cell
(ddNTP). Incorporating a ddNTP leads to the
into small fragments using restriction endo-
termination of second-strand synthesis. This
nucleases (see p. 258), and then incorporating
can occur whenever the corresponding dNTP
these into vector DNA. Suitable vectors for
ought to be incorporated. The illustration
gene libraries include bacteriophages, for ex-
shows this in detail using the example of
ample (“phages” for short). Phages are viruses
ddGTP. In this case, fragments are obtained
that only infect bacteria and are replicated by
that each include the primer plus three, six,
them (see p. 404). Gene libraries have the
eight, 13, or 14 additional nucleotides. Four
advantage that they can be searched for spe-
separate reactions, each with a different
cific DNA segments, using hybridization with
ddNTP, are carried out (c), and the products
oligonucleotides.
are placed side by side on a supporting mate-
The first step is to strongly dilute a small
rial. The fragments are then separated by gel
part of the library (105-106 phages in a small
electrophoresis
(see p. 76), in which they
volume), mix it with host bacteria, and plate
move in relation to their length.
out the mixture onto nutrient medium. The
Following visualization (d), the sequence of
bacteria grow and form a continuous cloudy
the fragments in the individual lanes is simply
layer of cells. Bacteria infected by phages
read from bottom to top (e) to directly obtain
grow more slowly. In their surroundings, the
the nucleotide sequence. A detail from such a
bacterial “lawn” is less dense, and a clearer
sequencing gel and the corresponding protein
circular zone known as a plaque forms. The
sequence are shown in Fig. 2.
bacteria in this type of plaque exclusively
In a more modern procedure, the four
contain the offspring of a single phage from
ddNTPs are covalently marked with fluores-
the library.
cent dyes, which produce a different color for
The next step is to make an impression of
each ddNTP on laser illumination. This allows
the plate on a plastic foil, which is then
the sequence in which the individual frag-
heated. This causes the phage DNA to adhere
ments appear at the lower end of the gel to
to the foil. When the foil is incubated with a
be continuously recorded and directly stored
DNA fragment that hybridizes to the DNA seg-
in digital form.
ment of interest (a gene probe), the probe
binds to the sites on the imprint at which
the desired DNA is attached. Binding of the
gene probe can be detected by prior radio-
active or other labeling of the probe. Phages
from the positive plaques in the original plate
are then isolated and replicated. Restriction
cleavage finally provides large amounts of the
desired DNA.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Genetic engineering
261
A. Gene libraries
Imprint on
Host cell
plastic membrane
Gene library
in phages
Gene probe
Plate
Hybrid-
with phage plaques
ization
Positive plaque
Washing,
detection
of the
gene probe
DNA
Gene
Gene probe
isolation
Phage
Restriction
repli-
Cloned gene
cation
Membrane
B. Sequencing of DNA
G A T C
5'
3'
a
3'
5'
Protein
sequence
Synthetic primer
Single-stranded preparation
Hybridization
DNA
5'
sequence
b
3'
3'
N
C
c
T
Phe
T
G
A
T
C
C
A
Tyr
+ 4 dNTP
+ 4 dNTP
+ 4 dNTP
+ 4 dNTP
T
+ ddGTP
+ ddATP
+ ddTTP
+ ddCTP
T
+ Polymerase
+ Polymerase
+ Polymerase
+ Polymerase
C
Ala
G
T
G
A T C
C
Thr
Example:
G
A
= ddG
A
A
Glu
G
A
A
Glu
G
G
d
Gel electrophoresis
T
Met
e
Visualization of the fragments
A
f
Reading off (
) : ACGATG ...
5'
N
1. Method
2. Sequence pattern
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
262
Molecular genetics
on their mass alone. The supporting material
PCR and protein expression
generally used in genetic engineering is a gel
of the polysaccharide agarose
(see p. 40).
A. Polymerase chain reaction (PCR)
Agarose gels are not very stable and are there-
The polymerase chain reaction (PCR) is an
fore poured horizontally into a plastic cham-
important procedure in genetic engineering
ber in which they are used for separation
that allows any DNA segment to be replicated
(top).
(amplified) without the need for restriction
To make the separated fragments visible,
enzymes, vectors, or host cells (see p. 258).
after running the procedure the gels are
However, the nucleotide sequence of the seg-
placed in solutions of ethidium bromide.
ment has to be known. Two oligonucleotides
This is an intercalator (see p. 254) that shows
(primers) are needed, which each hybridize
strong fluorescence in UV light after binding
with one of the strands at each end of the DNA
to DNA, although it barely fluoresces in an
segment to be amplified; also needed are suf-
aqueous solution. The result of separating
ficient quantities of the four deoxyribonucleo-
two PCR amplificates (lanes 1 and 2) is shown
side triphosphates and a special heat-tolerant
in the lower part of the illustration. Compar-
DNA polymerase. The primers are produced
ing their distances with those of poly-
by chemical synthesis, and the polymerase is
nucleotides of known lengths (lane 3; bp =
obtained from thermostable bacteria.
base pairs) yields lengths of approximately
First, the starter is heated to around 90 °C
800 bp for fragment 1 and 1800 bp for frag-
to separate the DNA double helix into single
ment 2. After staining, the bands can be cut
strands
(a; cf. p. 84). The mixture is then
out of the gel and the DNA can be extracted
cooled to allow hybridization of the primers
from them and used for further experiments.
(b). Starting from the primers, complemen-
tary DNA strands are now synthesized in
C. Overexpression of proteins
both directions by the polymerase (c). This
cycle (cycle 1) is repeated 20-30 times with
To treat some diseases, proteins are needed
the same reaction mixture (cycle 2 and sub-
that occur in such small quantities in the or-
sequent cycles). The cyclic heating and cool-
ganism that isolating them on a large scale
ing are carried out by computer-controlled
would not be economically feasible. Proteins
thermostats.
of this type can be obtained by overexpression
After only the third cycle, double strands
in bacteria or eukaryotic cells. To do this, the
start to form with a length equal to the dis-
corresponding gene is isolated from human
tance between the two primers. The propor-
DNA and cloned into an expression plasmid
tion of these approximately doubles during
as described on p. 258. In addition to the gene
each cycle, until almost all of the newly syn-
itself, the plasmid also has to contain DNA
thesized segments have the correct length.
segments that allow replication by the host
cell and transcription of the gene. After trans-
formation and replication of suitable host
B. DNA electrophoresis
cells, induction is used in a targeted fashion
The separation of DNA fragments by electro-
to trigger ef cient transcription of the gene.
phoresis is technically simpler than protein
Translation of the mRNA formed in the host
electrophoresis (see p. 78). The mobility of
cell then gives rise to large amounts of the
molecules in an electrical field of a given
desired protein. Human insulin (see p. 76),
strength depends on the size and shape of
plasminogen activators for dissolving blood
the molecules, as well as their charge. In con-
clots (see p. 292), and the growth hormone
trast to proteins, in which all three factors
somatotropin are among the proteins pro-
vary, the ratio of mass to charge in nucleic
duced in this way.
acids is constant, as all of the nucleotide com-
ponents have similar masses and carry one
negative charge. When electrophoresis is car-
ried out in a wide-meshed support material
that does not separate according to size and
shape, the mobility of the molecules depends
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Genetic engineering
263
A. Polymerase chain reaction (PCR)
B. DNA electrophoresis
Electrophoresis chamber Sample
DNA
a
Heat
b
Hybridize
Cycle
Primer
1
Power supply
De-novo synthesis
unit
c
(DNA polymerase)
Agarose gel
a
b
c
H3C
Cycle
Br N
2
NH2
Ethidium bromide
1
2
3
bp
a
b
c
1857
b
Cycle
1058
3
929
a
383
etc
C. Overexpression of proteins
Ribosome
Gene to be
Host cell
binding
expressed
site
Trans-
formation
Repli-
Inducible
cation
promotor
Origin of
Gene for
replication
antibiotic
resistance
Overexpressed
Protein
Cell
protein
purification
lysis
Expression plasmid
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
264
Molecular genetics
amplify a segment of this DNA with virus-
Genetic engineering in medicine
specific primers. In this way, an amplificate
Genetic engineering procedures are becom-
with a characteristic length can be obtained
ing more and more important in medicine
for each pathogen and identified using gel
for diagnostic purposes (A-C). New genetic
electrophoresis as described above.
approaches to the treatment of severe dis-
eases are still in the developmental stage
D. Gene therapy
(“gene therapy,” D).
Many diseases, such as hereditary metabolic
defects and tumors, can still not be ad-
A. DNA fingerprinting
equately treated. About 10 years ago, projects
were therefore initiated that aimed to treat
DNA fingerprinting is used to link small
diseases of this type by transferring genes
amounts of biological material—e. g., traces
into the affected cells (gene therapy). The
from the site of a crime—to a specific person.
illustration combines conceivable and already
The procedure now used is based on the fact
implemented approaches to gene therapy for
that the human genome contains non-coding
metabolic defects (left) and tumors (right).
repetitive DNA sequences, the length of which
None of these procedures has yet become
varies from individual to individual. Short
established in clinical practice.
tandem repeats (STRs) thus exist in which
If a mutation leads to failure of an enzyme
dinucleotides (e. g., -T-X-) are frequently re-
E1 (left), its substrate B will no longer be
peated. Each STR can occur in five to 15 differ-
converted into C and will accumulate. This
ent lengths (alleles), of which one individual
can lead to cell damage by B itself or by a
possesses only one or two. When the various
toxic product formed from it. Treatment
allele combinations for several STRs are de-
with intact E1 is not possible, as the proteins
termined after PCR amplification of the DNA
are not capable of passing through the cell
being investigated, a “genetic fingerprint” of
membrane. By contrast, it is in principle pos-
the individual from whom the DNA originates
sible to introduce foreign genes into the cell
is obtained. Using comparative material—e. g.,
using viruses as vectors (adenoviruses or ret-
saliva samples—definite identification is then
roviruses are mainly used). Their gene prod-
possible.
ucts could replace the defective E1 or convert
B into a harmless product. Another approach
B. Diagnosis of sickle-cell anemia using RFLP
uses the so-called antisense DNA (bottom
This example illustrates a procedure for diag-
right). This consists of polynucleotides that
nosing a point mutation in the β-globin gene
hybridize with the mRNA for specific cellular
that leads to sickle-cell anemia (see p. 248).
proteins and thereby prevent their transla-
The mutation in the first exon of the gene
tion. In the case shown, the synthesis of E2
destroys a cleavage site for the restriction
could be blocked, for example.
endonuclease MstII (see p. 258). When the
The main problem in chemotherapy for
DNA of healthy and diseased individuals is
tumors is the lack of tumor-specificity in the
cleaved with MstII, different fragments are
highly toxic cytostatic agents used
(see
produced in the region of the β-globin gene,
p. 402). Attempts are therefore being made
which can be separated by electrophoresis
to introduce into tumor cells genes with prod-
and then demonstrated using specific probes
ucts that are only released from a precursor to
(see p. 260). In addition, heterozygotic car-
form active cytostatics once they have
riers of the sickle-cell gene can be distin-
reached their target (left). Other gene prod-
guished from homozygotic ones.
ucts are meant to force the cells into apoptosis
(see p. 396) or make them more susceptible
C. Identification of viral DNA using RT-PCR
to attack by the immune system. To steer the
In viral infections, it is often dif cult to deter-
viral vectors to the tumor
(targeting), at-
mine the species of the pathogen precisely.
tempts are being made to express proteins
RT-PCR can be used to identify RNA viruses. In
on the virus surface that are bound by tu-
this procedure, reverse transcriptase
(see
mor-specific receptors. Fusion with a tumor-
p. 404) is used to transcribe the viral RNA
specific promoter could also help limit the
into dsDNA, and then PCR is employed to
effect of the foreign gene to the tumor cells.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Genetic engineering
265
A. DNA fingerprinting
B. Diagnosis of sickle-cell anemia using RFLP
STR, z.B.
Mst II
Mst II
TCTATCTGTCTG
DNA
1200 bp
201 bp
Normal
Extraction
gene
E1
I1
E2
Sickle-cell
gene
1401 bp
Evidence
Mst II
Mst II
PCR
Mutation
Locus 1
1.4
Locus 2
1 Normal (A/A)
1.2
2 Heterozygotic (A/S)
Locus 3
1
2
3
kbp
3 Homozygotic (S/S)
Amplificate
Comparative
C. Evidence of viral DNA using RT-PCR
fragments
RNA Hy-
ss-
ds-
brid DNA
DNA
Amplificate
Standards
P2
Alleles
Virus
1/3
Locus 1
PCR
1/4
P3
Locus 2
2/4
P1
3/3
Locus 3
Agarose gel
3/5
Reverse
DNA polymerase
Agarose gel
transcriptase
D. Gene therapy
Tumor-
Viral vector
Viral vector with
specific
with foreign DNA
foreign DNA
receptor
Normal
Tumor-
Tumor cell
body cell
specific
DNA
DNA
promoter
E1
mRNA
mRNA
Harmless
product
Immune
E1
Gene
system
product
Gene
product
E1
Accu-
Cytostatic
mulates
agent
A
B
E1
C
Apoptosis
E2
Genome
Toxic
Cell
Cell
Antisense
product
Lipo-
proliferation
death
DNA
some
1. For metabolic defects
2. For tumors
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
266
Tissues and organs
phosphate, and nucleosides (nucleobase pen-
Digestion: overview
tose). These cleavage products are resorbed
Most components of food (see p. 360) cannot
by the intestinal wall in the region of the
be resorbed directly by the organism. It is only
jejunum.
after they have been broken down into
Lipids are a special problem for digestion,
smaller molecules that the organism can
as they are not soluble in water. Before enzy-
take up the essential nutrients. Digestion re-
matic breakdown, they have to be emulsified
fers to the mechanical and enzymatic break-
by bile salts and phospholipids in the bile (see
down of food and the resorption of the result-
p. 314). At the water-lipid interface, pancre-
ing products.
atic lipase then attacks triacylglycerols with
the help of colipase (see p. 270). The cleavage
products include fatty acids, 2-monoacylgly-
A. Hydrolysis and resorption of food
cerols, glycerol, and phosphate from phospho-
components
lipid breakdown. After resorption into the
Following mechanical fragmentation of food
epithelial cells, fats are resynthesized from
during chewing in the mouth, the process of
fatty acids, glycerol and 2-monoacylglycerols
enzymatic degradation starts in the stomach.
and passed into the lymphatic system (see
For this purpose, the chyme is mixed with
p. 272). The lipids in milk are more easily
digestive enzymes that occur in the various
digested, as they are already present in emul-
digestive secretions or in membrane-bound
sion; on cleavage, they mostly provide short-
form on the surface of the intestinal epithe-
chain fatty acids.
lium (see p. 268). Almost all digestive en-
Inorganic components such as water, elec-
zymes are hydrolases (class 3 enzymes; see
trolytes, and vitamins are directly absorbed by
p. 88); they catalyze the cleavage of compo-
the intestine.
site bonds with the uptake of water.
High-molecular-weight indigestible com-
Proteins are first denatured by the
ponents, such as the fibrous components of
stomach’s hydrochloric acid (see p. 270), mak-
plant cell walls, which mainly consist of cel-
ing them more susceptible to attack by the
lulose and lignin, pass through the bowel un-
endopeptidases (proteinases) present in gas-
changed and form the main component of
tric and pancreatic juice. The peptides re-
feces, in addition to cells shed from the intes-
leased by endopeptidases are further de-
tinal mucosa. Dietary fiber makes a positive
graded into amino acids by exopeptidases. Fi-
contribution to digestion as a ballast material
nally, the amino acids are resorbed by the
by binding water and promoting intestinal
intestinal mucosa in cotransport with Na+
peristalsis.
ions (see p. 220). There are separate transport
The food components resorbed by the epi-
systems for each of the various groups of
thelial cells of the intestinal wall in the region
amino acids.
of the jejunum and ileum are transported
Carbohydrates mainly occur in food in the
directly to the liver via the portal vein. Fats,
form of polymers (starches and glycogen).
cholesterol, and lipid-soluble vitamins are
They are cleaved by pancreatic amylase into
exceptions. These are first released by the
oligosaccharides and are then hydrolyzed by
enterocytes in the form of chylomicrons (see
glycosidases, which are located on the surface
p. 278) into the lymph system, and only reach
of the intestinal epithelium, to yield mono-
the blood via the thoracic duct.
saccharides. Glucose and galactose are taken
up into the enterocytes by secondary active
cotransport with Na+ ions (see p. 220). In ad-
dition, monosaccharides also have passive
transport systems in the intestine.
Nucleic acids are broken down into their
components by nucleases from the pancreas
and small intestine (ribonucleases and deoxy-
ribonucleases). Further breakdown yields the
nucleobases (purine and pyrimidine deriva-
tives), pentoses
(ribose and deoxyribose),
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Digestion
267
A. Hydrolysis and resorption of dietary constituents
Proteins
Carbohydrates Nucleic acids
Lipids
Vitamins
Fiber
Inorganic
Bile salts
substances
HCl
Phospholipids
E n z y m a t i c h y d r o l y s i s
2-Monoacyl-
Cellulose
glycerol
Lignin
Fatty acids
Cholesterol
Nucleobases
Pentoses
Amino
Mono-
Phosphate
Glycerol
acids
saccharides
Nucleosides
phosphate
Glucose
Galactose
*
*
*
R e s o r p t i o n
Hydrophilic
Lipophilic
Portal vessel
Lymph system
Liver
Blood
Feces
*Resorption by active transport
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
268
Tissues and organs
Elastase mainly cleaves on the C side of the
Digestive secretions
aliphatic amino acids Gly, Ala, Val, and Ile.
Smaller peptides are attacked by carboxy-
A. Digestive juices
peptidases, which as exopeptidases cleave in-
Saliva. The salivary glands produce a slightly
dividual amino acids from the C-terminal end
alkaline secretion which—in addition to water
of the peptides (see p. 176).
and salts—contains glycoproteins (mucins) as
a-Amylase, the most important endoglyco-
lubricants,
antibodies,
and enzymes.
sidase in the pancreas, catalyzes the hydroly-
D-Amylase attacks polysaccharides, and a li-
sis of α1
4 bonds in the polymeric carbohy-
pase hydrolyzes a small proportion of the
drates starch and glycogen. This releases mal-
neutral fats. α-Amylase and lysozyme, a mu-
tose, maltotriose, and a mixture of other oli-
rein-cleaving enzyme (see p. 40), probably
gosaccharides.
serve to regulate the oral bacterial flora rather
Various pancreatic enzymes hydrolyze lip-
than for digestion (see p. 340).
ids, including lipase with its auxiliary protein
Gastric juice. In the stomach, the chyme is
colipase (see p. 270), phospholipase A2, and
mixed with gastric juice. Due to its hydro-
sterol esterase. Bile salts activate the lipid-
chloric acid content, this secretion of the gas-
cleaving enzymes through micelle formation
tric mucosa is strongly acidic (pH 1-3; see
(see below).
p. 270). It also contains mucus (mainly glyco-
Several hydrolases—particularly ribo-
proteins known as mucins), which protects
nuclease
(RNAse) and deoxyribonuclease
the mucosa from the hydrochloric acid, salts,
(DNAse)—break down the nucleic acids con-
and pepsinogen—the proenzyme (“zymogen”)
tained in food.
of the aspartate proteinase pepsin
(see
Bile. The liver forms a thin secretion (bile)
pp. 176, 270). In addition, the gastric mucosa
that is stored in the gallbladder after water
secretes what is known as “intrinsic factor”—a
and salts have been extracted from it. From
glycoprotein needed for resorption of vitamin
the gallbladder, it is released into the duode-
B12 ( “extrinsic factor”) in the bowel.
num. The most important constituents of bile
In the stomach, pepsin and related en-
are water and inorganic salts, bile acids and
zymes initiate the enzymatic digestion of pro-
bile salts (see p. 314), phospholipids, bile pig-
teins, which takes 1-3 hours. The acidic gas-
ments, and cholesterol. Bile salts, together
tric contents are then released into the duo-
with phospholipids, emulsify insoluble food
denum in batches, where they are neutralized
lipids and activate the lipases. Without bile,
by alkaline pancreatic secretions and mixed
fats would be inadequately cleaved, if at all,
with cystic bile.
resulting in
“fatty stool”
(steatorrhea). Re-
Pancreatic secretions. In the acinar cells,
sorption of fat-soluble vitamins would also
the pancreas forms a secretion that is alkaline
be affected.
due to its HCO3- content, the buffer capacity
Small-intestinal secretions. The glands of
of which is suf cient to neutralize the stom-
the small intestine (the Lieberkühn and Brun-
ach’s hydrochloric acid. The pancreatic secre-
ner glands) secrete additional digestive en-
tion also contains many enzymes that catalyze
zymes into the bowel. Together with enzymes
the hydrolysis of high-molecular-weight food
on the microvilli of the intestinal epithelium
components. All of these enzymes are hydro-
(peptidases, glycosidases, etc.), these en-
lases with pH optimums in the neutral or
zymes ensure almost complete hydrolysis of
weakly alkaline range. Many of them are
the food components previously broken
formed and secreted as proenzymes and are
down by the endoenzymes.
only activated in the bowel lumen
(see
p. 270).
Trypsin, chymotrypsin, and elastase are en-
dopeptidases that belong to the group of ser-
ine proteinases (see p. 176). Trypsin hydro-
lyzes specific peptide bonds on the C side of
the basic amino acids Arg and Lys, while chy-
motrypsin prefers peptide bonds of the apolar
amino acids Tyr, Trp, Phe, and Leu (see p. 94).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Digestion
269
A. Digestive juices
Saliva
Endoenzyme
Daily secretion 1.0-1.5 l
Exoenzyme
pH 7
Water
Moistens food
Salts
Mucus
Lubricant
Compo-
Function
nents
or substrate
Antibodies
Bind to bacteria
α-Amylases (3.2.1.1)
Cleave starch
Lysozyme (3.2.1.17) Attacks bacterial cell walls
Gastric juice
Daily secretion 2-3 l
pH 1
Bile
Water
Daily secretion 0.6 l
Salts
pH 6.9-7.7
HCl
Denatures proteins,
Water
kills bacteria
HCO3
Neutralizes gastric juice
Mucus
Protects stomach lining
Bile salts
Facilitate lipid digestion
Pepsins (3.4.23.1-3)
Cleave proteins
Phospholipids
Facilitate lipid digestion
Chymosin (3.4.23.4)
Precipitates casein
Bile pigments
Waste products
Triacylglycerol lipase (3.1.1.3)
Cleaves fats
Cholesterol
Waste product
Intrinsic factor
Protects vitamin B12
Pancreatic secretions
Daily secretion 0.7-2.5 l
pH 7.7 (7.5-8.8)
Water
HCO3
Neutralizes gastric juice
Trypsin (3.4.21.4)
Proteins
Chymotrypsin (3.4.21.1)
Proteins
Elastase (3.4.21.36)
Proteins
Carboxypeptidases (3.4.n.n)
Peptides
α-Amylase (3.2.1.1)
Starch and glycogen
Triacylglycerol lipase (3.1.1.3)
Fats
Co-Lipase
Cofactor for lipase
Phospholipase A2 (3.1.1.4)
Phospholipids
Sterol esterase (3.1.1.13)
Cholesterol esters
Ribonuclease (3.1.27.5)
RNA
Deoxyribonuclease I (3.1.21.1)
DNA
Secretions of the small intestine
Daily secretion unknown
pH 6.5-7.8
Aminopeptidases (3.4.11.n)
Peptides
Dipeptidases (3.4.13.n)
Dipeptides
α-Glucosidase (3.2.1.20)
Oligosaccharides
Oligo-1,6-glucosidase (3.2.1.10)
Oligosaccharides
β-Galactosidase (3.2.1.23)
Lactose
Sucrose α-glucosidase (3.2.1.48)
Sucrose
α, α-Trehalase (3.2.1.28)
Trehalose
Alkaline phosphatase (3.1.3.1)
Phosphoric acid esters
Polynucleotidases (3.1.3.n)
Nucleic acids, nucleotides
Nucleosidases (3.2.2.n)
Nucleosides
Phospholipases (3.1.n.n)
Phospholipids
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
270
Tissues and organs
stimulate pancreatic secretion and bile re-
Digestive processes
lease.
Gastric juice is the product of several cell
types. The parietal cells produce hydrochloric
B. Zymogen activation
acid, chief cells release pepsinogen, and acces-
sory cells form a mucin-containing mucus.
To prevent self-digestion, the pancreas re-
leases most proteolytic enzymes into the du-
odenum in an inactive form as proenzymes
A. Formation of hydrochloric acid
(zymogens). Additional protection from the
The secretion of hydrochloric acid (H+ and Cl-)
effects of premature activation of pancreatic
by the parietal cells is an active process that
proteinases is provided by proteinase inhibi-
uses up ATP and takes place against a concen-
tors in the pancreatic tissue, which inactivate
tration gradient (in the gastric lumen, with a
active enzymes by complex formation (right).
pH of 1, the H+ concentration is some 106
Trypsinogen plays a key role among the
times higher than in the parietal cells, which
proenzymes released by the pancreas. In the
have a pH of 7).
bowel, it is proteolytically converted into ac-
The precursors of the exported H+ ions are
tive trypsin (see p. 176) by enteropeptidase, a
carbon dioxide (CO2) and water (H2O). CO2
membrane enzyme on the surface of the en-
diffuses from the blood into the parietal cells,
terocytes. Trypsin then autocatalytically acti-
and in a reaction catalyzed by carbonate de-
vates additional trypsinogen molecules and
hydratase (carbonic anhydrase [2]), it reacts
the other proenzymes (left).
with H2O to form H+ and hydrogen carbonate
(HCO3-). The H+ ions are transported into the
C. Fat digestion
gastric lumen in exchange for K+ by a mem-
brane-bound H+/K+-exchanging ATPase [1] (a
Due to the “hydrophobic effect” (see p. 28),
transport ATPase of the P type; see p. 220).
water-insoluble neutral fats in the aqueous
The remaining hydrogen carbonate is re-
environment of the bowel lumen would ag-
leased into the interstitium in electroneutral
gregate into drops of fat in which most of the
antiport in exchange for chloride ions (Cl-),
molecules would not be accessible to pancre-
and from there into the blood. The Cl- ions
atic lipase. The amphipathic substances in bile
follow the secreted protons through a chan-
(bile acids, bile salts, phospholipids) create an
nel into the gastric lumen.
emulsion in which they occupy the surface of
The hydrochloric acid in gastric juice is
the droplets and thereby prevent them from
important for digestion. It activates pepsin-
coalescing into large drops. In addition, the
ogen to form pepsin (see below) and creates
bile salts, together with the auxiliary protein
an optimal pH level for it to take effect. It also
colipase, mediate binding of triacylglycerol
denatures food proteins so that they are more
lipase [1] to the emulsified fat droplets. Acti-
easily attacked by proteinases, and it kills
vation of the lipase is triggered by a confor-
micro-organisms.
mation change in the C-terminal domain of
Regulation. HCl secretion is stimulated by
the enzyme, which uncovers the active center.
the peptide hormone gastrin, the mediator
During passage through the intestines, the
histamine (see p. 380), and—via the neuro-
active lipase breaks down the triacylglycerols
transmitter acetylcholine—by the autonomous
in the interior of the droplets into free fatty
nervous system. The peptide somatostatin
acids and amphipathic monoacylglycerols.
and certain prostaglandins (see p. 390) have
Over time, smaller micelles develop
(see
inhibitory effects. Together with cholecysto-
p. 28), in the envelope of which monoacylgly-
kinin, secretin, and other peptides, gastrin
cerols are present in addition to bile salts and
belongs to the group of gastrointestinal hor-
phospholipids. Finally, the components of the
mones (see p. 370). All of these are formed in
micelles are resorbed by the enterocytes in
the gastrointestinal tract and mainly act in the
ways that have not yet been explained.
vicinity of the site where they are formed—
Monoacylglycerols and fatty acids are re-
i. e., they are paracrine hormones (see p. 372).
assembled into fats again (see p. 272), while
While gastrin primarily enhances HCl
the bile acids return to the liver (enterohe-
secretion, cholecystokinin and secretin mainly
patic circulation; see p. 314).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Digestion
271
A. Formation of hydrochloric acid
Interstitial
Parietal cell
Lumen of the stomach
fluid
Peptides
P
H2O
A
Pepsino-
Pepsins
P P
gens
3.4.23.1-3
Native
CO2
CO2
1
protein
K
K
Anion
H
H
exchanger
2
A
H Cl
P P P
≈ 0.1 mol · l-1
HCO3
HCO3
Cl
Cl
Denatured
Cl
Chloride channel
protein
Micro-organism
1
Carbonate dehydratase 4.2.1.1 [Zn2
]
2
H /K
-exchanging ATPase 3.6.1.36
B. Zymogen activation
Small bowel
Mucosal cell Entero-
Pancreatic secretion
peptidase
3.4.21.9
Trypsin
3.4.21.4
Various
zymogens
Trypsinogen
Trypsin inhibitor
Premature
Activated digestive
Zymogens:
trypsin
enzymes
Procarboxypeptidases
activation
Proelastase
Chymotrypsinogen
Prophospholipase A2
Digestion
Trypsinogen
Blocked trypsin
C. Fat digestion
1
Triacylglycerol lipase 3.1.1.3
Enterocyte
1
Micelle
Resorption
C-terminal
domain
1
Monoacyl-
Colipase
glycerol
Bile salt
Phospholipid
Triacylglycerol
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
272
Tissues and organs
acids. Individual amino acid groups have
Resorption
group-specific amino acid transporters, some
Enzymatic hydrolysis in the digestive tract
of which transport the amino acids into the
breaks down foodstuffs into their resorbable
enterocytes in cotransport with Na+ ions (sec-
components. Resorption of the cleavage prod-
ondary active transport), while others trans-
ucts takes place primarily in the small intes-
port them in an Na+-independent manner
tine. Only ethanol and short-chain fatty acids
through facilitated diffusion. Small peptides
are already resorbed to some extent in the
can also be taken up.
stomach.
The resorption process is facilitated by the
B. Lipids
large inner surface of the intestine, with its
brush-border cells. Lipophilic molecules pen-
Fats and other lipids are poorly soluble in
etrate the plasma membrane of the mucosal
water. The larger the accessible surface
cells by simple diffusion, whereas polar mol-
is—i. e., the better the fat is emulsified—the
ecules require transporters (facilitated diffu-
easier it is for enzymes to hydrolyze it (see
sion; see p. 218). In many cases, carrier-medi-
p. 270). Due to the special properties of milk,
ated cotransport with Na+ ions can be ob-
milk fats already reach the gastrointestinal
served. In this case, the difference in the con-
tract in emulsified form. Digestion of them
centration of the sodium ions (high in the
therefore already starts in the oral cavity
intestinal lumen and low in the mucosal cells)
and stomach, where lipases in the saliva and
drives the import of nutrients against a con-
gastric juice are available. Lipids that are less
centration gradient (secondary active trans-
accessible—e. g., from roast pork—are emulsi-
port; see p. 220). Failure of carrier systems in
fied in the small intestine by bile salts and bile
the gastrointestinal tract can result in dis-
phospholipids. Only then are they capable of
eases.
being attacked by pancreatic lipase [4] (see
p. 270).
Fats (triacylglycerols) are mainly attacked
A. Monosaccharides
by pancreatic lipase at positions 1 and 3 of the
The cleavage of polymeric carbohydrates by
glycerol moiety. Cleavage of two fatty acid
a-amylase
[1] leads to oligosaccharides,
residues gives rise to fatty acids and 2-mono-
which are broken down further by exoglyco-
acylglycerols, which are quantitatively the
sidases
(oligosaccharidases and disacchari-
most important products. However, a certain
dases [2]) on the membrane surface of the
amount of glycerol is also formed by complete
brush border. The monosaccharides released
hydrolysis. These cleavage products are re-
in this way then pass with the help of various
sorbed by a non-ATP-dependent process
sugar-specific transporters into the cells of
that has not yet been explained in detail.
intestinal epithelium. Secondary active
In the mucosal cells, long-chain fatty acids
transport serves for the uptake of glucose
are resynthesized by an ATP-dependent ligase
and galactose, which are transported against
[5] to form acyl-CoA and then triacylglycerols
a concentration gradient in cotransport with
(fats; see p. 170). The fats are released into the
Na+. The Na+ gradient is maintained on the
lymph in the form of chylomicrons
(see
basal side of the cells by Na+/K+-ATPase [3].
p. 278) and, bypassing the liver, are deposited
Another passive transporter then releases
in the thoracic duct—i. e., the blood system.
glucose and galactose into the blood.
Cholesterol also follows this route.
Fructose is taken up by a special type of trans-
By contrast, short-chain fatty acids (with
porter using facilitated diffusion.
chain lengths of less than 12 C atoms) pass
directly into the blood and reach the liver via
the portal vein. Resorbed glycerol can also
Amino acids (not illustrated)
take this path.
Protein degradation is initiated by proteina-
ses—by pepsins in the stomach and by trypsin,
chymotrypsin, and elastase in the small intes-
tine. The resulting peptides are then further
hydrolyzed by various peptidases into amino
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Digestion
273
A. Monosaccharides
2 K
3 Na
Poly-
saccharides
Na glucose
2 K
Glucose
3
symporter
ATP
ADP
transporter
P
i
Na
Na
Glucose
Glucose
Galactose
Galactose
1
Portal
*
vein
α-Amylase
Fructose
Fructose
Other mono-
Other mono-
saccharides
saccharides
Liver
1
α-Amylase 3.2.1.1
Disaccharidases
2
Oligosaccharidases
2
Na /K -exchanging
3
ATPase 3.6.1.37
Oligo-
saccharides
Intestinal
Secondary-active transport
epithelial cell
*
Facilitated diffusion
B. Lipids
Intestinal
Triacyl-
lumen
glycerol
Fat
Lymph
synthesis
P
4
Thoracic
Glyco-
duct
Fat
lysis
3x
synthesis
Diacyl-
S
A
glycerol
2x
Blood
A
P
P P
5
Short-chain
fatty acids
A
4
P P P
S
A
Portal
vein
Fatty acids
2-Mono-
acyl-
80%
glycerol
Resorption
4
Liver
Glycerol
20%
4
Triacylglycerol
lipase 3.1.1.3
Intestinal
epithelial cell
5
Fatty acid-CoA ligase 6.2.1.3
Stimulated by bile salts,
phospholipids, colipase and Ca2
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
274
Tissues and organs
The leukocytes include various types of
Blood: composition and functions
granulocyte, monocyte, and lymphocyte. All
Human blood constitutes about 8 % of the
of these have immune defense functions (see
body’s weight. It consists of cells and cell frag-
p. 294). The neutrophil granulocytes¸ mono-
ments in an aqueous medium, the blood
cytes, and the macrophages derived from
plasma. The proportion of cellular elements,
monocytes are phagocytes. They can ingest
known as hematocrit, in the total volume is
and degrade invading pathogens. The lympho-
approximately 45%.
cytes are divided into two groups, B lympho-
cytes and T lymphocytes. B lymphocytes
produce antibodies, while T lymphocytes reg-
A. Functions of the blood
ulate the immune response and destroy virus-
infected cells and tumor cells. Eosinophilic and
The blood is the most important transport
basophilic granulocytes have special tasks for
medium in the body. It serves to keep the
defense against animal parasites.
“internal milieu” constant (homeostasis) and
Thrombocytes are cell fragments that arise
it plays a decisive role in defending the body
in the bone marrow from large precursor
against pathogens.
cells, the megakaryocytes. Their task is to
Transport. The gases oxygen and carbon
promote hemostasis (see p. 290).
dioxide are transported in the blood. The
blood mediates the exchange of substances
between organs and takes up metabolic end
C. Blood plasma: composition
products from tissues in order to transport
The blood plasma is an aqueous solution of
them to the lungs, liver, and kidney for excre-
electrolytes, nutrients, metabolites, proteins,
tion. The blood also distributes hormones
vitamins, trace elements, and signaling sub-
throughout the organism (see p. 370).
stances. The fluid phase of coagulated blood is
Homeostasis. The blood ensures that a bal-
known as blood serum. It differs from the
anced distribution of water is maintained be-
plasma in that it lacks fibrin and other coag-
tween the vascular system, the cells (intra-
ulation proteins (see p. 290).
cellular space), and the extracellular space.
Laboratory assessment of the composition
The acid-base balance is regulated by the
of the blood plasma is often carried out in
blood in combination with the lungs, liver,
clinical chemistry. Among the electrolytes,
and kidneys (see p. 288). The regulation of
there is a relatively high concentration of
body temperature also depends on the con-
Na+, Ca2+, and Cl- ions in the blood in compar-
trolled transport of heat by the blood.
ison with the cytoplasm. By contrast, the con-
Defense. The body uses both non-specific
centrations of K+, Mg2+, and phosphate ions
and specific mechanisms to defend itself
are higher in the cells. Proteins also have a
against pathogens. The defense system in-
higher intracellular concentration. The elec-
cludes the cells of the immune system and
trolyte composition of blood plasma is similar
certain plasma proteins (see p. 294).
to that of seawater, due to the evolution of
Self-protection. To prevent blood loss
early forms of life in the sea. The solution
when a vessel is injured, the blood has sys-
known as “physiological saline” (NaCl at a con-
tems for stanching blood flow and coagulat-
centration of 0.15 mol L-1) is almost isotonic
ing the blood (hemostasis; see p. 290). The
with blood plasma.
dissolution of blood clots
(fibrinolysis) is
A list of particularly important metabolites
also managed by the blood itself (see p. 292).
in the blood plasma is given on the right.
B. Cellular elements
The solid elements in the blood are the eryth-
rocytes (red blood cells), leukocytes (white
blood cells), and thrombocytes (platelets).
The erythrocytes provide for gas transport
in the blood. They are discussed in greater
detail on pp. 280-285.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
275
A. Functions of the blood
B. Cellular elements
Blood gases:
O2
10 µm
CO2
Erythrocyte
5000 · 109 · l-1
Nutrients
59%
6.5%
Metabolites
Metabolic
wastes
Neutrophilic
Monocyte
Hormones
granulocyte
31%
Transport
Small
Large
Cell
lymphocyte
Extracellular
H2O
2.4%
space
0.6%
H
Water balance
OH
Body
Acid-base
temperature
balance
Homeostasis
Eosinophilic
Basophilic
granulocyte
Blood clotting
Leukocytes
7 · 109 · l-1
and fibrinolysis
-1
250 · 109 · l
Immune cells
Antibodies
Defense
Self defense
Thrombocytes
C. Blood plasma: composition
mM
Non-electrolytes
Uncharged
Concentration
Metabolite
Concentration (mM)
molecules
200
1.2
H2CO3
Glucose
3.6
- 6.1
Lactate
0.4
- 1.8
HCO
24-28
150
3
Pyruvate
0.07
- 0.11
Urea
3.5
- 9.0
136-145
Na
Cl
100-110
Uric acid
0.18
- 0.54
100
Creatinin
0.06
- 0.13
2
Amino acids
2.3
- 4.0
HPO
1.1-1.5
4
50
Ammonia
0.02
– 0.06
2
3.5-5.0
K
0.3-0.6
SO4
Lipids (total)
5.5
- 6.0 g · l-1
2.1-2.6
Ca2
Organic
0
acids
Triacylglycerols
1.0
- 1.3 g · l-1
0.6-1.0
Mg2
Proteins
Cholesterol
1.7
- 2.1 g · l
-1
Cations
Anions
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
276
Tissues and organs
as plasma cells (see p. 302) and peptide hor-
Plasma proteins
mones, which derive from endocrine gland
Quantitatively, proteins are the most impor-
cells.
tant part of the soluble components of the
With the exception of albumin, almost all
blood plasma. With concentrations of be-
plasma proteins are glycoproteins. They carry
tween 60 and 80 g L-1, they constitute ap-
oligosaccharides in N-and O-glycosidic bonds
proximately 4% of the body’s total protein.
(see p. 44). N-acetylneuraminic acid
(sialic
Their tasks include transport, regulation of
acid; see p. 38) often occurs as a terminal
the water balance, hemostasis, and defense
carbohydrate among sugar residues.
against pathogens.
Neuraminidases (sialidases) on the surface of
the vascular endothelia gradually cleave the
sialic acid residues and thereby release ga-
A. Plasma proteins
lactose units on the surfaces of the proteins.
Some 100 different proteins occur in human
These asialoglycoproteins (“asialo-” = without
blood plasma. Based on their behavior during
sialic acid) are recognized and bound by gal-
electrophoresis (see below), they are broadly
actose receptors on hepatocytes. In this way,
divided into five fractions: albumins and α1-,
the liver takes up aged plasma proteins by
α2-, β- and γ-globulins. Historically, the dis-
endocytosis and breaks them down. The oli-
tinction between the albumins and globulins
gosaccharides on the protein surfaces thus
was based on differences in the proteins’
determine the half-life of plasma proteins,
solubility
-albumins are soluble in pure
which is a period of days to weeks.
water, whereas globulins only dissolve in
In healthy individuals, the concentration of
the presence of salts.
plasma proteins is constant. Diseases in or-
The most frequent protein in the plasma, at
gans that are involved in protein synthesis
around 45 g L-1, is albumin. Due to its high
and breakdown can shift the protein pattern.
concentration, it plays a crucial role in main-
For example, via cytokines (see p. 392), se-
taining the blood’s colloid osmotic pressure
vere injuries trigger increased synthesis of
and represents an important amino acid re-
acute-phase proteins, which include C-reac-
serve for the body. Albumin has binding sites
tive protein, haptoglobin, fibrinogen, comple-
for apolar substances and therefore functions
ment factor C-3, and others. The concentra-
as a transport protein for long-chain fatty
tions of individual proteins are altered in
acids, bilirubin, drugs, and some steroid hor-
some diseases (known as dysproteinemias).
mones and vitamins. In addition, serum albu-
min binds Ca2+ and Mg2+ ions. It is the only
B. Carrier electrophoresis
important plasma protein that is not glycosy-
lated.
Proteins and other electrically charged mac-
The albumin fraction also includes trans-
romolecules can be separated using electro-
thyretin (prealbumin), which together with
phoresis (see also pp. 78, 262). Among the
other proteins transports the hormone thy-
various procedures used, carrier electropho-
roxine and its metabolites.
resis on cellulose acetate foil (CAF) is partic-
The table also lists important globulins in
ularly simple. Using this method, serum pro-
blood plasma, with their mass and function.
teins—which at slightly alkaline pH values all
The α- and β-globulins are involved in the
move towards the anode, due to their excess
transport of lipids (lipoproteins; see p. 278),
of negative charges—can be separated into
hormones, vitamins, and metal ions. In addi-
the five fractions mentioned. After the pro-
tion, they provide coagulation factors, pro-
teins have been stained with dyes, the result-
tease inhibitors, and the proteins of the com-
ing bands can be quantitatively assessed us-
plement system (see p. 298). Soluble antibod-
ing densitometry.
ies (immunoglobulins; see p. 300) make up
the γ-globulin fraction.
Synthesis and degradation. Most plasma
proteins are synthesized by the liver. Excep-
tions to this include the immunoglobulins,
which are secreted by B lymphocytes known
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
277
A. Plasma proteins
Group
Protein
Mr
in kDa
Function
Albumins:
Transthyretin
50-66
Transport of thyroxin and triiodothyronin
Albumin: 45 g · l-1
67
Maintenance of osmotic pressure; transport of
fatty acids, bilirubin, bile acids, steroid hor-
mones, pharmaceuticals and inorganic ions.
Antitrypsin
51
Inhibition of trypsin and other proteases
α1-Globulins:
Antichymotrypsin
58-68
Inhibition of chymotrypsin
Lipoprotein (HDL)
200-400
Transport of lipids
Prothrombin
72
Coagulation factor II, thrombin precursor
(3.4.21.5)
Transcortin
51
Transport of cortisol, corticosterone and
progesterone
Acid glycoprotein
44
Transport of progesterone
Thyroxin-binding globulin
54
Transport of iodothyronins
Ceruloplasmin
135
Transport of copper ions
α2-Globulins:
Antithrombin III
58
Inhibition of blood clotting
Haptoglobin
100
Binding of hemoglobin
Cholinesterase (3.1.1.8)
ca. 350
Cleavage of choline esters
Plasminogen
90
Precursor of plasmin (3.4.21.7), breakdown
of blood clots
Macroglobulin
725
Binding of proteases, transport of zinc ions
Retinol-binding protein
21
Transport of vitamin A
Vitamin D-binding protein
52
Transport of calciols
β-Globulins:
Lipoprotein (LDL)
2.000-4.500
Transport of lipids
Transferrin
80
Transport of iron ions
Fibrinogen
340
Coagulation factor I
Sex hormone-
binding globulin
65
Transport of testosterone and estradiol
Transcobalamin
38
Transport of vitamin B12
C-reactive protein
110
Complement activation
γ-Globulins:
IgG
150
Late antibodies
IgA
162
Mucosa-protecting antibodies
IgM
900
Early antibodies
IgD
172
B-lymphocyte receptors
IgE
196
Reagins
B. Electrophoresis
Buffer-saturated strip
Cellulose-acetate sheet
of filter paper
soaked with buffer
+
–
52-58%
Serum
+
sample
-
2.4-4.4%
Anode
Cathode
6.1-10.1%
Electro-
phoresis
8.5-14.5%
10-21%
Staining
Densitometry
Cellulose-acetate sheet
Albumins α1- α2- β- γ-Globulins
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
278
Tissues and organs
cosa and reach the blood via the lymphatic
Lipoproteins
system (see p. 266). In the peripheral vessel-
Most lipids are barely soluble in water, and
s—particularly in muscle and adipose tis-
many have amphipathic properties. In the
sue—lipoprotein lipase [1] on the surface of
blood, free triacylglycerols would coalesce
the vascular endothelia hydrolyzes most of
into drops that could cause fat embolisms.
the triacylglycerols. Chylomicron breakdown
By contrast, amphipathic lipids would be de-
is activated by the transfer of apoproteins E
posited in the blood cells’ membranes and
and C from HDL. While the fatty acids released
would dissolve them. Special precautions are
and the glycerol are taken up by the cells, the
therefore needed for lipid transport in the
chylomicrons gradually become converted
blood. While long-chain fatty acids are bound
into chylomicron remnants, which are ulti-
to albumin and short-chain ones are dissolved
mately removed from the blood by the liver.
in the plasma (see p. 276), other lipids are
transported in lipoprotein complexes, of
VLDLs, IDLs, and LDLs are closely related to
which there several types in the blood
one another. VLDLs formed in the liver (see
plasma, with different sizes and composition.
p. 312) transport triacylglycerols, cholesterol,
and phospholipids to other tissues. Like chy-
lomicrons, they are gradually converted into
A. Composition of lipoprotein complexes
IDL and LDL under the influence of lipoprotein
Lipoproteins are spherical or discoid aggre-
lipase [1]. This process is also stimulated by
gates of lipids and apoproteins. They consist
HDL. Cells that have a demand for cholesterol
of a nucleus of apolar lipids (triacylglycerols
bind LDL through an interaction between
and cholesterol esters) surrounded by a sin-
their LDL receptor and ApoB-100, and then
gle-layered shell approximately 2 nm thick of
take up the complete particle through recep-
amphipathic lipids (phospholipids and cho-
tor-mediated endocytosis. This type of trans-
lesterol; the example shown here is LDL).
port is mediated by depressions in the mem-
The shell, in which the apoproteins are also
brane (“coated pits”), the interior of which is
deposited, gives the surfaces of the particles
lined with the protein clathrin. After LDL
polar properties and thereby prevents them
binding, clathrin promotes invagination of
from aggregating into large particles. The
the pits and pinching off of vesicles (“coated
larger the lipid nucleus of a lipoprotein
vesicles”). The clathrin then dissociates off and
is—i. e., the larger the number of apolar lipids
is reused. After fusion of the vesicle with ly-
it contains—the lower its density is.
sosomes, the LDL particles are broken down
Lipoproteins are classified into five groups.
(see p. 234), and cholesterol and other lipids
In order of decreasing size and increasing
are used by the cells.
density, these are: chylomicrons, VLDLs
(very-low-density lipoproteins), IDLs (inter-
The HDLs also originate in the liver. They
mediate-density lipoproteins), LDLs
(low-
return the excess cholesterol formed in the
density lipoproteins), and HDLs (high-density
tissues to the liver. While it is being trans-
lipoproteins). The proportions of apoproteins
ported, cholesterol is acylated by lecithin cho-
range from 1% in chylomicrons to over 50% in
lesterol acyltransferase (LCAT). The cholesterol
HDLs. These proteins serve less for solubility
esters formed are no longer amphipathic and
purposes, but rather function as recognition
can be transported in the core of the lipopro-
molecules for the membrane receptors and
teins. In addition, HDLs promote chylomicron
enzymes that are involved in lipid exchange.
and VLDL turnover by exchanging lipids and
apoproteins with them (see above).
B. Transport functions
The classes of lipoproteins differ not only in
their composition, but also in the ways in
which they originate and function.
The chylomicrons take care of the transport
of triacylglycerols from the intestine to the
tissues. They are formed in the intestinal mu-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
279
A. Composition of lipoprotein complexes
Lipoprotein
Lipoprotein
Composition
coat
core
10 nm
VLDL
Chylomicrons
B-100, C-III, E, C-II
C-III, B-48, C-II,
C-II, E, A-I
LDL
B-100
Apo-B-100
Characteristic
apoproteins
IDL
B-100, E, C-III
HDL
A-I, A-III, C-III
Apoprotein
Lipoprotein
Density (g · cm-3)
Free cholesterol
Chylomicrons
<0.95
VLDL
0.950
- 1.006
Phospholipid
IDL
1.006
- 1.019
Cholesterol ester
LDL
1.019
- 1.063
Triacylglycerols
HDL
1.063
- 1.210
B. Transport functions
Intestine
Dietary lipids
Endogenous lipids
Extrahepatic
tissues
Receptor-
Liver
*
mediated
*
*
endocytosis
Cholesterol
Cholesterol
LDL
*
Fats
*
*
Fatty acids
Chylomicrons
Cholesterol
2
return
HDL
IDL
Chylomicron
remnants
1
HDL
VLDL
1
Muscle
Free
Exchange
fatty
+
of lipids and
acids
Adipose tissue
apoproteins
Free
Free
HDL
fatty
fatty
acids
acids
Free
fatty
Lipoprotein lipase
Lecithin-cholesterol acyltransferase
acids
1
2
3.1.1.34
(LCAT) 2.3.1.43
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
280
Tissues and organs
the R form. The T form (for tense; left) and
Hemoglobin
has a much lower O2 af nity than the R form
The most important task of the red blood cells
(for relaxed; right).
(erythrocytes) is to transport molecular oxy-
Binding of O2 to one of the subunits of the T
gen (O2) from the lungs into the tissues, and
form leads to a local conformational change
carbon dioxide (CO2) from the tissues back
that weakens the association between the
into the lungs. To achieve this, the higher
subunits. Increasing O2 partial pressure thus
organisms require a special transport system,
means that more and more molecules convert
since O2 is poorly soluble in water. For exam-
to the higher-af nity R form. This coopera-
ple, only around 3.2 mL O2 is soluble in 1 L
tive interaction between the subunits in-
blood plasma. By contrast, the protein hemo-
creases the O2 af nity of Hb with increasing
globin (Hb), contained in the erythrocytes,
O2
concentrations—i. e., the O2
saturation
can bind a maximum of 220 mL O2
per
curve is sigmoidal (see p. 282).
liter—70 times the physically soluble amount.
Various allosteric effectors influence the
The Hb content of blood, at 140-180 g L-1
equilibrium between the T and R forms and
in men and 120-160 g L-1 in women, is
thereby regulate the O2 binding behavior of
twice as high as that of the plasma proteins
hemoglobin (yellow arrows). The most impor-
(50-80 g L-1). Hb is therefore also responsi-
tant effectors are CO2, H+, and 2,3-bisphospho-
ble for the majority of the blood proteins’ pH
glycerate (see p. 282).
buffer capacity (see p. 288).
Further information
A. Hemoglobin: structure
As mentioned above, hemoglobin in adults
In adults, hemoglobin (HbA; see below) is a
consists of two α- and two β-chains. In addi-
heterotetramer consisting of two α-chains and
tion to this main form (HbA1, α2β2), adult
two β-chains, each with masses of 16 kDa.
blood also contains small amounts of a second
The α- and β-chains have different sequences,
form with a higher O2 af nity in which the β-
but are similarly folded. Some 80% of the
chains are replaced by δ-chains (HbA2, α2δ2).
amino acid residues form D-helices, which
Two other forms occur during embryonic and
are identified using the letters A-H.
fetal development. In the first three months,
Each subunit carries a heme group (for-
embryonic hemoglobins are formed, with the
mula on p. 106), with a central bivalent iron
structure ζ2ε2 and α2ε2. Up to the time of
ion. When O2
binds to the heme iron
birth, fetal hemoglobin then predominates
(Oxygenation of Hb) and when O2 is released
(HbF, α2γ2), and it is gradually replaced by
(Deoxygenation), the oxidation stage of the
HbA during the first few months of life. Em-
iron does not change. Oxidation of Fe2+ to
bryonic and fetal hemoglobins have higher O2
Fe3+ only occurs occasionally. The oxidized
af nities than HbA, as they have to take up
form, methemoglobin, is then no longer able
oxygen from the maternal circulation.
to bind O2. The proportion of Met-Hb is kept
low by reduction (see p. 284) and usually
amounts to only 1-2%.
Four of the six coordination sites of the iron
in hemoglobin are occupied by the nitrogen
atoms of the pyrrol rings, and another is oc-
cupied by a histidine residue of the globin
(the proximal histidine). The iron’s sixth site
is coordinated with oxygen in oxyhemoglobin
and with H2O in deoxyhemoglobin.
B. Hemoglobin: allosteric effects
Like aspartate carbamoyltransferase
(see
p. 116), Hb can exist in two different states
(conformations), known as the T form and
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
281
A. Hemoglobin: structure
NH3
NH3
Proximal
Distal
β
β
histidine
O2
histidine
COO
A
E
F
H
B
D
G
C
B
G
H
D C
E
A
F
α
α
NH3
COO
F helix
Heme
E helix
Hemoglobin A
(α2 β2) M: 65 kDa
B. Hemoglobin: allosteric effects
pO2
pCO2
, pH
[BPG]
4
O2
HbT ·
BPG
HbR · (O2)4
H
BPG
T
R
pO2
pCO2
, pH
[BPG]
α
α
α
α
β
β
β
β
O2 bond
BPG
weakens
stabilizes
association
T form
O2 affinity 70
times higher
than in T form
Deoxyhemoglobin
Oxyhemoglobin
T form
R form
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
282
Tissues and organs
B. Hemoglobin and CO2 transport
Gas transport
Hemoglobin is also decisively involved in the
Most tissues are constantly dependent on a
transport of carbon dioxide (CO2) from the
supply of molecular oxygen (O2) to maintain
tissues to the lungs.
their oxidative metabolism. Due to its poor
Some 5% of the CO2 arising in the tissues is
solubility, O2
is bound to hemoglobin for
covalently bound to the N terminus of hemo-
transport in the blood (see p. 280). This not
globin and transported as carbaminohemoglo-
only increases the oxygen transport capacity,
bin (not shown). About 90% of the CO2 is first
but also allows regulation of O2 uptake in the
converted in the periphery into hydrogen car-
lungs and O2 release into tissues.
bonate (HCO3-), which is more soluble (bot-
tom). In the lungs (top), CO2 is regenerated
A. Regulation of O2 transport
again from HCO3- and can then be exhaled.
These two processes are coupled to the
When an enzyme reacts to effectors (sub-
oxygenation and deoxygenation of Hb.
strates, activators, or inhibitors) with confor-
Deoxy-Hb is a stronger base than oxy-Hb. It
mational changes that increase or reduce its
therefore binds additional protons
(about
activity, it is said to show allosteric behavior
0.7 H+ per tetramer), which promotes the for-
(see p. 116). Allosteric enzymes are usually
mation of HCO3- from CO2 in the peripheral
oligomers with several subunits that mutually
tissues. The resulting HCO3- is released into
influence each other.
the plasma via an antiporter in the erythro-
Although hemoglobin is not an enzyme (it
cyte membrane in exchange for Cl-, and
releases the bound oxygen without changing
passes from the plasma to the lungs. In the
it), it has all the characteristics of an allosteric
lungs, the reactions described above then
protein. Its effectors include oxygen, which as
proceed in reverse order: deoxy-Hb is oxy-
a positive homotropic effector promotes its
genated and releases protons. The protons
own binding. The O2 saturation curve of he-
shift the HCO3/CO2 equilibrium to the left
moglobin is therefore markedly sigmoidal in
and thereby promote CO2 release.
shape (2, curve 2). The non-sigmoidal satura-
O2 binding to Hb is regulated by H+ ions
tion curve of the muscular protein myoglobin
(i. e., by the pH value) via the same mecha-
is shown for comparison (curve 1). The struc-
nism. High concentrations of CO2 such as
ture of myoglobin (see p. 336) is similar to
those in tissues with intensive metabolism
that of a subunit of hemoglobin, but as a
locally increase the H+ concentration and
monomer it does not exhibit any allosteric
thereby reduce hemoglobin’s O2
af nity
behavior.
(Bohr effect; see above). This leads to in-
CO2, H+, and a special metabolite of ery-
creased O2 release and thus to an improved
throcytes—2,3-bisphosphoglycerate
(BPG)—
oxygen supply.
act as heterotropic effectors of hemoglobin.
The adjustment of the equilibrium be-
BPG is synthesized from 1,3-bisphosphogly-
tween CO2 and HCO3- is relatively slow in
cerate, an intermediate of glycolysis
(see
the uncatalyzed state. It is therefore acceler-
p. 150), and it can be returned to glycolysis
ated in the erythrocytes by carbonate dehy-
again by breakdown into 2-phosphoglycerate
dratase (carbonic anhydrase) [1])—an enzyme
(1), with loss of an ATP.
that occurs in high concentrations in the
BPG binds selectively to deoxy-Hb, thereby
erythrocytes.
increasing its amount of equilibrium. The re-
sult is increased O2 release at constant pΟ2. In
the diagram, this corresponds to a right shift
of the saturation curve (2, curve 3). CO2 and
H+ act in the same direction as BPG. Their
influence on the position of the curve has
long been known as the Bohr effect.
The effects of CO2 and BPG are additive. In
the presence of both effectors, the saturation
curve of isolated Hb is similar to that of whole
blood (curve 4).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
283
A. Regulation of O2 transport
Bisphosphoglycerate
1.0
1
mutase 5.4.2.4
Bisphosphoglycerate
1,3-Bisphospho-
2
phosphatase3.1.3.13
0.8
glycerate
1
ADP
2,3-
0.6
1
2
3
4
ATP
Bisphospho-
glycerate (BPG)
pH = 7.40
3-Phospho-
glycerate
2
0.4
O
O
1
Myoglobin
C
2
Hemoglobin (Hb)
0.2
3
Hb + BPG
P
H
C O
P
4
Hb + CO2 + BPG
2-Phospho-
glycerate
H2C
O
P
0.0
0
10
20
30
40
50
O2 partial pressure (mmHg)
1. BPG metabolism
2. Saturation curves
D. Hemoglobin and CO2 transport
Lungs
CO2
O2
CO
O2
2
CO2
O2
Lungs
CO2
+
H2O
[H2CO3]
1
HCO3
HCO3
+
H - Hb
Hb -O2
+
[H2CO3]
Cl
Cl
pO2
100 mmHg
Blood
Erythrocyte
Heart
Cl
Cl
HCO3
HCO3
+ H - Hb
Hb - O2
+
[H2CO3]
pO2
CO2
+
H2O
1
[H2CO3]
30-40 mmHg
CO2
O2
Veins
Arteries
Tissue
CO2
O2
1
Carbonate dehydratase [Zn2
] 4.2.1.1
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
284
Tissues and organs
antioxidants include vitamins C and E (see
Erythrocyte metabolism
pp. 364, 368), coenzyme Q (see p. 104), and
Cells living in aerobic conditions are depen-
several carotenoids (see pp. 132, 364). Biliru-
dent on molecular oxygen for energy produc-
bin, which is formed during heme degrada-
tion. On the other hand, O2 constantly gives
tion (see p. 194), also serves for protection
rise to small quantities of toxic substances
against oxidation.
known as reactive oxygen species
(ROS).
Glutathione, a tripeptide that occurs in
These substances are powerful oxidation
high concentrations in almost all cells, is par-
agents or extremely reactive free radicals
ticularly important. Glutathione (sequence:
(see p. 32), which damage cellular structures
Glu-Cys-Gly) contains an atypical γ-peptide
and functional molecules. Due to their role in
bond between Glu and Cys. The thiol group of
O2 transport, the erythrocytes are constantly
the cysteine residue is redox-active. Two mol-
exposed to high concentrations of O2 and are
ecules of the reduced form (GSH, top) are
therefore particularly at risk from ROS.
bound to the disulfide (GSSG, bottom) during
oxidation.
A. Reactive oxygen species
C. Erythrocyte metabolism
The dioxygen molecule (O2) contains two un-
paired electrons—i. e., it is a diradical. Despite
Erythrocytes also have systems that can in-
this, O2 is relatively stable due to its special
activate ROS (superoxide dismutase, catalase,
electron arrangement. However, if the mole-
GSH). They are also able to repair damage
cule takes up an extra electron (a), the highly
caused by ROS. This requires products that
reactive superoxide radical ( O2-) arises. An-
are supplied by the erythrocytes’ mainte-
other reduction step (b) leads to the peroxide
nance metabolism, which basically only in-
anion (O22-), which easily binds protons and
volves anaerobic glycolysis (see p. 150) and
thus becomes hydrogen peroxide (H2O2). In-
the pentose phosphate pathway (PPP; see
clusion of a third electron (c) leads to cleavage
p. 152).
of the molecule into the ions O2- and O-.
The ATP formed during glycolysis serves
While O2- can form water by taking up two
mainly to supply Na+/K+-ATPase, which main-
protons, protonation of O- provides the ex-
tains the erythrocytes’ membrane potential.
tremely dangerous hydroxy radical ( OH). A
The allosteric effector 2,3-BPG (see p. 282) is
fourth electron transfer and subsequent pro-
also derived from glycolysis. The PPP supplies
tonation also convert O- into water.
NADPH+H+, which is needed to regenerate
The synthesis of ROS can be catalyzed by
glutathione (GSH) from GSSG with the help
iron ions, for example. Reaction of O2 with
of glutathione reductase [3]. GSH, the most
FMN or FAD (see p. 32) also constantly pro-
important antioxidant in the erythrocytes,
duces ROS. By contrast, reduction of O2 by
serves as a coenzyme for glutathione peroxi-
cytochrome c-oxidase (see p. 140) is “clean,”
dase [5]. This selenium-containing enzyme
as the enzyme does not release the intermedi-
detoxifies H2O2 and hydroperoxides, which
ates. In addition to antioxidants (B), enzymes
arise during the reaction of ROS with unsatu-
also provide protection against ROS: superox-
rated fatty acids in the erythrocyte mem-
ide dismutase [1] breaks down (“dispropor-
brane. The reduction of methemoglobin
tionates”) two superoxide molecules into O2
(Hb Fe3+) to Hb (Hb Fe2+, [4]) is carried out
and the less damaging H2O2. The latter is in
by GSH or ascorbate by a non-enzymatic
turn disproportionated into O2 and H2O by
pathway; however, there are also NAD(P)H-
heme-containing catalase [2].
dependent Met-Hb reductases.
B. Biological antioxidants
To protect them against ROS and other radi-
cals, all cells contain antioxidants. These are
reducing agents that react easily with oxida-
tive substances and thus protect more impor-
tant molecules from oxidation. Biological
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
285
A. Reactive oxygen species
B. Biological antioxidants
Quinols
α-Tocopherol (vitamin E)
Molecular
O2
and enols
Ubiquinol (coenzyme Q)
oxygen
Ascorbic acid (vitamin C)
e
Carotenoids
β-Carotin
a
Dispro-
Lycopin
portionation
Others
Glutathione
Bilirubin
Superoxide
O2
1
radical
1. Examples
Glu
Cys
Gly
e
b
H
O
H O
Hydrogen
peroxide
OOC
C
(CH2)2
C N
C C N CH2
COO
NH3
H
CH2
H
2
O2
H2O2
2
2 Glutathione
SH
2 (GSH)
Hydroxy
e
2H
Oxidation
Reduction
radical
c
H
O
H O
2
OH
O
O
H2O
OOC
C
(CH2)2
C N
C C
N CH2
COO
NH
H
CH
H
3
2
d
Water
S
Glutathione
H
2H
disulfide (GSSG)
S
e
NH
H
CH2
H
3
OOC
C
(CH2)2
C N
C C
N CH2
COO
H
O
H O
1
Superoxide dismutase
2
Catalase
1.15.1.1
1.11.1.6
2. Glutathione
C. Erythrocyte metabolism
Glutathione reductase
Methemoglobin
Glutathione peroxidase
3
4
5
[FAD] 1.6.4.2
reductase
[Se] 1.11.1.9/12
Glucose 6-
Pentose phosphate pathway
Glucose
phosphate
Glucose
N
A
N
A
N
A
N
A
P
P
P
P
Glycolysis
A
P P
3
3
3 Na
P
2 K
2 GSH
GSSG
2 GSH
GSSG
Na /K -
A
O2
ATPase
P P P
BPG
4
5
2 Lactate
R-O-O-R'
R -OH
2
3+
2
2+
2 Lactate
R'-OH
Peroxide
Hb
Met-Hb
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
286
Tissues and organs
very tightly. Similar iron transport proteins
Iron metabolism
are found in secretions such as saliva, tears,
and milk; these are known as lactoferrins
A. Distribution of iron
(bottom right). Transferrin and the lactofer-
Iron (Fe) is quantitatively the most important
rins maintain the concentration of free iron in
trace element (see p. 362). The human body
body fluids at values below 10-10 mol L-1.
contains 4-5 g iron, which is almost exclusively
This low level prevents bacteria that require
present in protein-bound form. Approximately
free iron as an essential growth factor from
three-quarters of the total amount is found in
proliferating in the body. Like LDLs
(see
heme proteins (see pp. 106, 192), mainly he-
p. 278), transferrin and the lactoferrins are
moglobin and myoglobin. About 1% of the
taken up into cells by receptor-mediated
iron is bound in iron-sulfur clusters
(see
endocytosis.
p. 106), which function as cofactors in the
Excess iron is incorporated into ferritin and
respiratory chain, in photosynthesis, and in
stored in this form in the liver and other
other redox chains. The remainder consists
organs. The ferritin molecule consists of 24
of iron in transport and storage proteins
subunits and has the shape of a hollow sphere
(transferrin, ferritin; see B).
(bottom left). It takes up Fe2+ ions, which in
the process are oxidized to Fe3+ and then
deposited in the interior of the sphere as fer-
B. Iron metabolism
rihydrate. Each ferritin molecule is capable of
Iron can only be resorbed by the bowel in
storing several thousand iron ions in this way.
bivalent form (i. e., as Fe2+). For this reason,
In addition to ferritin, there is another storage
reducing agents in food such as ascorbate
form, hemosiderin, the function of which is
(vitamin C; see p. 368) promote iron uptake.
not yet clear.
Via transporters on the luminal and basal side
of the enterocytes, Fe2+ enters the blood,
Further information
where it is bound by transferrin. Part of the
iron that is taken up is stored in the bowel in
Disturbances of the iron metabolism are fre-
the form of ferritin (see below). Heme groups
quent and can lead to severe disease pictures.
can also be resorbed by the small intestine.
Iron deficiency is usually due to blood loss,
Most of the resorbed iron serves for the
or more rarely to inadequate iron uptake.
formation of red blood cells in the bone mar-
During pregnancy, increased demand can
row (erythropoiesis, top). As discussed on
also cause iron deficiency states. In severe
p. 192, it is only in the final step of hem bio-
cases, reduced hemoglobin synthesis can
synthesis that Fe2+ is incorporated by ferro-
lead to anemia (“iron-deficiency anemia”). In
chelatase into the previously prepared tetra-
these patients, the erythrocytes are smaller
pyrrol framework.
and have less hemoglobin. As their membrane
In the blood, 2.5-3.0 g of hemoglobin iron
is also altered, they are prematurely elimi-
circulates as a component of the erythrocytes
nated in the spleen.
(top right). Over the course of several months,
Disturbances resulting from raised iron
the flexibility of the red blood cells constantly
concentrations are less frequent. Known as
declines due to damage to the membrane and
hemochromatoses, these conditions can
cytoskeleton. Old erythrocytes of this type are
have genetic causes, or may be due to re-
taken up by macrophages in the spleen and
peated administration of blood transfusions.
other organs and broken down. The organic
As the body has practically no means of ex-
part of the heme is oxidized into bilirubin (see
creting iron, more and more stored iron is
p. 194), while the iron returns to the plasma
deposited in the organs over time in patients
pool. The quantity of heme iron recycled per
with untreated hemochromatosis, ultimately
day is much larger than the amount resorbed
leading to severe disturbances of organ func-
by the intestines.
tion.
Transferrin, a β-globulin with a mass of
80 kDa, serves to transport iron in the blood.
This monomeric protein consists of two sim-
ilar domains, each of which binds an Fe2+ ion
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
287
A. Distribution of iron
Heme enzyme (<1%)
O2
Myo-
Transport and
globin
storage forms
(6%)
(26%)
Hemo-
Iron-sulfur
Oxygenated heme
globin
clusters
Fe4S4-Cluster
(66%)
(< 1%)
1. Heme iron
2. Non-heme iron
B. Iron metabolism
120 - 150 mg
Free iron
Heme
Transferrin
Erythro-
Ferritin
poiesis
E.g.,
Hemosiderin
ascorbate
Bone
Fe3+
marrow
XH
2
Store
Fe2+
X
2500 - 3000 mg
Fe2+
Erythro-
Fe2+
Heme
cytes
Intestine
Trans-
4 mg
Ferritin
ferrin
Erythro-
cyte
degra-
dation
Blood
Concentration
Fe2+
serum
of free iron in
the blood
< 10-10 mol . l -1
Bilirubin
Spleen
Insoluble
Ferritin
Hemosiderin
150 - 200 mg
Storage tissue
Apoferritin (section)
Lactoferrin
(liver, etc.)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
288
Tissues and organs
the urine, the H+ ions are buffered by NH3 and
Acid-base balance
phosphate.
A. Hydrogen ion concentration in the blood
plasma
C. Buffer systems in the plasma
The H+ concentration in the blood and ex-
The buffering capacity of a buffer system de-
tracellular space is approximately
40 nM
pends on its concentration and its pKa value.
(4
10-8 mol L-1). This corresponds to a pH
The strongest effect is achieved if the pH
of 7.40. The body tries to keep this value con-
value corresponds to the buffer system’s pKa
stant, as large shifts in pH are incompatible
value (see p. 30). For this reason, weak acids
with life.
with pKa values of around 7 are best suited for
The pH value is kept constant by buffer
buffering purposes in the blood.
systems that cushion minor disturbances in
The most important buffer in the blood is
the acid-base balance (C). In the longer term,
the CO2/bicarbonate buffer. This consists of
the decisive aspect is maintaining a balanced
water, carbon dioxide (CO2, the anhydride of
equilibrium between H+ production and up-
carbonic acid H2CO3), and hydrogen carbo-
take and H+ release. If the blood’s buffering
nate (HCO3-, bicarbonate). The adjustment
capacity is not suf cient, or if the acid-base
of the balance between CO2 and HCO3- is
balance is not in equilibrium—e. g., in kidney
accelerated by the zinc-containing enzyme
disease or during hypoventilation or hyper-
carbonate dehydratase
(carbonic anhydrase
ventilation—shifts in the plasma pH value
[1]; see also p. 282). At the pH value of the
can occur. A reduction by more than 0.03
plasma, HCO3- and CO2 are present in a ratio
units is known as acidosis, and an increase is
of about 20 : 1. However, the CO2 in solution
called alkalosis.
in the blood is in equilibrium with the gaseous
CO2 in the pulmonary alveoli. The CO2/HCO3-
system is therefore a powerful open buffer
B. Acid-base balance
system, despite having a not entirely optimal
Protons are mainly derived from two sour-
pKa value of 6.1. Faster or slower respiration
ces—free acids in the diet and sulfur-contain-
increases or reduces CO2 release in the lungs.
ing amino acids. Acids taken up with food—
This shifts the CO2/HCO3- ratio and thus the
e. g., citric acid, ascorbic acid, and phosphoric
plasma pH value (respiratory acidosis or alka-
acid—already release protons in the alkaline
losis). In this way, respiration can compensate
pH of the intestinal tract. More important for
to a certain extent for changes in plasma pH
proton balance, however, are the amino acids
values. However, it does not lead to the ex-
methionine and cysteine, which arise from
cretion of protons.
protein degradation in the cells. Their S atoms
Due to their high concentration, plasma
are oxidized in the liver to form sulfuric acid,
proteins—and hemoglobin in the erythro-
which supplies protons by dissociation into
cytes in particular—provide about one-quar-
sulfate.
ter of the blood’s buffering capacity. The buf-
During anaerobic glycolysis in the muscles
fering effect of proteins involves contribu-
and erythrocytes, glucose is converted into
tions from all of the ionizable side chains. At
lactate, releasing protons in the process (see
the pH value of blood, the acidic amino acids
p. 338). The synthesis of the ketone bodies
(Asp, Glu) and histidine are particularly effec-
acetoacetic acid and 3-hydroxybutyric acid
tive.
in the liver (see p. 312) also releases protons.
The second dissociation step in phosphate
Normally, the amounts formed are small and
(H2PO4/HPO42-) also contributes to the buf-
of little influence on the proton balance. If
fering capacity of the blood plasma. Although
acids are formed in large amounts, however
the pKa value of this system is nearly optimal,
(e. g., during starvation or in diabetes mellitus;
its contribution remains small due to the low
see p. 160), they strain the buffer systems and
total concentration of phosphate in the blood
can lead to a reduction in pH (metabolic
(around 1 mM).
acidoses; lactacidosis or ketoacidosis).
Only the kidney is capable of excreting pro-
tons in exchange for Na+ ions (see p. 326). In
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
289
A. Hydrogen ion concentration in the blood plasma
pH value of the plasma
nM
60
50
40
30
20
Logarithmic scale
[H
]
Linear scale
pH
7.2
7.4
7.6
Acidosis
Alkalosis
Hypoventilation, increased production,
Hyperventilation
decreased excretion of H
increased H
excretion
B. Acid-base balance
Proteins
Thiol groups
Fatty acids
Glucose
Acids
H2 SO4
Ketone bodies
H + anions
2H
+ SO42
H + anions
H + lactate
CO2
+ H2O
HCO3
+
H
Only overproduction leads to acidosis
Lung
Lung influences
pH value by CO2
excretion
Urine (
/day)
~60 mmol H
C. Buffer systems in the plasma
Carbon dioxide-
CO2
+ H2O
1
H2 CO3
H + HCO3
pKa = 6.1
bicarbonate buffer
1.2 mM
24 mM
Prot · H
Prot + H
Protein buffer
pKa = 4 - 12
200 - 240 g Protein · l-1
HCO3 /CO2 + H2O
75%
Prot/Prot· H
24%
2
H2PO4
HPO
+ H
Phosphate buffer
4
pKa = 6.8
1 mM
HPO
1%
42
/H2PO4
Buffering capacity
1
Carbonate dehydratase 4.2.1.1
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
290
Tissues and organs
leads in five steps via factors XIIa, XIa, IXa,
Blood clotting
and Xa to the activation of prothrombin. The
Following injury to blood vessels, hemostasis
significance of this pathway in vivo has been
ensures that blood loss is minimized. Initially,
controversial since it was found that a genetic
thrombocyte activation leads to contraction
deficiency in factor XII does not lead to coag-
of the injured vessel and the formation of
ulation disturbances.
a loose clot consisting of thrombocytes
Both pathways depend on the presence of
(hemostasis). Slightly later, the action of the
activated thrombocytes, on the surface of
enzyme thrombin leads to the formation and
which several reactions take place. For exam-
deposition in the thrombus of polymeric fi-
ple, the prothrombinase complex (left) forms
brin (coagulation, blood clotting). The coagu-
when factors Xa and II, with the help of Va,
lation process is discussed here in detail.
bind via Ca2+ ions to anionic phospholipids in
the thrombocyte membrane. For this to hap-
pen, factors II and X have to contain the non-
A. Blood clotting
proteinogenic amino acid J-carboxygluta-
The most important reaction in blood clotting
mate (Gla; see p. 62), which is formed in the
is the conversion, catalyzed by thrombin, of
liver by post-translational carboxylation of
the soluble plasma protein fibrinogen (factor
the factors. The Gla residues are found in
I) into polymeric fibrin, which is deposited as
groups in special domains that create contacts
a fibrous network in the primary thrombus.
to the Ca2+ ions. Factors VII and IX are also
Thrombin (factor IIa) is a serine proteinase
linked to membrane phospholipids via Gla
(see p. 176) that cleaves small peptides from
residues.
fibrinogen. This exposes binding sites that
Substances that bind Ca2+ ions (e. g., citrate)
spontaneously allow the fibrin molecules to
prevent Gla-containing factors from attaching
aggregate into polymers. Subsequent covalent
to the membrane and therefore inhibit
cross-linking of fibrin by a transglutaminase
coagulation. Antagonists of vitamin K, which
(factor XIII) further stabilizes the thrombus.
is needed for synthesis of the Gla residues
Normally, thrombin is present in the blood
(see p. 364) also have anticoagulatory effects.
as an inactive proenzyme (see p. 270). Pro-
These include dicumarol, for example.
thrombin is activated in two different ways,
Active thrombin not only converts fibrino-
both of which represent cascades of enzy-
gen into fibrin, but also indirectly promotes
matic reactions in which inactive proenzymes
its own synthesis by catalyzing the activation
(zymogens, symbol: circle) are proteolytically
of factors V and VIII. In addition, it catalyzes
converted into active proteinases (symbol:
the activation of factor XIII and thereby trig-
sector of a circle). The proteinases activate
gers the cross-linking of the fibrin.
the next proenzyme in turn, and so on. Sev-
Regulation of blood clotting (not shown).
eral steps in the cascade require additional
To prevent the coagulation reaction from be-
protein factors (factors III, Va and VIIIa) as
coming excessive, the blood contains a num-
well as anionic phospholipids (PL; see below)
ber of anticoagulant substances, including
and Ca2+ ions. Both pathways are activated by
highly effective proteinase inhibitors. For
injuries to the vessel wall.
example, antithrombin III binds to various ser-
In the extravascular pathway (right), tissue
ine proteinases in the cascade and thereby
thromboplastin (factor III), a membrane pro-
inactivates them. Heparin, an anticoagulant
tein in the deeper layers of the vascular wall,
glycosaminoglycan (see p. 346), potentiates
activates coagulation factor VII. The activated
the effect of antithrombin III. Thrombomodu-
form of this (VIIa) autocatalytically promotes
lin, which is located on the vascular endothe-
its own synthesis and also generates the ac-
lia, also inactivates thrombin. A glycoprotein
tive factors IXa and Xa from their precursors.
known as Protein C ensures proteolytic deg-
With the aid of factor VIIIa, PL, and Ca2+, factor
radation of factors V and VIII. As it is activated
IXa produces additional Xa, which finally—
by thrombin, coagulation is shut down in this
with the support of Va, PL, and Ca2+—releases
way.
active thrombin.
The intravascular pathway (left) is prob-
ably also triggered by vascular injuries. It
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
291
A. Blood clotting
Kallikrein
Intravascular pathway
Extravascular pathway
?
~3 min
~10 s
Endothelium
Deeper
wall
XII
XIIa
layers
III
Function
unclear
Thromboplastin
XI
XIa
IX
IXa
VIIa
VII
VIII
VIIIa
+
Ca
2
PL
X
Xa
V
Va
+
Ca
2
PL
II
IIa
Thrombin
Fibrinogen
Fibrin
I
Prothrombinase complex
Fibrin
polymer
II
Thrombin
Xa
IIa
XIII
XIIIa
K2
EGF2
K1
EGF1
Va
Gla domain
Ca2
Ca2
Ca2
Ca2
Thrombocyte
Polymeric
membrane
fibrin network
Coagulation factors
l Fibrinogen
lX Christmas factor* 3.4.21.22
ll Prothrombin* 3.4.21.5
X Stuart-Prower factor* 3.4.21.6
lll Tissue factor/thromboplastin
Xl Plasma thromboplastin antecedent* (PTA) 3.4.21.27
lV Ca2
Xll Hageman factor* 3.4.21.38
V Proaccelerin
Xlll Fibrin-stabilizing factor* 2.3.2.13
Vl Synonym for Va
Vll Proconvertin* 3.4.21.21
Proenzyme
*
Vlll Antihemophilic factor A
Contains γ-carboxyglutamate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
292
Tissues and organs
possess. For example, carriers of blood group
Fibrinolysis, blood groups
A form antibodies against antigen B (“anti-B”),
while carriers of group B form antibodies
A. Fibrinolysis
against antigen A (“anti-A”). Individuals with
The fibrin thrombus resulting from blood
blood group 0 form both types, and those
clotting
(see p. 290) is dissolved again by
with blood group AB do not form any of these
plasmin, a serine proteinase found in the
antibodies.
blood plasma. For this purpose, the pre-
If blood from blood group A is transfused
cursor plasminogen first has to be proteolyti-
into the circulation of an individual with
cally activated by enzymes from various tis-
blood group B, for example, then the anti-A
sues. This group includes the plasminogen
present there binds to the A antigens. The
activator from the kidney (urokinase) and tis-
donor erythrocytes marked in this way are
sue plasminogen activator (t-PA) from vascular
recognized and destroyed by the complement
endothelia. By contrast, the plasma protein
system (see p. 298). In the test tube, aggluti-
α2-antiplasmin, which binds to active plasmin
nation of the erythrocytes can be observed
and thereby inactivates it, inhibits fibrinoly-
when donor and recipient blood are incom-
sis.
patible.
Urokinase, t-PA, and streptokinase, a bac-
The recipient’s serum should not contain
terial proteinase with similar activity, are
any antibodies against the donor erythro-
used clinically to dissolve thrombi following
cytes, and the donor serum should not con-
heart attacks. All of these proteins are ex-
tain any antibodies against the recipient’s
pressed recombinantly in bacteria
(see
erythrocytes. Donor blood from blood group
p. 262).
0 is unproblematic, as its erythrocytes do not
possess any antibodies and therefore do not
react with anti-A or anti-B in the recipient’s
B. Blood groups: the ABO system
blood. Conversely, blood from the AB group
During blood transfusions, immune reactions
can only be administered to recipients with
can occur that destroy the erythrocytes trans-
the AB group, as these are the only ones with-
fused from the donor. These reactions result
out antibodies.
from the formation of antibodies (see p. 300)
In the Rh system (not shown), proteins on
directed to certain surface structures on the
the surface of the erythrocytes act as antigens.
erythrocytes. Known as blood group antigens,
These are known as “rhesus factors,” as the
these are proteins or oligosaccharides that can
system was first discovered in rhesus mon-
differ from individual to individual. More than
keys.
20 different blood group systems are now
The rhesus D antigen occurs in 84% of all
known. The ABO system and the Rh system
white individuals, who are therefore “Rh-pos-
are of particular clinical importance.
itive.” If an Rh-positive child is born to an Rh-
In the ABO system, the carbohydrate parts
negative mother, fetal erythrocytes can enter
of glycoproteins or glycolipids act as antigens.
the mother’s circulation during birth and lead
In this relatively simple system, there are four
to the formation of antibodies (IgG) against
blood groups (A, B, AB, and 0). In individuals
the D antigen. This initially has no acute ef-
with blood groups A and B, the antigens con-
fects on the mother or child. Complications
sist of tetrasaccharides that only differ in their
only arise when there is a second pregnancy
terminal sugar (galactose or N-acetylgalactos-
with an Rh-positive child, as maternal anti-D
amine). Carriers of the AB blood group have
antibodies cross the placenta to the fetus even
both antigens (A and B). Blood group 0 arises
before birth and can trigger destruction of the
from an oligosaccharide (the H antigen) that
child’s Rh-positive erythrocytes (fetal erythro-
lacks the terminal residue of antigens A and B.
blastosis).
The molecular causes for the differences be-
tween blood groups are mutations in the gly-
cosyl transferases that transfer the terminal
sugar to the core oligosaccharide.
Antibodies are only formed against anti-
gens that the individual concerned does not
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Blood
293
A. Fibrinolysis
Plasminogen
Plasminogen
Tissue
activator
plasminogen
3.4.21.73
activator
3.4.21.68
Plasmin
Soluble
Fibrin
3.4.21.7
proteins
thrombus
Inactive plasmin
α2-antiplasmin
B. Blood groups: the AB0 system
Blood group
A
B
AB
0
Genotypes
AA and A0
BB and B0
AB
00
A
B
H
H
A
B
A A
B
H
A
Antigens
A
A
B
B
H
H
H
B
B
B
A
B
B
H
H
A
B
A
A
B
A
H
Antibodies
anti-B
anti-A
anti-A
in blood
anti-B
Frequency in
40%
16%
4%
40%
central Europe
Erythrocyte
Oligosaccharide
GP
Glycoprotein with oligosaccharide
N-acetyl-D-glucosamine
Glyco-
lipid
D-Galactose
Membrane
D-Fucose
protein
GP
N-acetyl-D-galactosamine
1
Fucosyltransferase 2.4.1.69/152
N-acetyl-galactosaminyl
2
GDP
transferase 2.4.1.40
3
Galactosyltransferase 2.4.1.37
1
GDP
GP
UDP
UDP
GP
UDP
UDP
GP
3
2
B antigen
H antigen (blood group 0)
A antigen
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
294
Tissues and organs
A. Simplified diagram of the immune
Immune response
response
Viruses, bacteria, fungi, and parasites that en-
Pathogens that have entered the body—e. g.,
ter the body of vertebrates of are recognized
viruses (top)—are taken up by antigen-pre-
and attacked by the immune system. Endog-
senting cells (APCs) and proteolytically de-
enous cells that have undergone alterations—
graded (1). The viral fragments produced in
e. g., tumor cells—are also usually recognized
this way are then presented on the surfaces of
as foreign and destroyed. The immune system
these cells with the help of special membrane
is supported by physiological changes in
proteins (MHC proteins; see p. 296) (2). The
infected tissue, known as inflammation. This
APCs include B lymphocytes, macrophages,
reaction makes it easier for the immune cells
and dendritic cells such as the skin’s Langer-
to reach the site of infection.
hans cells.
Two different systems are involved in the
The complexes of MHC proteins and viral
immune response. The innate immune system
fragments displayed on the APCs are recog-
is based on receptors that can distinguish
nized by T cells that carry a receptor that
between bacterial and viral surface structures
matches the antigen (“T-cell receptors”; see
or foreign proteins (known as antigens) and
p. 296) (3). Binding leads to activation of the T
those that are endogenous. With the help of
cell concerned and selective replication of it
these receptors, phagocytes bind to the patho-
(4, “clonal selection”). The proliferation of im-
gens, absorb them by endocytosis, and break
mune cells is stimulated by interleukins (IL).
them down. The complement system (see
These are a group of more than 20 signaling
p. 298) is also part of the innate system.
substances belonging to the cytokine family
The acquired (adaptive) immune system is
(see p. 392), with the help of which immune
based on the ability of the lymphocytes to
cells communicate with each other. For exam-
form highly specific antigen receptors “on
ple, activated macrophages release IL-1 (5),
suspicion,” without ever having met the cor-
while T cells stimulate their own replication
responding antigen. In humans, there are sev-
and that of other immune cells by releasing
eral billion different lymphocytes, each of
IL-2 (6).
which carries a different antigen receptor. If
Depending on their type, activated T cells
this type of receptor recognizes “its” cognate
have different functions. Cytotoxic T cells
antigen, the lymphocyte carrying it is acti-
(green) are able to recognize and bind virus-
vated and then plays its special role in the
infected body cells or tumor cells (7). They
immune response.
then drive the infected cells into apoptosis
In addition, a distinction is made between
(see p. 396) or kill them with perforin, a pro-
cellular and humoral immune responses. The
tein that perforates the target cell’s plasma
T lymphocytes (T cells) are responsible for cel-
membrane (8).
lular immunity. They are named after the thy-
B lymphocytes, which as APCs present viral
mus, in which the decisive steps in their dif-
fragments on their surfaces, are recognized by
ferentiation take place. Depending on their
helper T cells (blue) or their T cell receptors
function, another distinction is made be-
(9). Stimulated by interleukins, selective clo-
tween cytotoxic T cells (green) and helper T
nal replication then takes place of B cells that
cells
(blue). Humoral immunity is based on
carry antigen receptors matching those of the
the activity of the B lymphocytes (B cells, light
pathogen (10). These mature into plasma cells
brown), which mature in the bone marrow.
(11) and finally secrete large amounts of
After activation by T cells, B cells are able to
soluble antibodies (12).
release soluble forms of their specific antigen
receptors, known as antibodies (see p. 300),
into the blood plasma. The immune system’s
“memory” is represented by memory cells.
These are particularly long-lived cells that
can arise from any of the lymphocyte types
described.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Immune response
295
A. Simplified scheme of the immune response
Virus
MHC protein
(class I)
Antigen-presenting cells
MHC protein
B lymphocyte
Macrophage
(class II)
1
Antibody
(IgM)
2
2
Macrophage
MHC protein
(class I) with
T cell receptor
viral peptide
for MHC-protein
(class II)
T lymphocytes
MHC protein
3
3
(class II) with
virus fragment
Resting
Resting
helper cells
cytotoxic
T cells
10
IL- 1
IL
Activated
5
IL- 1
helper cell
IL- 2
6
IL- 2
11
4
6
4
9
Plasma
Activated
Activated
cell
helper cells
cytotoxic
T cells
12
Any body
cell
8
7
Antibody
(IgG)
Memory cells
IL = interleukin
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
296
Tissues and organs
that any two individuals carry the same set of
T-cell activation
MHC proteins—except for monozygotic twins.
For the selectivity of the immune response
Class I MHC proteins occur in almost all
(see p. 294), the cells involved must be able
nucleated cells. They mainly interact with cy-
to recognize foreign antigens and proteins on
totoxic T cells and are the reason for the re-
other immune cells safely and reliably. To do
jection of transplanted organs. Class I MHC
this, they have antigen receptors on their cell
proteins are heterodimers (αβ). The β subunit
surfaces and co-receptors that support recog-
is also known as β2-microglobulin.
nition.
Class II MHC proteins also consist of two
peptide chains, which are related to each
other. MHC II molecules are found on all anti-
A. Antigen receptors
gen-presenting cells in the immune system.
Many antigen receptors belong to the immu-
They serve for interaction between these cells
noglobulin superfamily. The common charac-
and CD4-carrying T helper cells.
teristic of these proteins is that they are made
up from “immunoglobulin domains.” These
B. T-cell activation
are characteristically folded substructures
consisting of 70-110 amino acids, which are
The illustration shows an interaction between
also found in soluble immunoglobulins (Ig;
a virus-infected body cell (bottom) and a CD8-
see p. 300). The illustration shows schemati-
carrying cytotoxic T lymphocyte (top). The
cally a few of the important proteins in the Ig
infected cell breaks down viral proteins in
superfamily. They consist of constant regions
its cytoplasm (1) and transports the peptide
(brown or green) and variable regions (or-
fragments into the endoplasmic reticulum
ange). Homologous domains are shown in
with the help of a special transporter (TAP)
the same colors in each case. All of the recep-
(2). Newly synthesized class I MHC proteins
tors have transmembrane helices at the C
on the endoplasmic reticulum are loaded
terminus, which anchor them to the mem-
with one of the peptides (3) and then trans-
branes. Intramolecular and intermolecular di-
ferred to the cell surface by vesicular trans-
sulfide bonds are also usually found in pro-
port (4). The viral peptides are bound on the
teins belonging to the Ig family.
surface of the α2 domain of the MHC protein
Immunoglobulin M (IgM), a membrane
in a depression formed by an insertion as a
protein on the surface of B lymphocytes,
“floor” and two helices as “walls” (see smaller
serves to bind free antigens to the B cells. By
illustration).
contrast, T cell receptors only bind antigens
Supported by CD8 and other co-receptors,
when they are presented by another cell as a
a T cell with a matching T cell receptor binds
complex with an MHC protein (see below).
to the MHC peptide complex (5; cf. p. 224).
Interaction between MHC-bound antigens
This binding activates protein kinases in the
and T cell receptors is supported by co-recep-
interior of the T cell, which trigger a chain of
tors. This group includes CD8, a membrane
additional reactions (signal transduction; see
protein that is typical in cytotoxic T cells. T
p. 388). Finally, destruction of the virus-in-
helper cells use CD4 as a co-receptor instead
fected cell by the cytotoxic T lymphocytes
(not shown). The abbreviation “CD” stands for
takes place.
“cluster of differentiation.” It is the term for a
large group of proteins that are all located on
the cell surface and can therefore be identi-
fied by antibodies. In addition to CD4 and
CD8, there are many other co-receptors on
immune cells (not shown).
The MHC proteins are named after the
“major histocompatibility complex”—the DNA
segment that codes for them. Human MHC
proteins are also known as HLA antigens (“hu-
man leukocyte-associated” antigens). Their
polymorphism is so large that it is unlikely
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Immune response
297
A. Antigen receptors
Variable part
H chain
Constant part
L chain
Disulfide bond
Characteristic
domain of the
superfamily
Ig domain
•
•
α Chain
β Chain
α Chain
α Chain
β Chain
•
•
44 kDa
37 kDa
44 kDa
33 kDa
28 kDa
•
•
•
•
•
•
•
α2
•
α1
•
•
•
•
•
•
•
•
β2 Micro-
•
•
•
•
•
•
•
•
•
•
globulin
•
•
•
•
•
•
12 kDa
α3
C terminus
T cell receptor
IgM
MHC protein
MHC protein
CD8
(class I)
(class II)
B. T cell activation
Viral peptide
Cytotoxic T cell
T cell activity
CD8
T cell
receptor
Virus
Presented
MHC
5
viral peptide
protein
(class I)
Endoplasmic
ATP
ADP + P
Vesicular
reticulum
4
transport
1
Proteo-
lysis
3
2
TAP Viral peptides
Virus-infected body cell
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
298
Tissues and organs
The classic pathway is triggered by the for-
Complement system
mation of factor C1 at IgG or IgM on the sur-
The complement system is part of the innate
face of microorganisms (left). C1 is an 18-part
immune system (see p. 294). It supports non-
molecular complex with three different com-
specific defense against microorganisms. The
ponents (C1q, C1r, and C1s). C1q is shaped like
system consists of some 30 different proteins,
a bunch of tulips, the “flowers” of which bind
the “complement factors,” which are found in
to the Fc region of antibodies (left). This acti-
the blood and represent about
4% of all
vates C1r, a serine proteinase that initiates the
plasma proteins there. When inflammatory
cascade of the classic pathway. First, C4 is
reactions occur, the complement factors enter
proteolytically activated into C4b, which in
the infected tissue and take effect there.
turn cleaves C2 into C2a and C2b. C4B and
The complement system works in three
C2a together form C3 convertase [1], which
different ways:
finally catalyzes the cleavage of C3 into C3a
Chemotaxis. Various complement factors
and C3b. Small amounts of C3b also arise from
attract immune cells that can attack and
non-enzymatic hydrolysis of C3.
phagocytose pathogens.
The alternative pathway starts with the
Opsonization. Certain complement factors
binding of factors C3b and B to bacterial lipo-
(“opsonins”) bind to the pathogens and
polysaccharides (endotoxins). The formation
thereby mark them as targets for phagocytos-
of this complex allows cleavage of B by factor
ing cells (e. g., macrophages).
D, giving rise to a second form of C3 conver-
Membrane attack. Other complement fac-
tase (C3bBb).
tors are deposited in the bacterial membrane,
Proteolytic cleavage of factor C3 provides
where they create pores that lyse the patho-
two components with different effects. The
gen (see below).
reaction exposes a highly reactive thioester
group in C3b, which reacts with hydroxyl or
amino groups. This allows C3b to bind cova-
A. Complement activation
lently to molecules on the bacterial surface
The reactions that take place in the comple-
(opsonization, right). In addition, C3b initiates
ment system can be initiated in several ways.
a chain of reactions leading to the formation
During the early phase of infection, lipopoly-
of the membrane attack complex (see below).
saccharides and other structures on the sur-
Together with C4a and C5a (see below), the
face of the pathogens trigger the alternative
smaller product C3a promotes the inflamma-
pathway
(right). If antibodies against the
tory reaction and has chemotactic effects.
pathogens become available later, the anti-
The “late” factors C5 to C9 are responsible
gen-antibody complexes formed activate the
for the development of the membrane attack
classic pathway (left). Acute-phase proteins
complex (bottom). They create an ion-perme-
(see p. 276) are also able to start the comple-
able pore in the bacterial membrane, which
ment cascade (lectin pathway, not shown).
leads to lysis of the pathogen. This reaction is
Factors C1 to C4 (for “complement”) belong
triggered by C5 convertase [2]. Depending on
to the classic pathway, while factors B and D
the type of complement activation, this en-
form the reactive components of the alterna-
zyme has the structure C4b2a3b or C3bBb3b,
tive pathway. Factors C5 to C9 are responsible
and it cleaves C5 into C5a and C5b. The com-
for membrane attack. Other components not
plex of C5b and C6 allows deposition of C7 in
shown here regulate the system.
the bacterial membrane. C8 and numerous C9
As in blood coagulation (see p. 290), the
molecules—which form the actual pore—then
early components in the complement system
bind to this core.
are serine proteinases, which mutually acti-
vate each other through limited proteolysis.
They create a self-reinforcing enzyme cas-
cade. Factor C3, the products of which are
involved in several functions, is central to
the complement system.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Immune response
299
A. Complement activation
Pathogen surface
Immune complex
Lipopolysaccharide
C4
C2
B
C1
C2b
D
C4b
C2a
Bb
Classic pathway
C3 convertase
Alternative pathway
C2aC4b
C3bBb
1
Collagen-
Is
C3
activated
like
filament
S C
when
O
C1q
C3
C3b
binds
1
to Ab
C3a
C3a
C1r*
and
C3b
C1s*
S C
H2O
O
NH2
Chemo-
Opson-
Affinity
C1q
taxis
ization
for
C3b’
antibody
SH C
domain
Fc
NHO
Bacterium
Membrane
1
C3/C5 convertase
attack
3.4.21.43/47
C6
C7
C8
C9
C5
2
C5b
Bacterial membrane
Membrane
attack complex
2
C5 convertase 3.4.21.47
C2aC3bC4b or (C3b)2 Bb
Structure of the membrane attack complex
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
300
Tissues and organs
Disulfide bonds link the two heavy chains to
Antibodies
each other and also link the heavy chains to
Soluble antigen receptors, which are formed
the light chains. Inside the domains, there are
by activated B cells (plasma cells; see p. 294)
also disulfide bonds that stabilize the tertiary
and released into the blood, are known as
structure. The domains are approximately 110
antibodies. They are also members of the im-
amino acids (AA) long and are homologous
munoglobulin family (Ig; see p. 296). Anti-
with each other. The antibody structure evi-
bodies are an important part of the humoral
dently developed as a result of gene duplica-
immune defense system. They have no anti-
tion. In its central region, known as the
microbial properties themselves, but support
“hinge” region, the antibodies are highly mo-
the cellular immune system in various ways:
bile.
1. They bind to antigens on the surface of
pathogens and thereby prevent them from
B. Classes of immunoglobulins
interacting with body cells
(neutralization;
see p. 404, for example).
Human immunoglobulins are divided into
2. They link single-celled pathogens into
five classes. IgA (with two subgroups), IgD,
aggregates (immune complexes), which are
IgE, IgG (with four subgroups), and IgM are
more easily taken up by phagocytes (aggluti-
defined by their H chains, which are desig-
nation).
nated by the Greek letters α, δ, ε, γ, and µ. By
3. They activate the complement system
contrast, there are only two types of L chain
(see p. 298) and thereby promote the innate
(κ and λ). IgD and IgE (like IgG) are tetramers
immune defense system (opsonization).
with the structure H2L2. By contrast, soluble
In addition, antibodies have become indis-
IgA and IgM are multimers that are held
pensable aids in medical and biological diag-
together by disulfide bonds and additional
nosis (see p. 304).
J peptides (joining peptides).
The antibodies have different tasks. IgMs
are the first immunoglobulins formed after
A. Domain structure of immunoglobulin G
contact with a foreign antigen. Their early
Type G immunoglobulins (IgG) are quantita-
forms are located on the surface of B cells
tively the most important antibodies in the
(see p. 296), while the later forms are se-
blood, where they form the fraction of γ-glob-
creted from plasma cells as pentamers. Their
ulins (see p. 276). IgGs (mass 150 kDa) are
action targets microorganisms in particular.
tetramers with two heavy chains (H chains;
Quantitatively, IgGs are the most important
red or orange) and two light chains (L chains;
immunoglobulins (see the table showing se-
yellow). Both H chains are glycosylated (vio-
rum concentrations). They occur in the blood
let; see also p. 43).
and interstitial fluid. As they can pass the
The proteinase papain cleaves IgG into two
placenta with the help of receptors, they can
Fab fragments and one Fc fragment. The Fab
be transferred from mother to fetus. IgAs
(“antigen-binding”) fragments, which each
mainly occur in the intestinal tract and in
consist of one L chain and the N-terminal
body secretions. IgEs are found in low con-
part of an H chain, are able to bind antigens.
centrations in the blood. As they can trigger
The Fc (“crystallizable”) fragment is made up
degranulation of mast cells (see p. 380), they
of the C-terminal halves of the two H chains.
play an important role in allergic reactions.
This segment serves to bind IgG to cell sur-
The function of IgDs is still unexplained. Their
faces, for interaction with the complement
plasma concentration is also very low.
system and antibody transport.
Immunoglobulins are constructed in a
modular fashion from several immunoglobu-
lin domains (shown in the diagram on the
right in Ω form). The H chains of IgG contain
four of these domains (VH, CH1, CH2, and
CH3) and the L chains containtwo (CL and VL).
The letters C and V designate constant or
variable regions.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Immune response
301
A. Domain structure of immunoglobulin G
Antigen-
Heavy
binding
chain
site
“Hinge” region
(450 AA)
Variable
V
H
domain
Light
chain
CH1
(212 AA)
Fab fragment
VL
S
S
CL
S
S
Fab
Cleavage site
Oligo-
for papain
saccharide
CH2
3.4.22.2
Light chain
Oligo-
Fc
saccharide
Disulfide
CH3
bond
C-terminal end
Heavy chain
B. Classes of immunoglobulins
IgA
IgD
IgE
IgG
IgM
360-720 kDa
172 kDa
196 kDa
150 kDa
935 kDa
J
J
IgG
IgA
13.5
3.5
IgM
IgD
IgE
1.5
0.03
0.00005
Serum concentration (g · l-1)
Chain:
H
α
δ
ε
γ
µ
L
κ or λ
κ or λ
κ or λ
κ or λ
κ or λ
Structure:
(α2 κ2)
δ2 κ2
ε2 κ2
γ2 κ2
(µ2 κ2)5 J
nJ
(α2 λ2)
δ2 λ2
ε2 λ2
γ2 λ2
(µ2 λ2)5 J
nJ
n =1, 2 or 3
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
302
Tissues and organs
maturation give rise to randomly combined
Antibody biosynthesis
new genes (“mosaic genes”).
3. Somatic mutation. During differentiation
The acquired (adaptive) immune system (see
of B cells into plasma cells, the coding genes
p. 294) is based on the ability of the lympho-
mutate. In this way, the “primordial” germ-
cytes to keep an extremely large repertoire of
line genes can become different somatic genes
antigen receptors and soluble antibodies
in the individual B cell clones.
ready for use, so that even infections involv-
ing new types of pathogen can be combated.
The wide range of immunoglobulins (Ig) are
C. Biosynthesis of a light chain
produced by genetic recombination and addi-
We can look at the basic features of the ge-
tional mutations during the development and
netic organization and synthesis of immuno-
maturation of the individual lymphocytes.
globulins using the biosynthesis of a mouse κ
chain as an example. The gene segments for
this light chain are designated L, V, J, and C.
A. Variability of immunoglobulins
They are located on chromosome 6 in the
germ-line DNA (on chromosome 2 in humans)
It is estimated that more than 108 different
and are separated from one another by in-
antibody variants occur in every human
trons (see p. 242) of different lengths.
being. This variability affects both the heavy
Some 150 identical L segments code for the
and the light chains of immunoglobulins.
signal peptide
(“leader sequence,”
17-20
There are five different types of heavy (H)
amino acids) for secretion of the product
chain, according to which the antibody classes
(see p. 230). The V segments, of which there
are defined (α, δ, ε, γ, µ), and two types of light
are 150 different variants, code for most of the
(L) chain (κ and λ; see p. 300). The various Ig
variable domains (95 of the 108 amino acids).
types that arise from combinations of these
L and V segments always occur in pairs—in
chains are known as isotypes. During immu-
tandem, so to speak. By contrast, there are
noglobulin biosynthesis, plasma cells can
only five variants of the J segments (joining
switch from one isotype to another (“gene
segments) at most. These code for a peptide
switch”). Allotypic variation is based on the
with 13 amino acids that links the variable
existence of various alleles of the same
part of the κ chains to the constant part. A
gene—i. e., genetic differences between indi-
single C segment codes for the constant part
viduals. The term idiotypic variation refers to
of the light chain (84 amino acids).
the fact that the antigen binding sites in the
During the differentiation of B lympho-
Fab fragments can be highly variable. Idiotypic
cytes, individual V/J combinations arise in
variation affects the variable domains (shown
each B cell. One of the 150 L/V tandem seg-
here in pink) of the light and heavy chains. At
ments is selected and linked to one of the five
certain sites—known as the hypervariable re-
J segments. This gives rise to a somatic gene
gions (shown here in red)—variation is partic-
that is much smaller than the germline gene.
ularly wide; these sequences are directly in-
Transcription of this gene leads to the forma-
volved in the binding of the antigen.
tion of the hnRNA for the κ chain, from which
introns and surplus J segments are removed
by splicing (see p. 246). Finally, the completed
B. Causes of antibody variety
mRNA still contains one each of the L-V-J-C
There are three reasons for the extremely
segments and after being transported into the
wide variability of antibodies:
cytoplasm is available for translation. The
1. Multiple genes. Various genes are avail-
subsequent steps in Ig biosynthesis follow
able to code for the variable protein domains.
the rules for the synthesis of membrane-
Only one gene from among these is selected
bound or secretory proteins (see p. 230).
and expressed.
2. Somatic recombination. The genes are
divided into several segments, of which there
are various versions. Various (“untidy”) com-
binations of the segments during lymphocyte
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Immune response
303
A. Variability of immunoglobulins
Constant
Variable
domain
domain
Hyper-
κ or λ
variable
regions
α,δ,ε,γ
or µ
Constant
domain
Isotypic
Allotypic
Idiotypic
B. Origins of antibody variety
C. Biosynthesis of a light chain
V
L
V
L
L
L
C
1
V2
V3
V4
V5
V6
V
1
V2
V3
Vn
J1
J2
J3
J4
DNA
Intro
Is removed
Germ
line
DNA
V/J Recombination
Protein
1. Multiple genes
V
L
L
L
C
J1
J2
J3
J4
V1
V2
V3
J2
J3
Germ line
B cell
Point mutations
DNA
DNA
during B cell
maturation
Somatic
DNA
Transcription
in B cell
L
C
V3
J2
J3
AAA
Protein
hnRNA
2. Somatic recombination
Splicing
Germ line
DNA
Selection and
L
C
V3
J2
linkage of
gene segments
Somatic
AAA
DNA
mRNA
Translation
V3 J
H3N
Protein
2
C
Protein
3. Somatic mutation
COO
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
304
Tissues and organs
or in vivo by producing ascites fluid in mice
Monoclonal antibodies,
(6).
immunoassay
B. Immunoassay
A. Monoclonal antibodies
Immunoassays are semiquantitative proce-
Monoclonal antibodies (MABs) are secreted
dures for assessing substances with low con-
by immune cells that derive from a single
centrations. In principle, immunoassays can
antibody-forming cell
(from a single cell
be used to assess any compound against
clone). This is why each MAB is directed
which antibodies are formed.
against only one specific epitope of an immu-
The basis for this procedure is the anti-
nogenic substance, known as an “antigenic
gen-antibody “reaction”—i. e., specific binding
determinant.” Large molecules contain several
of an antibody to the molecule being assayed.
epitopes, against which various antibodies are
Among the many different immunoassay
formed by various B cells. An antiserum con-
techniques that have been developed—e. g.,
taining a mixture of all of these antibodies is
radioimmunoassay (RIA), and chemolumines-
described as being polyclonal.
cence immunoassay (CIA)—a version of the
To obtain MABs, lymphocytes isolated from
enzyme-linked immunoassay (EIA) is shown
the spleen of immunized mice (1) are fused
here.
with mouse tumor cells (myeloma cells, 2).
The substance to be assayed—e. g., the hor-
This is necessary because antibody-secreting
mone thyroxine in a serum sample—is pipet-
lymphocytes in culture have a lifespan of only
ted into a microtiter plate (1), the walls of
a few weeks. Fusion of lymphocytes with tu-
which are coated with antibodies that specif-
mor cells gives rise to cell hybrids, known as
ically bind the hormone. At the same time, a
hybridomas, which are potentially immortal.
small amount of thyroxine is added to the
Successful fusion (2) is a rare event, but the
incubation to which an enzyme known as
frequency can be improved by adding poly-
the “tracer” (1) has been chemically coupled.
ethylene glycol (PEG). To obtain only success-
The tracer and the hormone being assayed
fully fused cells, incubation is required for an
compete for the small number of antibody
extended period in a primary culture with
binding sites available. After binding has
HAT medium (3), which contains hypoxan-
taken place (2), all of the unbound molecules
thine, aminopterin, and thymidine. Amino-
are rinsed out. The addition of a substrate
pterin, an analogue of dihydrofolic acid, com-
solution for the enzyme (a chromogenic solu-
petitively inhibits dihydrofolate reductase and
tion) then triggers an indicator reaction (3),
thus inhibits the synthesis of dTMP
(see
the products of which can be assessed using
p. 402). As dTMP is essential for DNA synthe-
photometry (4).
sis, myeloma cells cannot survive in the pres-
The larger the amount of enzyme that can
ence of aminopterin. Although spleen cells
bind to the antibodies on the container’s
are able to circumvent the inhibitory effect
walls, the larger the amount of dye that is
of aminopterin by using hypoxanthine and
produced. Conversely, the larger the amount
thymidine, they have a limited lifespan and
of the substance being assayed that is present
die. Only hybridomas survive culture in HAT
in the sample, the smaller the amount of
medium, because they possess both the im-
tracer that can be bound by the antibodies.
mortality of the myeloma cells and the spleen
Quantitative analysis can be carried out
cells’ metabolic side pathway.
through parallel measurement using stan-
Only a few fused cells actually produce
dards with a known concentration.
antibodies. To identify these cells, the hybrid-
omas have to be isolated and replicated by
cloning (4). After the clones have been tested
for antibody formation, positive cultures are
picked out and selected by further cloning (5).
This results in hybridomas that synthesize
monoclonal antibodies. Finally, MAB produc-
tion is carried out in vitro using a bioreactor,
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Immune response
305
A. Monoclonal antibodies
Immunized
Antigen
Hybridoma cells
Antibody test:
mouse
Positive
cultures
A B C
A B
With
HAT
PEG
medium
Immunization
Spleen cells
C
1.
Fusion
Primary culture
2.
3.
Myeloma cells
Cell culture
Mouse with
ascites
Subcloning
4.
Cloning
5.
C
C C
Production of
C
C
monoclonal
C
C
antibodies
6.
Culture of
Selection
C
C
B
B
A
A
a selected
of positive
C
B
A
clone
clones
Clone
Cell culture
B. Immunoassay
Enzyme:
peroxidase
Sample
Tracer
Antibodies
E
Incubate
Microtiter plate
1.
2.
Absorption
Working range
C
10-6-10-12 M
Read
E
Wash
absorption
F
Substrate: H2O2
Concentration
Chromogen: C
F
C
C
F
E
E
C
C
C
C
C
F
F
Incubate
4.
3.
C
F
Peroxidase
Chromogen + H2O2
Dye
+ H2O
1.11.1.7
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
306
Tissues and organs
the form of the polysaccharide glycogen or
Liver: functions
converted into fatty acids. When there is a
Weighing 1.5 kg, the liver is one of the largest
drop in the blood glucose level, the liver re-
organs in the human body. Although it only
leases glucose again by breaking down glyco-
represents 2-3% of the body’s mass, it ac-
gen. If the glycogen store is exhausted, glu-
counts for 25-30% of oxygen consumption.
cose can also be synthesized by gluconeogen-
esis from lactate, glycerol, or the carbon skel-
eton of amino acids (see p. 310).
A. Diagram of a hepatocyte
Lipid metabolism. The liver synthesizes
The 3
1011 cells in the liver—particularly the
fatty acids from acetate units. The fatty acids
hepatocytes, which make up 90% of the cell
formed are then used to synthesize fats and
mass—are the central location for the body’s
phospholipids, which are released into the
intermediary metabolism. They are in close
blood in the form of lipoproteins. The liver’s
contact with the blood, which enters the liver
special ability to convert fatty acids into ke-
from the portal vein and the hepatic arteries,
tone bodies and to release these again is also
flows through capillary vessels known as si-
important (see p. 312).
nusoids, and is collected again in the central
Like other organs, the liver also synthesizes
veins of the hepatic lobes. Hepatocytes are
cholesterol, which is transported to other tis-
particularly rich in endoplasmic reticulum,
sues as a component of lipoproteins. Excess
as they carry out intensive protein and lipid
cholesterol is converted into bile acids in the
synthesis. The cytoplasm contains granules of
liver or directly excreted with the bile (see
insoluble glycogen. Between the hepatocytes,
p. 314).
there are bile capillaries through which bile
Amino acid and protein metabolism. The
components are excreted.
liver controls the plasma levels of the amino
acids. Excess amino acids are broken down.
With the help of the urea cycle (see p. 182),
B. Functions of the liver
the nitrogen from the amino acids is con-
The most important functions of the liver are:
verted into urea and excreted via the kidneys.
1. Uptake of nutrients supplied by the in-
The carbon skeleton of the amino acids enters
testines via the portal vein.
the intermediary metabolism and serves for
2. Biosynthesis of endogenous compounds
glucose synthesis or energy production. In
and storage, conversion, and degradation of
addition, most of the plasma proteins are syn-
them into excretable molecules (metabolism).
thesized or broken down in the liver (see
In particular, the liver is responsible for the
p. 276).
biosynthesis and degradation of almost all
Biotransformation. Steroid hormones and
plasma proteins.
bilirubin, as well as drugs, ethanol, and other
3. Supply of the body with metabolites and
xenobiotics are taken up by the liver and in-
nutrients.
activated and converted into highly polar me-
4. Detoxification of toxic compounds by
tabolites by conversion reactions (see p. 316).
biotransformation.
Storage. The liver not only stores energy
5. Excretion of substances with the bile.
reserves and nutrients for the body, but also
certain mineral substances, trace elements,
and vitamins, including iron, retinol, and vi-
C. Hepatic metabolism
tamins A, D, K, folic acid, and B12.
The liver is involved in the metabolism of
practically all groups of metabolites. Its func-
tions primarily serve to cushion fluctuations
in the concentration of these substances in
the blood, in order to ensure a constant sup-
ply to the peripheral tissues (homeostasis).
Carbohydrate metabolism. The liver takes
up glucose and other monosaccharides from
the plasma. Glucose is then either stored in
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Liver
307
A. Diagram of a hepatocyte
Erythrocyte
Disse
Sinusoid
space
Lipoproteins
Glycogen
***
*
*
Golgi
complex
*
Nucleus
Rough ER
Smooth ER
Microbody
*
Mitochondrion
*
*
Lysosome
*
Desmosome
Biliary capillary
B. Functions of the liver
Vena cava
Supply
Metabolism
3
Detoxifica-
tion
Biosynthesis
B
Biotrans-
Storage
S
formation
Conversion
C
4
and
Degradation
2
Portal vein
1
5
Uptake
Excretion
Gallbladder
From the gastrointestinal
Bile duct
tract, pancreas, spleen
Intestine
C. Liver metabolism
Carbohydrate
Lipid
Amino acid
Biotrans-
metabolism
metabolism
metabolism
formation
Glucose
BSC
Fatty acids
BC
Amino acids
C
Steroid hormones
EC
Galactose
C
Fats
BC
Urea
B
Bile pigments
EC
Fructose
C
Ketone bodies
B
Ethanol
C
Mannose
C
Cholesterol
BEC
Drugs
EC
Pentoses
BC
Bile acids
BE
Plasma
Lactate
C
Vitamins
SC
proteins
Glycerol
BC
Glycogen
BSC
Lipoproteins
BC
Albumin
BC
B
Biosynthesis
Coagulation
C
Conversion and
factors
BC
degradation
Hormones
C
E
Excretion
Enzymes
BC
S
Storage
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
308
Tissues and organs
acids, ketone bodies). By contrast, the central
Buffer function in organ meta-
nervous system (CNS) is dependent on glu-
bolism
cose. It is only able to utilize ketone bodies
after a prolonged phase of hunger (B).
All of the body’s tissues have a constant re-
quirement for energy substrates and nu-
trients. The body receives these metabolites
B. Postabsorptive state
with food, but the supply is irregular and in
When the food supply is interrupted, the
varying amounts. The liver acts here along
postabsorbtive state quickly sets in. The pan-
with other organs, particularly adipose tissue,
creatic A cells now release increased amounts
as a balancing buffer and storage organ.
of glucagon, while the B cells reduce the
In the metabolism, a distinction is made
amount of insulin they secrete. The reduced
between the absorptive state (well-fed state)
insulin/glucagon quotient leads to switching
immediately after a meal and the postabsorb-
of the intermediary metabolism. The body
tive state
(state of starvation), which starts
now falls back on its energy reserves. To do
later and can merge into hunger. The switch-
this, it breaks down storage substances (gly-
ing of the organ metabolism between the two
cogen, fats, and proteins) and shifts energy-
phases depends on the concentration of en-
supplying metabolites between the organs.
ergy-bearing metabolites in the blood (plas-
The liver first empties its glycogen store
ma level). This is regulated jointly by hor-
(glycogenolysis; see p. 156). It does not use
mones and by the autonomic nervous system.
the released glucose itself, however, but sup-
plies the other tissues with it. In particular,
the brain, adrenal gland medulla, and eryth-
A. Absorptive state
rocytes depend on a constant supply of glu-
The absorptive state continues for 2-4 hours
cose, as they have no substantial glucose re-
after food intake. As a result of food digestion,
serves themselves. When the liver’s glycogen
the plasma levels of glucose, amino acids, and
reserves are exhausted after
12-24 hours,
fats (triacylglycerols) temporarily increase.
gluconeogenesis begins (see p. 154). The pre-
The endocrine pancreas responds to this by
cursors for this are derived from the muscu-
altering its hormone release—there is an in-
lature (amino acids) and adipose tissue (glyc-
crease in insulin secretion and a reduction in
erol from fat degradation). From the fatty
glucagon secretion. The increase in the insu-
acids that are released (see below), the liver
lin/glucagon quotient and the availability of
starts to form ketone bodies (ketogenesis; see
substrates trigger an anabolic phase in the
p. 312). These are released into the blood and
tissues—particularly liver, muscle, and adi-
serve as important energy suppliers during
pose tissues.
the hunger phase. After 1-2 weeks, the CNS
The liver forms increased amounts of gly-
also starts to use ketone bodies to supply part
cogen and fats from the substrates supplied.
of its energy requirements, in order to save
Glycogen is stored, and the fat is released into
glucose.
the blood in very low density lipoproteins
In muscle, the extensive glycogen reserves
(VLDLs).
are exclusively used for the muscles’ own
Muscle also refills its glycogen store and
requirements (see p. 320). The slowly initi-
synthesizes proteins from the amino acids
ated protein breakdown in muscle supplies
supplied.
amino acids for gluconeogenesis in the liver.
Adipose tissue removes free fatty acids
In adipose tissue, glucagon triggers lipoly-
from the lipoproteins, synthesizes triacylgly-
sis, releasing fatty acids and glycerol. The fatty
cerols from them again, and stores these in
acids are used as energy suppliers by many
the form of insoluble droplets.
types of tissue (with the exception of brain
During the absorptive state, the heart and
and erythrocytes). An important recipient of
neural tissue mainly use glucose as an energy
the fatty acids is the liver, which uses them for
source, but they are unable to establish any
ketogenesis.
substantial energy stores. Heart muscle cells
are in a sense “omnivorous,” as they can also
use other substances to produce energy (fatty
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Liver
309
A. Absorptive state
Glycogen
Proteins
Nervous
tissue
Muscle
Glycogen
Fat
Liver
Triacylglycerols
Glucose
Amino acids
Fat
High blood
Heart
Adipose tissue
glucose level
Insulin
Glucagon
Storage compound
Glucose
Amino
Intestine
acids
Pancreas
B. Postabsorptive state
Erythrocytes
Gluconeogenesis
Glycogen
Glucose
Amino acids
Proteins
Nervous
tissue
Muscle
Glycogen
Do not have
energy
Ketone bodies
reserves
Liver
Fatty acids
Triacylglycerols
Adipose tissue
Insulin
Heart
Glucagon
Glucose
amino
Intestine
acids
Pancreas
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
310
Tissues and organs
and therefore cannot be converted into oxalo-
Carbohydrate metabolism
acetic acid, the precursor for gluconeogenesis.
Besides fatty acids and ketone bodies, glucose
is the body’s most important energy supplier.
B. Fructose and galactose metabolism
The concentration of glucose in the blood (the
“blood glucose level”) is maintained at
Fructose is mainly metabolized by the liver,
4-6 mM (0.8-1.0 g L-1) by precise regula-
which channels it into glycolysis (left half of
tion of glucosesupplying and glucose-utilizing
the illustration).
processes. Glucose suppliers include the in-
A special ketohexokinase [1] initially phos-
testines (glucose from food), liver, and kid-
phorylates fructose into fructose
1-phos-
neys. The liver plays the role of a “glucostat”
phate. This is then cleaved by an aldolase
(see p. 308).
[2], which is also fructose-specific, to yield
The liver is also capable of forming glucose
glycerone
3-phosphate
(dihydroxyacetone
by converting other sugars—e. g., fructose and
phosphate) and glyceraldehyde. Glycerone
galactose—or by synthesizing from other me-
3-phosphate is already an intermediate of
tabolites. The conversion of lactate to glucose
glycolysis
(center), while glyceraldehyde
in the Cori cycle (see p. 338) and the conver-
can be phosphorylated into glyceraldehyde
sion of alanine to glucose with the help of the
3-phosphate by triokinase [3].
alanine cycle (see p. 338) are particularly im-
To a smaller extent, glyceraldehyde is also
portant for the supply of erythrocytes and
reduced to glycerol [4] or oxidized to glycer-
muscle cells.
ate, which can be channeled into glycolysis
Transporters in the plasma membrane of
following phosphorylation (not shown). The
hepatocytes allow insulin-independent trans-
reduction of glyceraldehyde
[4] uses up
port of glucose and other sugars in both di-
NADH. As the rate of degradation of alcohol
rections. In contrast to muscle, the liver pos-
in the hepatocytes is limited by the supply of
sesses the enzyme glucose-6-phosphatase,
NAD+, fructose degradation accelerates alco-
which can release glucose from glucose-6-
hol degradation (see p. 320).
phosphate.
Outside of the liver, fructose is channeled
into the sugar metabolism by reduction at C-2
to yield sorbitol and subsequent dehydration
A. Gluconeogenesis: overview
at C-1 to yield glucose (the polyol pathway;
Regeneration of glucose (up to 250 g per day)
not shown).
mainly takes place in the liver. The tubule
Galactose is also broken down in the liver
cells of the kidney are also capable of carrying
(right side of the illustration). As is usual with
out gluconeogenesis, but due to their much
sugars, the metabolism of galactose starts
smaller mass, their contribution only repre-
with a phosphorylation to yield galactose
sents around 10% of total glucose formation.
1-phosphate [5]. The connection to the glu-
Gluconeogenesis is regulated by hormones.
cose metabolism is established by C-4 epime-
Cortisol, glucagon, and epinephrine promote
rization to form glucose 1-phosphate. How-
gluconeogenesis, while insulin inhibits it
ever, this does not take place directly. Instead,
(see pp. 158, 244).
a transferase [6] transfers a uridine 5 -mono-
The main precursors of gluconeogenesis in
phosphate (UMP) residue from uridine di-
the liver are lactate from anaerobically work-
phosphoglucose
(UDPglucose) to galactose
ing muscle cells and from erythrocytes,
1-phosphate. This releases glucose 1-phos-
glucogenic amino acids from the digestive
phate, while galactose 1-phosphate is con-
tract and muscles
(mainly alanine), and
verted into uridine diphosphogalactose (UDP-
glycerol from adipose tissue. The kidney
galactose). This then is isomerized into UDP-
mainly uses amino acids for gluconeogenesis
glucose. The biosynthesis of galactose also fol-
(Glu, Gln; see p. 328).
lows this reaction pathway, which is freely
In mammals, fatty acids and other suppli-
reversible up to reaction [5]. Genetic defects
ers of acetyl CoA are not capable of being used
of enzymes [5] or [6] can lead to the clinical
for gluconeogenesis, as the acetyl residues
picture of galactosemia.
formed during β-oxidation in the tricarbox-
ylic acid cycle (see p. 132) are oxidized to CO2
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Liver
311
A. Gluconeogenesis: overview
Gluco-
Amino acids
neogenesis
Muscle
Liver
Kidney
Cortisol
Glucagon
Epinephrine
Insulin
Lactate
Glycogen
10%
Amino acids
Gluco-
Fructose
Glucose
neogenesis
90%
Galactose
Conversion
Glycerol
Plasma
concentration
4 - 6 mM
Fatty acids
Not possible
in mammals
Adipose tissue
B. Fructose and galactose metabolism
HOCH2
Fructose
Glucose
Galactose
HO
O
H
ATP
OH
H
OH
ATP
1
H
5
H
OH
ADP
ADP
Fructose 1-
P
Glucose 6-
P
Glucose 1-
P
6
Galactose 1-
P
2
Fructose 6-
P
UDP-Glucose
UDP-Galactose
Fructose 1,6-bis-
P
7
Glycer-
Glycer-
Glycerone 3-
P
aldehyde
aldehyde 3-
P
Glycolysis
3
NADH+H +
4
Pyruvate
+
NAD
ATP
ADP
Glycerol
Ketohexokinase
Triokinase
Galactokinase
UDPglucose
1
3
5
7
2.7.1.3
2.7.1.28
2.7.1.6
4-epimerase
5.1.3.2
Fructose-bisphosphate
Aldehyde reductase
Hexose-1-phosphate
2
4
6
aldolase 4.1.2.13
1.1.1.21
uridyltransferase
2.7.7.12
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
312
Tissues and organs
B. Biosynthesis of ketone bodies
Lipid metabolism
At high concentrations of acetyl-CoA in the
The liver is the most important site for the
liver mitochondria, two molecules condense
formation of fatty acids, fats (triacylglycerols),
to form acetoacetyl CoA [1]. The transfer of
ketone bodies, and cholesterol. Most of these
another acetyl group
[2] gives rise to
products are released into the blood. In con-
3-hydroxy-3-methylglutaryl-CoA (HMG CoA),
trast, the triacylglycerols synthesized in adi-
which after release of acetyl CoA [3] yields
pose tissue are also stored there.
free acetoacetate (Lynen cycle). Acetoacetate
can be converted to 3-hydroxybutyrate by
A. Lipid metabolism
reduction [4], or can pass into acetone by
nonenzymatic decarboxylation
[5]. These
Lipid metabolism in the liver is closely linked
three compounds are together referred to as
to the carbohydrate and amino acid metabo-
“ketone bodies,” although in fact 3-hydroxy-
lism. When there is a good supply of nutrients
butyrate is not actually a ketone. As reaction
in the resorptive (wellfed) state (see p. 308),
[3] releases an H+ ion, metabolic acidosis can
the liver converts glucose via acetyl CoA into
occur as a result of increased ketone body
fatty acids. The liver can also take up fatty
synthesis (see p. 288).
acids from chylomicrons, which are supplied
The ketone bodies are released by the liver
by the intestine, or from fatty acid-albumin
into the blood, in which they are easily solu-
complexes (see p. 162). Fatty acids from both
ble. Blood levels of ketone bodies therefore
sources are converted into fats and phospho-
rise during periods of hunger. Together with
lipids. Together with apoproteins, they are
free fatty acids, 3-hydroxybutyrate and ace-
packed into very-low-density lipoproteins
toacetate are then the most important energy
(VLDLs; see p. 278) and then released into
suppliers in many tissues (including heart
the blood by exocytosis. The VLDLs supply
muscle). Acetone cannot be metabolized and
extrahepatic tissue, particularly adipose tis-
is exhaled via the lungs or excreted with
sue and muscle.
urine.
In the postresorptive state (see p. 292)—
To channel ketone bodies into the energy
particularly during fasting and starva-
metabolism, acetoacetate is converted with
tion—the lipid metabolism is readjusted and
the help of succinyl CoA into succinic acid
the organism falls back on its own reserves. In
and acetoacetyl CoA, which is broken down
these conditions, adipose tissue releases fatty
by β-oxidation into acetyl CoA (not shown;
acids. They are taken up by the liver and are
see p. 180).
mainly converted into ketone bodies (B).
If the production of ketone bodies exceeds
Cholesterol can be derived from two sour-
the demand for them outside the liver, there
ces—food or endogenous synthesis from ace-
is an increase in the concentration of ketone
tyl-CoA. A substantial percentage of endo-
bodies in the plasma (ketonemia) and they are
genous cholesterol synthesis takes place in
also eventually excreted in the urine (ketonu-
the liver. Some cholesterol is required for
ria). Both phenomena are observed after pro-
the synthesis of bile acids (see p. 314). In ad-
longed starvation and in inadequately treated
dition, it serves as a building block for cell
diabetes mellitus. Severe ketonuria with ke-
membranes (see p. 216), or can be esterified
toacidosis can cause electrolyte shifts and
with fatty acids and stored in lipid droplets.
loss of consciousness, and is therefore life-
The rest is released together into the blood in
threatening (ketoacidotic coma).
the form of lipoprotein complexes (VLDLs)
and supplies other tissues. The liver also con-
tributes to the cholesterol metabolism by tak-
ing up from the blood and breaking down
lipoproteins that contain cholesterol and cho-
lesterol esters (HDLs, IDLs, LDLs; see p. 278).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Liver
313
A. Lipid metabolism
Intestine
Adipose tissue
Muscle
Glucose
Fatty acids
Acetyl CoA
Ketone bodies
Ketone
bodies
Apolipo-
Fats
proteins
Fats
Fatty acids
Phospholipids
VLDL
Cholesterol
Cholesterol
Chylomicron
residues
HDL
Cholesterol
Bile
IDL
esters
acids
LDL
Bile
Peripheral
tissues
Absorptive state
Postabsorptive state
A. Biosynthesis of ketone bodies
H
O
H
O
H O
H OH H
O
3
1
H
C
C
S
H
C
C
C C
S
H
C
C
C C
S
A
A
A
H
H
H
H
H
H
C
H
COO
3-Hydroxy-
Acetyl CoA
1
Acetacetyl CoA
2
3-methyl-
glutaryl CoA
Acetyl CoA
CoA
Acetyl CoA CoA
3
H OH H
H O H
H +
H
O
H
3
1
H
C C
C
COO
H
C C
C
COO
H
C
C
C
H
H H
H
H
H
H
H
3-Hydroxybutyrate
4
Acetacetate
5
Acetone
N
A
N
A
CO2
1
Acetyl-CoA-C-acyltransferase 2.3.1.16
3
Hydroxymethylglutaryl-CoA lyase 4.1.3.4
2
Hydroxymethylglutaryl-CoA synthase 4.1.3.5
4
3-Hydroxybutyrate dehydrogenase 1.1.1.30
5
Nonenzymatic reaction
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
314
Tissues and organs
cleavage of the OH group at C-7 (see B).
Bile acids
They are therefore referred to as secondary
Bile is an important product released by the
bile acids.
hepatocytes. It promotes the digestion of fats
from food by emulsifying them in the small
B. Metabolism of bile salts
intestine (see p. 2770). The emulsifying com-
ponents of bile, apart from phospholipids,
Bile salts are exclusively synthesized in the
mainly consist of bile acids and bile salts
liver (see A). The slowest step in their biosyn-
(see below). The bile also contains free cho-
thesis is hydroxylation at position 7 by a 7-D-
lesterol, which is excreted in this way (see
hydroxylase. Cholic acid and other bile acids
p. 312).
inhibit this reaction (end-product inhibition).
In this way, the bile acids present in the liver
regulate the rate of cholesterol utilization.
A. Bile acids and bile salts
Before leaving the liver, a large proportion
Bile acids are steroids consisting of 24 C atoms
of the bile acids are activated with CoA and
carrying one carboxylate group and several
then conjugated with the amino acids glycine
hydroxyl groups. They are formed from cho-
or taurine (2; cf. A). In this way, cholic acid
lesterol in the liver via an extensive reaction
gives rise to glycocholic acid and taurocholic
pathway (top). Cytochrome P450 enzymes in
acid. The liver bile secreted by the liver be-
the sER of hepatocytes are involved in many
comes denser in the gallbladder as a result of
of the steps (seep. 318). Initially, the choles-
the removal of water (bladder bile; 3).
terol double bond is removed. Monooxyge-
Intestinal bacteria produce enzymes that
nases then introduce one or two additional
can chemically alter the bile salts (4). The
OH groups into the sterane framework. Fi-
acid amide bond in the bile salts is cleaved,
nally, the side chain is shortened by three C
and dehydroxylation at C-7 yields the corre-
atoms, and the terminal C atom is oxidized to
sponding secondary bile acids from the pri-
a carboxylate group.
mary bile acids (5). Most of the intestinal bile
It is important that the arrangement of the
acids are resorbed again in the ileum (6) and
A and B rings is altered from trans to cis dur-
returned to the liver via the portal vein (en-
ing bile acid synthesis (see p. 54). The result
terohepatic circulation). In the liver, the sec-
of this is that all of the hydrophilic groups in
ondary bile acids give rise to primary bile
the bile acids lie on one side of the molecule.
acids again, from which bile salts are again
Cholesterol, which is weakly amphipathic
produced. Of the 15-30 g bile salts that are
(top), has a small polar “head” and an ex-
released with the bile per day, only around
tended apolar “tail.” By contrast, the much
0.5g therefore appears in the feces. This ap-
more strongly amphipathic bile acid mole-
proximately corresponds to the amount of
cules (bottom) resemble disks with polar top
daily de novo synthesis of cholesterol.
sides and apolar bottom sides. At physiolog-
ical pH values, the carboxyl groups are almost
Further information
completely dissociated and therefore nega-
tively charged.
The cholesterol excreted with the bile is
Cholic acid and chenodeoxycholic acid,
poorly water-soluble. Together with phos-
known as the primary bile acids, are quanti-
pholipids and bile acids, it forms micelles
tatively the most important metabolites of
(see p. 270), which keep it in solution. If the
cholesterol. After being biosynthesized, they
proportions of phospholipids, bile acids and
are mostly activated with coenzyme A and
cholesterol shift, gallstones can arise. These
then conjugated with glycine or the non-pro-
mainly consist of precipitated cholesterol
teinogenic amino acid taurine (see p. 62). The
(cholesterol stones), but can also contain
acid amides formed in this way are known as
Ca2+ salts of bile acids and bile pigments (pig-
conjugated bile acids or bile salts. They are
ment stones).
even more amphipathic than the primary
products.
Deoxycholic acid and lithocholic acid are
only formed in the intestine by enzymatic
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Liver
315
A. Bile acids and bile salts
Cholesterol
Bile acids
HO- in position
Primary bile acids
— Cholic acid
C-3
C-7
C-12
— Chenodeoxycholic
C-3
C-7
-
acid
Secondary bile acids
— Deoxycholic acid
C-3
-
C-12
— Lithocholic acid
C-3
-
-
C
D
A
B
3
Bile salts
HO
O
C
14 steps
OH
N COO
H
Cholic acid
COO
Glycine
OH
Glycocholic acid
12
HO
OH
C
D
O
A
B
3
7
C
SO3
OH
N
HO
OH
H
Taurine
Taurocholic acid
HO
OH
Bile salts = conjugated bile acids
B. Metabolism of bile salts
R
1
Primary
Cholesterol
bile acids
Intestine
2
Glycine, taurine
Bile salts
15 - 30 g
3
Gallbladder
Bile
Bile salts
Breakdown of bile salts
Glycine,
by intestinal bacteria
4
taurine
Secondary
Primary
bile acids
bile acids
5
Resorption
6
Enterohepatic
circulation
0,5 g
Feces
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
316
Tissues and organs
Most oxidation reactions are catalyzed by
Biotransformations
cytochrome P450
systems
(see p. 318).
The body is constantly taking up foreign sub-
These monooxygenases are induced by
stances (= xenobiotics) from food or through
their substrates and show wide specificity.
contact with the environment, via the skin
The substrate-specific enzymes of the ste-
and lungs. These substances can be natural
roid metabolism (see p. 376) are exceptions
in origin, or may have been synthetically pro-
to this.
duced by humans. Many of these substances
Phase II reactions (conjugate formation). Type
are toxic, particularly at high concentrations.
II reactions couple their substrates (bilirubin,
However, the body has effective mechanisms
steroid hormones, drugs, and products of
for inactivating and then excreting foreign
phase I reactions) via ester or amide bonds
substances through biotransformations. The
to highly polar negatively charged molecules.
mechanisms of biotransformation are similar
The enzymes involved are transferases, and
to those with which endogenous substances
their products are known as conjugates.
such as bile pigments and steroid hormones
The most common type of conjugate for-
are enzymatically converted. Biotransforma-
mation is coupling with glucuronate (GlcUA)
tions mainly take place in the liver.
as an O-or N-glucuronide. The coenzyme for
the reaction is uridine diphosphate glucuro-
A. Biotransformations
nate, the
“active glucuronate”
(see p. 110).
Phase I reactions (interconversion reactions).
Coupling with the polar glucuronate makes
Type I reactions introduce functional groups
an apolar
(hydrophobic) molecule more
into inert, apolar molecules or alter functional
strongly polar, and it becomes suf ciently
groups that are already present. In many
water-soluble and capable of being excreted.
cases, this is what first makes it possible for
Example (3) shows the glucuronidation of
foreign substances to conjugate with polar
tetrahydrocortisol, a metabolite of the gluco-
molecules via phase II reactions (see below).
corticoid cortisol (see p. 374).
Phase I reactions usually reduce the biological
The biosynthesis of sulfate esters with the
activity or toxicity of a substance (“detoxifica-
help of phosphoadenosine phosphosulfate
tion”). However, some substances only be-
(PAPS), the “active sulfate”, (see p. 110) and
come biologically active as a result of the
amide formation with glycine and glutamine
interconversion reaction (see, for example,
also play a role in conjugation. For example,
benzo[a]pyrene, p. 256) or become more toxic
benzoic acid is conjugated with glycine to
after interconversion than the initial sub-
form the more soluble and less toxic hippuric
stance (“toxification”).
acid (N-benzoylglycine; see p. 324).
Important phase I biotransformation reac-
In contrast with unconjugated compounds,
tions include:
the conjugates are much more water-soluble
and capable of being excreted. The conjugates
• Hydrolytic cleavages of ether, ester, and
are eliminated from the liver either by the
peptide bonds. Example (1) shows hydrol-
biliary route—i. e., by receptor-mediated ex-
ysis of the painkiller acetylsalicylic acid.
cretion into the bile—or by the renal route,
• Oxidations. Hydroxylations, epoxide for-
via the blood and kidneys by filtration.
mation, sulfoxide formation, dealkylation,
deamination. For example, benzene is oxi-
dized into phenol, and toluene (methylben-
Further information
zene) is oxidized into benzoic acid.
To detoxify heavy metals, the liver contains
• Reductions. Reduction of carbonyl, azo-, or
metallothioneins, a group of cysteine-rich pro-
nitro- compounds, dehalogenation.
teins with a high af nity for divalent metal
• Methylations. Example (2) illustrates the
ions such as Cd2+, Cu2+, Hg2+, and Zn2+. These
inactivation of the catecholamine norepi-
metal ions also induce the formation of metal-
nephrine by methylation of a phenolic OH
lothioneins via a special metal-regulating el-
group (see p. 334).
ement (MRE) in the gene's promoter (see
• Desulfurations. The reactions take place in
p. 244).
the hepatocytes on the smooth endoplas-
mic reticulum.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Liver
317
A. Biotransformations
Acetic acid
H2O
CH3-COOH
Foreign
Endogenous
O
substances:
substances
C
1
Drugs
Steroid hormones
O
CH
3
OH
Preservatives
and other low
COO
COO
Plasticizers
molecular weight
Pigments
substances
Acetylsalicylic acid
Salicylate
Pesticides etc.
Bile pigments
1. Hydrolysis of a drug
Poorly
soluble,
biologically
S-adenosyl
S-adenosyl homocysteine
active, some
methionine
toxic
H3N
CH2
H3N
CH2
Substrate
Transformation
induction
reactions:
Phase I
HO
CH
HO
CH
reactions
Hydrolytic
2
cleavage,
Epoxide formation
Dealkylation
OH
OCH3
Deamination
Reduction
OH
OH
Methylation
Desulfuration
Norepinephrine
O-methyl
norepinephrine
2. Methylation
of a hormone/neurotransmitter
Transformation
Tetrahydrocortisol
CH2OH
product
CO
HO
OH
Conjugate
formation:
Glucuronidation
HO
Phase II
Esterification
H
Substrate
reactions
with sulfate
Amidation
UDP-GlcUA
induction
with Gly and Glu
3
CH2OH
CO
UDP
HO
OH
COO
H
O
H
H
Water soluble,
OH
H
Conjugate
inactive,
HO
O
non-toxic
H
H
OH
Tetrahydrocortisol glucuronide
3. Glucuronidation of a hormone
Arylesterase
Catechol O-methyl-
1
2
3.1.1.2
transferase 2.1.1.6
Bile
Urine
3
Glucuronosyltransferase 2.4.1.17
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
318
Tissues and organs
Only a few examples of the numerous Cyt
Cytochrome P450 systems
P450-dependent reactions are shown here.
During the first phase of biotransformation in
Hydroxylation of aromatic rings (a) plays a
the liver, compounds that are weakly chemi-
central part in the metabolism of medicines
cally reactive are enzymatically hydroxylated
and steroids. Aliphatic methyl groups can also
(see p. 316). This makes it possible for them to
be oxidized to hydroxyl groups (b). Epoxi-
be conjugated with polar substances. The hy-
dation of aromatics (c) by Cyt P450 yields
droxylating enzymes are generally mono-
products that are highly reactive and often
oxygenases that contain a heme as the redox-
toxic. For example, the mutagenic effect of
active coenzyme (see p. 106). In the reduced
benzo[a]pyrene (see p. 244) is based on this
form, the heme can bind carbon monoxide
type of interconversion in the liver. In Cyt
(CO), and it then shows characteristic light
P450 dependent dealkylations (d), alkyl sub-
absorption at 450 nm. This was what led to
stituents of O, N, or S atoms are released as
this enzyme group being termed cytochrome
aldehydes.
P450 (Cyt P450).
Cyt P450 systems are also involved in many
B. Reaction mechanism
other metabolic processes—e. g., the biosyn-
thesis of steroid hormones (see p. 172), bile
The course of Cyt P450 catalysis is in principle
acids
(see p. 314), and eicosanoids
(see
well understood. The most important func-
p. 390), as well as the formation of unsatu-
tion of the heme group consists of converting
rated fatty acids (see p. 409). The liver’s red-
molecular oxygen into an especially reactive
dish-brown color is mainly due to the large
atomic form, which is responsible for all of the
amounts of P450 enzymes it contains.
reactions described above.
[1] In the resting state, the heme iron is
trivalent. Initially, the substrate binds near the
A. Cytochrome P450-dependent mono oxy-
heme group.
genases: reactions
[2] Transfer of an electron from FADH2 re-
Cyt P450-dependent monooxygenases cata-
duces the iron to the divalent form that is able
lyze reductive cleavage of molecular oxygen
to bind an O2 molecule (2).
(O2). One of the two oxygen atoms is trans-
[3] Transfer of a second electron and a
ferred to the substrate, while the other is re-
change in the valence of the iron reduce the
leased as a water molecule. The necessary re-
bound O2 to the peroxide.
ducing equivalents are transferred to the actual
[4] A hydroxyl ion is now cleaved from this
monooxygenase by an FAD-containing auxili-
intermediate. Uptake of a proton gives rise to
ary enzyme from the coenzyme NADPH+H+.
H2O and the reactive form of oxygen men-
Cyt P450
enzymes occur in numerous
tioned above. In this ferryl radical, the iron
forms in the liver, steroid-producing glands,
is formally tetravalent.
and other organs. The substrate specificity of
[5] The activated oxygen atom inserts itself
liver enzymes is low. Apolar compounds con-
into a C-H bond in the substrate, thereby
taining aliphatic or aromatic rings are partic-
forming an OH group.
ularly easily converted. These include endog-
[6] Dissociation of the product returns the
enous substances such as steroid hormones,
enzyme to its initial state.
as well as medical drugs, which are inacti-
vated by phase I reactions. This is why Cyt
P450 enzymes are of particular interest in
pharmacology. The degradation of ethanol in
the liver is also partly catalyzed by Cyt P450
enzymes (the “microsomal ethanol-oxidizing
system”; see p. 304). As alcohol and drugs are
broken down by the same enzyme system,
the effects of alcoholic drinks and medical
drugs can sometimes be mutually en-
hancing—even sometimes to the extent of
becoming life-threatening.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Liver
319
A. Cytochrome P450-dependent monooxygenases: reactions
R
Apolar substrate
CH2
H
O
H
3
C
O O
F
A
N
A
O
P
O
R
2H
1
R
Fe
H
H
O
C
O
H
O
F
A
N
A
HO
H2O
CH
2
P
(H ) O
O
O
O
Oxygenated product
R
1
Monooxygenase
a) Hydroxylation
b) Hydroxylation c) Epoxidation
d) Dealkylation
1.14.n.n [heme P450]
aromatic
alipathic
B. Reaction mechanism
HO
H
Product
Substrate
S
Fe3
6
1
Resting
S
Fe3
state
S
Fe3
H
O
H
F
A
N
A
P
O2
e
5
2
N
A
F
A
P
O
“Activated
e
S
Fe4
O
oxygen”
S
Fe2
O
Cyt P450
H
reductase
H
4
3
OH
S
Fe3
O
H2O
H
H
H
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
320
Tissues and organs
The rate of ethanol degradation in the liver
Ethanol metabolism
is limited by alcohol dehydrogenase activity.
The amount of NAD+ available is the limiting
A. Blood ethanol level
factor. As the maximum degradation rate is
Ethanol (EtOH, “alcohol”) naturally occurs in
already reached at low concentrations of
fruit in small quantities. Alcoholic drinks
ethanol, the ethanol level therefore declines
contain much higher concentrations. Their
at a constant rate (zero-order kinetics). The
alcohol content is usually given as percent
calorific value of ethanol is 29.4 kJ g-1. Alco-
by volume. To estimate alcohol uptake and
holic drinks—particularly in alcoholics—can
the blood alcohol level, it is useful to convert
therefore represent a substantial proportion
the amount to grams of ethanol (density
of dietary energy intake.
0.79 kg L-1). For example, a bottle of beer
(0.5 L at
4% v/v alcohol) contains
C. Liver damage due to alcohol
20 mL = 16 g of ethanol, while a bottle of
wine (0.7 L at
12 % v/v alcohol) contains
Alcohol is a socially accepted drug of abuse in
84 mL = 66 g ethanol.
Western countries. Due to the high potential
Ethanol is membrane-permeable and is
for addiction to develop, however, it is ac-
quickly resorbed. The maximum blood level
tually a “hard” drug and has a much larger
is already reached within 60-90 min after
number of victims than the opiate drugs, for
drinking. The resorption rate depends on var-
example. In the brain, ethanol is deposited in
ious conditions, however. An empty stomach,
membranes due to its amphipathic proper-
a warm drink (e. g., mulled wine), and the
ties, and it influences receptors for neuro-
presence of sugar and carbonic acid (e. g., in
transmitters (see p. 352). The effect of GABA
champagne) promote ethanol resorption,
is enhanced, while that of glutamate declines.
whereas a heavy meal reduces it. Ethanol is
High ethanol consumption over many
rapidly distributed throughout the body. A
years leads to liver damage. For a healthy
large amount is taken up by the muscles and
man, the limit is about 60 g per day, and for
brain, but comparatively little by adipose tis-
a woman about 50 g. However, these values
sue and bones. Roughly 70% of the body is
are strongly dependent on body weight,
accessible to alcohol. Complete resorption of
health status, and other factors.
the ethanol contained in one bottle of beer
Ethanol-related high levels of NADH+H+
(16 g) by a person weighing 70 kg (distribu-
and acetyl-CoA in the liver lead to increased
tion in
70 kg
70/100 = 49 kg) leads to a
synthesis of neutral fats and cholesterol.
blood alcohol level of
0.33
per thousand
However, since the export of these in the
(7.2 mM). The lethal concentration of alcohol
form of VLDLs (see p. 278) is reduced due to
is approximately 3.5 per thousand (76 mM).
alcohol, storage of lipids occurs (fatty liver).
This increase in the fat content of the liver
(from less than 5% to more than 50% of the
B. Ethanol metabolism
dry weight) is initially reversible. However, in
The major site of ethanol degradation is the
chronic alcoholism the hepatocytes are in-
liver, although the stomach is also able to me-
creasingly replaced by connective tissue.
tabolize ethanol. Most ethanol is initially oxi-
When liver cirrhosis occurs, the damage to
dized by alcohol dehydrogenase to form etha-
the liver finally reaches an irreversible stage,
nal (acetaldehyde). A further oxidization, cata-
characterized by progressive loss of liver
lyzed by aldehyde dehydrogenase, leads to ace-
functions.
tate. Acetate is then converted with the help
of acetate-CoA ligase to form acetyl CoA, using
ATP and providing a link to the intermediary
metabolism. In addition to cytoplasmic alco-
hol dehydrogenase, catalase and inducible
microsomal alcohol oxidase
(“MEOS”; see
p. 318) also contribute to a lesser extent to etha-
nol degradation. Many of the enzymes men-
tioned above are induced by ethanol.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Liver
321
A. Blood ethanol level
Vol.%
Blood alcohol level
[‰]
Whiskey
Cognac
40
1,0
Maximum after 60 - 90 min
Corn liquor
Constant decline
30
Cherry brandy
ca. 0.015 % · h-1
0,5
Rapid increase
Advocaat
20
due to easy
resorption
Red wine
10
White wine
Strong beer
0,0
Beer
1
2
3
4
5
6[h]
Water
Vodka (55 Vol.%) 0.75 g EtOH · kg-1
B. Ethanol metabolism
A c
c
u m u l a t e s
N
A
N
A
N
A
N
A
H
H
H
H
O
H
O
H C C O
1
H C
C
2
H
C
C
H H
H
H
H
O
Ethanol
Ethanal
Acetate
A
P P P
O2
2 H2O
S
‚MEOS‘
A
[Cyt P-450]
3
A
P
N
A
N
A
P P
P
P
H O
H
C
C
S
1
Alcohol dehydrogenase 1.1.1.1 [Zn
2
]
Fatty acid
A
and
H
2
Aldehyde dehydrogenase 1.2.1.3
cholesterol
Tricarboxylic
biosynthesis
acid cycle
Acetyl CoA
3
Acetate-CoA ligase 6.2.1.1
C. Liver damage due to alcohol
> 50-60 g EtOH
> 160 g EtOH
daily
daily
Healthy
Fatty
Cirrhotic
liver
liver
liver
Abstinence
Death
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
322
Tissues and organs
transported back into the blood by resorption,
Kidney: functions
to prevent losses of valuable metabolites and
A. Functions of the kidneys
electrolytes. In the proximal tubule, organic
metabolites (e. g., glucose and other sugars,
The kidneys’ main function is excretion of
amino acids, lactate, and ketone bodies) are
water and water-soluble substances (1). This
recovered by secondary active transport (see
is closely associated with their role in regulat-
p. 220). There are several group-specific
ing the body’s electrolyte and acid-base bal-
transport systems for resorbing amino acids,
ance (homeostasis, 2; see pp. 326 and 328).
with which hereditary diseases can be
Both excretion and homeostasis are subject to
associated
(e. g., cystinuria, glycinuria, and
hormonal control. The kidneys are also in-
Hartnup’s disease). HCO3-, Na+, phophate,
volved in synthesizing several hormones (3;
and sulfate are also resorbed by ATP-depend-
see p. 315). Finally, the kidneys also play a role
ent (active) mechanisms in the proximal tu-
in the intermediary metabolism (4), particu-
bule. The later sections of the nephron mainly
larly in amino acid degradation and gluconeo-
serve for additional water recovery and regu-
genesis (see p. 154).
lated resorption of Na+ and Cl- (see pp. 326,
The kidneys are extremely well-perfused
328). These processes are controlled by hor-
organs, with about 1500 L of blood flowing
mones (aldosterone, vasopressin).
through them every day. Approximately 180 L
Secretion. Some excretable substances are
of primary urine is filtered out of this. Re-
released into the urine by active transport in
moval of water leads to extreme concentra-
the renal tubules. These substances include H+
tion of the primary urine (to approximately
and K+ ions, urea, and creatinine, as well as
one-hundredth of the initial volume). As a
drugs such as penicillin.
result, only a volume of 0.5-2.0 L of final
Clearance. Renalclearanceisusedasaquan-
urine is excreted per day.
titative measure of renal function. It is defined
as the plasma volume cleared of a given sub-
B. Urine formation
stance per unit of time. Inulin, a fructose poly-
The functional unit of the kidney is the neph-
saccharide with a mass of ca. 6 kDa (see p. 40)
ron. It is made up of the Malpighian bodies or
that is neither actively excreted nor resorbed
renal corpuscles (consisting of Bowman’s cap-
but is freely filtered, has a clearance of
sules and the glomerulus), the proximal tu-
120mL min-1 in healthy individuals.
bule, Henle’s loop, and the distal tubule,
which passes into a collecting duct. The hu-
Further information
man kidney contains around one million
nephrons. The nephrons form urine in the
Concentrating urine and transporting it
following three phases.
through membranes are processes that re-
Ultrafiltration. Ultrafiltration of the blood
quire large amounts of energy. The kidneys
plasma in the glomerulus gives rise to primary
therefore have very high energy demands. In
urine, which is isotonic with plasma. The pores
the proximal tubule, the ATP needed is ob-
in the glomerular basal membrane, which are
tained from oxidative metabolism of fatty
made up of type IV collagen (see p. 344), have
acids, ketone bodies, and several amino acids.
an effective mean diameter of 2.9 nm. This al-
To a lesser extent, lactate, glycerol, and citric
lows all plasma components with a molecular
acid are also used. In the distal tubule and
mass of up to about 15 kDa to pass through
Henle’s loop, glucose is the main substrate
unhindered. At increasing masses, molecules
for the energy metabolism. The endothelial
are progressively held back; at masses greater
cells in the proximal tubule are also capable
than 65 kDa, they are completely unable to
of gluconeogenesis. The substrates for this are
enter the primary urine. This applies to almost
mainly the carbohydrate skeletons of amino
all plasma proteins—which in addition, being
acids. Their amino groups are used as ammo-
anions, are repelled by the negative charge in
nia for buffering urine (see p. 311). Enzymes
the basal membrane.
for peptide degradation and the amino acid
Resorption. All low-molecular weight
metabolism occur in the kidneys at high lev-
plasma components enter the primary urine
els of activity
(e. g., amino acid oxidases,
via glomerular filtration. Most of these are
amine oxidases, glutaminase).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Kidney
323
A. Functions of the kidneys
1. Excretion
3. Hormone synthesis
Water
Salts
Metabolic
wastes
Erythropoietin
Foreign
V
H
Calcitriol
substances
4. Metabolism
Glucose
Blood
2. Homeostasis
Acid-base balance
Gluconeo-
Electrolyte balance
Amino
genesis
acids
NH3
Urine
B. Urine formation
Distal
Resorption
tubule
Afferent
Glomerulus
arteriole
Proximal
Regulated
tubule
secretion
Ultra-
filtration
Final
urine
Collecting
Secretion
duct
Efferent
Bowman’s
arteriole
capsule
Renal corpuscle
Henle’s
loop
Ultrafiltration
Secretion
Resorption
Regulated
resorption
All solute
H
Glucose
plasma
K
Lactate
components
smaller
Drugs
2-Oxoacids
of H2O
than
Amino acids
3 nm = 15 kDaUric acid
Creatinine
Na , K , Ca2 , Mg2
2
2
Cl
, SO
HCO
4 , HPO4,
3
Water, etc.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
324
Tissues and organs
Modified amino acids, which occur in special
Urine
proteins such as hydroxyproline in collagen
and 3-methylhistidine in actin and myosin,
A. Urine
can be used as indicators of the degradation
Water and water-soluble compounds are ex-
of these proteins.
creted with the urine. The volume and com-
Other components of the urine are conju-
position of urine are subject to wide variation
gates with sulfuric acid, glucuronic acid, gly-
and depend on food intake, body weight, age,
cine, and other polar compounds that are
sex, and living conditions such as tempera-
synthesized in the liver by biotransformation
ture, humidity, physical activity, and health
(see p. 316). In addition, metabolites of many
status. As there is a marked circadian rhythm
hormones (catecholamines, steroids, seroto-
in urine excretion, the amount of urine and its
nin) also appear in the urine and can provide
composition are usually given relative to a 24-
information about hormone production. The
hour period.
proteohormone chorionic gonadotropin (hCG,
A human adult produces 0.5-2.0 L urine
mass ca. 36 kDa), which is formed at the onset
per day, around 95 % of which consists of
of pregnancy, appears in the urine due to its
water. The urine usually has a slightly acidic
relatively small size. Evidence of hCG in the
pH value (around 5.8). However, the pH value
urine provides the basis for an immunological
of urine is strongly affected by metabolic sta-
pregnancy test.
tus. After ingestion of large amounts of plant
The yellow color of urine is due to uro-
food, it can increase to over 7.
chromes, which are related to the bile pig-
ments produced by hemoglobin degradation
(see p. 194). If urine is left to stand long
B. Organic components
enough, oxidation of the urochromes may
Nitrogen-containing compounds are among
lead to a darkening in color.
the most important organic components of
urine. Urea, which is mainly synthesized in
C. Inorganic components
the liver (urea cycle; see p. 182), is the form in
which nitrogen atoms from amino acids are
The main inorganic components of the urine
+
excreted. Breakdown of pyrimidine bases also
are the cations Na+, K+, Ca2+, Mg2+, and NH4
produces a certain amount of urea
(see
and the anions Cl-, SO42-, and HPO42-, as well
p. 190). When the nitrogen balance is con-
as traces of other ions. In total, Na+ and Cl-
stant, as much nitrogen is excreted as is taken
represent about two-thirds of all the electro-
up (see p. 174), and the amount of urea in the
lytes in the final urine. Calcium and magne-
urine therefore reflects protein degradation:
sium occur in the feces in even larger quanti-
70 g protein in food yields approximately
ties. The amounts of the various inorganic
30 g urea in the urine.
components of the urine also depend on the
Uric acid is the end product of the purine
composition of the diet. For example, in
metabolism. When uric acid excretion via the
acidosis there can be a marked increase in
kidneys is disturbed, gout can develop (see
the excretion of ammonia (see p. 326). Excre-
p. 190). Creatinine is derived from the muscle
tion of Na+, K+, Ca2+, and phosphate via the
metabolism, where it arises spontaneously
kidneys is subject to hormonal regulation (see
and irreversibly by cyclization of creatine
p. 330).
and creatine phosphate (see p. 336). Since
the amount of creatinine an individual ex-
Further information
cretes per day is constant (it is directly pro-
portional to muscle mass), creatinine as an
Shifts in the concentrations of the physiolog-
endogenous substance can be used to mea-
ical components of the urine and the appear-
sure the glomerular filtration rate. The
ance of pathological urine components can be
amount of amino acids excreted in free form
used to diagnose diseases. Important exam-
is strongly dependent on the diet and on the
ples are glucose and ketone bodies, which are
ef ciency of liver function. Amino acid deriv-
excreted to a greater extent in diabetes
atives are also found in the urine (e. g., hippu-
mellitus (see p. 160).
rate, a detoxification product of benzoic acid).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Kidney
325
A. Urine
O
O
H
C
C
N
N COO
HN
C
H
C
O
C
C
N
Hippurate
O
N
H
0.15 g
H
Uric acid
0.3-2.0 g
NH2
from purine
H3C
C
degradation
Creatine
N
NH2
0.05-0.10 g
H2C
O
from muscle
C
metabolism
O
O
C
H2N
NH2
Creatinine
H3C
NH2
1.0-1.5 g
from creatine
N
C
Urea
and creatine
20-35 g
phosphate
H2C
N
from proteins
C
and amino acids
O
24-h urine
Volume: 0.5
-2l/day
pH: 5.8 (4.8 - 7.5)
Density: 1.015 - 1.022 kg/l
Glucose
Ketone bodies
Osmolarity: 50 -1300 mosmol/kg
< 0.16 g
< 3 g
Solids: 50 - 72 g/day
Proteins
Amino acids
< 0.15 g
1-3 g
C. Inorganic constituents
B. Organic constituents
Daily
excretion
(mmol)
Dissociation
dependent on pH
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
326
Tissues and organs
B. Ammonia excretion
Functions in the acid-base balance
Approximately 60 mmol of protons are ex-
Along with the lungs, the kidneys are partic-
creted with the urine every day. Buffering
ularly involved in keeping the pH value of the
systems in the urine catch a large proportion
extracellular fluid constant (see p. 288). The
of the H+ ions, so that the urine only becomes
contribution made by the kidneys particularly
weakly acidic (down to about pH 4.8).
involves resorbing HCO3- and actively excret-
An important buffer in the urine is the
ing protons.
hydrogen phosphate/dihydrogen phosphate
system (HPO42-/H2PO4-). In addition, ammo-
A. Proton excretion
nia also makes a vital contribution to buffer-
ing the secreted protons.
The renal tubule cells are capable of secreting
Since plasma concentrations of free am-
protons (H+) from the blood into the urine
monia are low, the kidneys release NH3 from
against a concentration gradient, despite the
glutamine and other amino acids. At
fact that the H+ concentration in the urine is
0.5-0.7 mM, glutamine is the most important
up to a thousand times higher than in the
amino acid in the plasma and is the preferred
blood. To achieve this, carbon dioxide (CO2)
form for ammonia transport in the blood. The
is taken up from the blood and—together with
kidneys take up glutamine, and with the help
water (H2O) and with the help of carbonate
of glutaminase [4], initially release NH3 from
dehydratase (carbonic anhydrase, [1])—con-
the amide bond hydrolytically. From the glu-
verted into hydrogen carbonate (“bicarbo-
tamate formed, a second molecule of NH3 can
nate,” HCO3-) and one H+. Formally, this yields
be obtained by oxidative deamination with
carbonic acid H2CO3 as an intermediate, but it
the help of glutamate dehydrogenase [5] (see
is not released during the reaction.
p. 178). The resulting 2-oxoglutarate is fur-
The hydrogen carbonate formed in car-
ther metabolized in the tricarboxylic acid
bonic anhydrase returns to the plasma, where
cycle. Several other amino acids—alanine in
it contributes to the blood’s base reserve. The
particular, as well as serine, glycine, and
proton is exported into the urine by secondary
aspartate—can also serve as suppliers of am-
active transport in antiport for Na+ (bottom
monia.
right). The driving force for proton excretion,
Ammonia can diffuse freely into the urine
as in other secondary active processes, is the
through the tubule membrane, while the am-
Na+ gradient established by the ATPase in-
monium ions that are formed in the urine are
volved in the Na+/K+ exchange (“Na+/K+ AT-
charged and can no longer return to the cell.
Pase”, see p. 220). This integral membrane
Acidic urine therefore promotes ammonia ex-
protein on the basal side (towards the blood)
cretion, which is normally 30-50 mmol per
of tubule cells keeps the Na+ concentration in
day. In metabolic acidosis (e. g., during fasting
the tubule cell low, thereby maintaining Na+
or in diabetes mellitus), after a certain time
inflow. In addition to this secondary active H+
increased induction of glutaminase occurs in
transport mechanism, there is a V-type H+-
the kidneys, resulting in increased NH3 excre-
transporting ATPase in the distal tubule and
tion. This in turn promotes H+ release and
collecting duct (see p. 220).
thus counteracts the acidosis. By contrast,
An important function of the secreted H+
when the plasma pH value shifts towards
ions is to promote HCO3- resorption
(top
alkaline values (alkalosis), renal excretion of
right). Hydrogen carbonate, the most impor-
ammonia is reduced.
tant buffering base in the blood, passes into
the primary urine quantitatively, like all ions.
In the primary urine, HCO3- reacts with H+
ions to form water and CO2, which returns
by free diffusion to the tubule cells and from
there into the blood. In this way, the kidneys
also influence the CO2/HCO3- buffering bal-
ance in the plasma.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Kidney
327
A. Proton secretion
Blood plasma
Urine
From the
Tubule cell
final urine
CO2
CO2
H
H2O
2O
HCO
3
CO
2
Carbonate dehydratase
1
4.2.1.1 [Zn
2
]
1
Na
/K
-exchanging ATPase
2
3.6.1.37
1
H
H
Secondary active
HCO
3
HCO
transport of H
3
A
P
P
P
3Na
3Na
Na
Na
2
2K
2K
A
P P P
B. Ammonia excretion
Glutamine
Glutamine
H2O
Ala
Ser
Amino acid
Gly
with the highest
Diffusion
3
Asp
plasma
concentration
NH3
NH3
Glutamate
Glutamate
H
H
N
A
H2O
NH4
NH4
4
N
A
2-Oxoglutarate
Excretion of
30 - 50 mmol/day
3
Glutaminase 3.5.1.2
Tricarboxylic
Glutamat-
4
acid cycle
Dehydrogenase 1.4.1.2
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
328
Tissues and organs
with glucose and amino acids (see p. 322).
Electrolyte and water recycling
These two pathways are responsible for
60-70% of total Na+ resorption. In the ascend-
A. Electrolyte and water recycling
ing part of Henle’s loop, there is another
Electrolytes and other plasma components
transporter
(shown at the bottom right),
with low molecular weights enter the primary
which functions electroneutrally and takes
urine by ultrafiltration (right). Most of these
up one Na+ ion and one K+ ion together with
substances are recovered by energy-depen-
two Cl- ions. This symport is also dependent
dent resorption (see p. 322). The extent of
on the activity of Na+/K+ ATPase [2], which
the resorption determines the amount that
pumps the Na+ resorbed from the primary
ultimately reaches the final urine and is ex-
urine back into the plasma in exchange for K+.
creted. The illustration does not take into ac-
The steroid hormone aldosterone
(see
count the zoning of transport processes in the
p. 55) increases Na+ reuptake, particularly in
kidney (physiology textbooks may be referred
the distal tubule, while atrial natriuretic
to for further details).
peptide (ANP) originating from the cardiac
Calcium and phosphate ions. Calcium
atrium reduces it. Among other effects, aldo-
(Ca2+) and phosphate ions are almost com-
sterone induces Na+/K+ ATPase and various
pletely resorbed from the primary urine by
Na+ transporters on the luminal side of the
active transport (i. e., in an ATP-dependent
cells.
fashion). The proportion of Ca2+ resorbed is
Water. Water resorption in the proximal
over 99%, while for phosphate the figure is
tubule is a passive process in which water
80-90%. The extent to which these two elec-
follows the osmotically active particles, par-
trolytes are resorbed is regulated by the three
ticularly the Na+ ions. Fine regulation of water
hormones parathyrin, calcitonin, and calci-
excretion (diuresis) takes place in the collect-
triol.
ing ducts, where the peptide hormone vaso-
The peptide hormone parathyrin (PTH),
pressin (antidiuretic hormone, ADH) operates.
which is produced by the parathyroid gland,
This promotes recovery of water by stimulat-
stimulates Ca2+ resorption in the kidneys and
ing the transfer of aquaporins (see p. 220)
at the same time inhibits the resorption of
into the plasma membrane of the tubule cells
phosphate. In conjunction with the effects of
via V2 receptors. A lack of ADH leads to the
this hormone in the bones and intestines (see
disease picture of diabetes insipidus, in which
p. 344), this leads to an increase in the plasma
up to 30 L of final urine is produced per day.
level of Ca2+ and a reduction in the level of
phosphate ions.
B. Gluconeogenesis
Calcitonin, a peptide produced in the C cells
of the thyroid gland, inhibits the resorption of
Apart from the liver, the kidneys are the only
both calcium and phosphate ions. The result is
organs capable of producing glucose by
an overall reduction in the plasma level of both
neosynthesis
(gluconeogenesis; see p. 154).
ions. Calcitonin is thus a parathyrin antago-
The main substrate for gluconeogenesis in
nist relative to Ca2+.
the cells of the proximal tubule is glutamine.
The steroid hormone calcitriol, which is
In addition, other amino acids and also lac-
formed in the kidneys (see p. 304), stimulates
tate, glycerol, and fructose can be used as
the resorption of both calcium and phosphate
precursors. As in the liver, the key enzymes
ions and thus increases the plasma level of
for gluconeogenesis are induced by cortisol
both ions.
(see p. 374). Since the kidneys also have a
Sodium ions. Controlled resorption of Na+
high level of glucose consumption, they only
from the primary urine is one of the most
release very little glucose into the blood.
important functions of the kidney. Na+ resorp-
tion is highly effective, with more than 97%
being resorbed. Several mechanisms are in-
volved: some of the Na+ is taken up passively
in the proximal tubule through the junctions
between the cells (paracellularly). In addition,
there is secondary active transport together
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Kidney
329
A. Electrolyte and water recycling
Blood plasma
Urine
Tubule cell
A
P P P
Ca2
Ca2
Ca2
1
A
Ca2 -transporting
1
P P P
ATPase
3.6.1.38
Na /K -exchanging
2
ATPase 3.6.1.37
Parathyrin
Calcitonin
Calcitriol
H2PO4
H2PO4
H2PO4
H2O
H2O
Water follows
osmotic
ANP
Vasopressin
gradient
(adiuretin)
K
Aldosterone
A
Aldosterone
P P P
3Na
3Na
Na
2
2K
2K
P
A
P P
2 Cl
2 Cl
B. Gluconeogenesis
Glucose
Glucose
Free
Cortisol
Gluconeogenesis
diffusion
Glycerol
Fructose
NH3
NH3
Amino acids
2-Oxoglutarate
H
Glutamine
Glutamine
NH4
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
330
Tissues and organs
the α2-globulin group
(see p. 276), which
Renal hormones
like almost all plasma proteins is synthesized
in the liver. The decapeptide cleaved off by
A. Renal hormones
renin is called angiotensin I. Further cleavage
In addition to their involvement in excretion
by peptidyl dipeptidase A (angiotensin-con-
and metabolism, the kidneys also have endo-
verting enzyme, ACE), a membrane enzyme
crine functions. They produce the hormones
located on the vascular endothelium in the
erythropoietin and calcitriol and play a deci-
lungs and other tissues, gives rise to the
sive part in producing the hormone angioten-
octapeptide angiotensin II [3], which acts as
sin II by releasing the enzyme renin. Renal
a hormone and neurotransmitter. The lifespan
prostaglandins (see p. 390) have a local effect
of angiotensin II in the plasma is only a few
on Na+ resorption.
minutes, as it is rapidly broken down by other
Calcitriol (vitamin D hormone, 1α,25-dihy-
peptidases (angiotensinases [4]), which occur
droxycholecalciferol) is a hormone closely re-
in many different tissues.
lated to the steroids that is involved in Ca2+
The plasma level of angiotensin II is mainly
homeostasis (see p. 342). In the kidney, it is
determined by the rate at which renin is re-
formed from calcidiol by hydroxylation at C-1.
leased by the kidneys. Renin is synthesized by
The activity of calcidiol-1-monooxygenase [1]
juxtaglomerular cells, which release it when
is enhanced by the hormone parathyrin
sodium levels decline or there is a fall in blood
(PTH).
pressure.
Erythropoietin is a peptide hormone that is
Effects of angiotensin II. Angiotensin II has
formed predominantly by the kidneys, but
effects on the kidneys, brain stem, pituitary
also by the liver. Together with other factors
gland, adrenal cortex, blood vessel walls, and
known as “colony-stimulating factors” (CSF;
heart via membrane-located receptors. It in-
see p. 392), it regulates the differentiation of
creases blood pressure by triggering vasocon-
stem cells in the bone marrow.
striction (narrowing of the blood vessels). In
Erythropoietin release is stimulated by hy-
the kidneys, it promotes the retention of Na+
poxia (low pO2). Within hours, the hormone
and water and reduces potassium secretion.
ensures that erythrocyte precursor cells in the
In the brain stem and at nerve endings in the
bone marrow are converted to erythrocytes,
sympathetic nervous system, the effects of
so that their numbers in the blood increase.
angiotensin II lead to increased tonicity (neu-
Renal damage leads to reduced erythropoie-
rotransmitter effect). In addition, it triggers
tin release, which in turn results in anemia.
the sensation of thirst. In the pituitary gland,
Forms of anemia with renal causes can now
angiotensin II stimulates vasopressin release
be successfully treated using erythropoietin
(antidiuretic hormone) and corticotropin
produced by genetic engineering techniques.
(ACTH) release. In the adrenal cortex, it in-
The hormone is also administered to dialysis
creases the biosynthesis and release of aldo-
patients. Among athletes and sports profes-
sterone, which promotes sodium and water
sionals, there have been repeated cases of
retention in the kidneys. All of the effects of
erythropoietin being misused for doping pur-
angiotensin II lead directly or indirectly to
poses.
increased blood pressure, as well as increased
sodium and water retention. This important
hormonal system for blood pressure regula-
B. Renin-angiotensin system
tion can be pharmacologically influenced by
The peptide hormone angiotensin II is not
inhibitors at various points:
synthesized in a hormonal gland, but in the
• Using angiotensinogen analogs that inhibit
blood. The kidneys take part in this process by
renin.
releasing the enzyme renin.
• Using angiotensin I analogs that competi-
Renin [2] is an aspartate proteinase (see
tively inhibit the enzyme ACE [3].
p. 176). It is formed by the kidneys as a pre-
• Using hormone antagonists that block the
cursor
(prorenin), which is proteolytically
binding of angiotensin II to its receptors.
activated into renin and released into the
blood. In the blood plasma, renin acts on
angiotensinogen, a plasma glycoprotein in
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Kidney
331
A. Renal hormones
Differentiation
Maturation
Stem cell
Erythrocyte
Calcidiol 1-monooxygenase
OH
1
[heme P450]1.14.13.13
25
2
Renin 3.4.23.15
Erythropoietin
18.4 kDa
Peptidyl-dipeptidase A
3
[Zn2
]
3.4.15.1 (ACE)
Hormone
CH2
3
1
pO
2
HO
OH
Blood
Angiotensinogen
57 kDa
Prorenin
Calcitriol
Calcitriol
2
Renin
Renin
42 kDa
Parathyrin
1
Enzyme
Digestive
Angiotensin I
tract
Calcidiol
Kidneys
Bones
3
Calcidiol
from liver
Angiotensin II
Hormone
B. Renin angiotensin system
Na
Angiotensinogen
Effects
Blood pressure
from liver
Kidneys:
2
Na retention
O retention
H2
Protein
Residual
protein
Decapeptide
Angiotensin I
Brain:
Release of
corticotropin
ACE inhibitors
3
and adiuretin,
Thirst
Dipeptide
Octapeptide
Angiotensin II
DRVYIHPF
Adrenals:
Aldosterone
Hormone
production
4
Blood vessels:
Vasoconstriction
Blood pressure
4
Peptidases 3.4.n.n
Degradation products
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
332
Tissues and organs
(64 kDa) attaches to F-actin as a rod-like
Muscle contraction
dimer and connects approximately seven ac-
The musculature is what makes movements
tin units with each other. The heterotrimer
possible. In addition to the skeletal muscles,
troponin (78 kDa) is bound to one end of
which can be contracted voluntarily, there are
tropomyosin.
also the autonomically activated heart muscle
In addition to the above proteins, a number
and smooth muscle, which is also involuntary.
of other proteins are also typical of
In all types of muscle, contraction is based on
muscle—including titin (the largest known
an interplay between the proteins actin and
protein), D- and E-actinin, desmin, and
myosin.
vimentin.
A. Organization of skeletal muscle
B. Mechanism of muscle contraction
Striated muscle consists of parallel bundles of
The sliding filament model describes the
muscle fibers. Each fiber is a single large mul-
mechanism involved in muscle contraction.
tinucleate cell. The cytoplasm in these cells
In this model, sarcomeres become shorter
contains myofibrils 2-3 µm thick that can ex-
when the thin and thick filaments slide along-
tend over the full length of the muscle fiber.
side each other and telescope together, with
The striation of the muscle fibers is charac-
ATP being consumed. During contraction, the
teristic of skeletal muscle. It results from the
following reaction cycle is repeated several
regular arrangement of molecules of differing
times:
density. The repeating contractile units, the
sarcomeres, are bounded by Z lines from
[1 ] In the initial state, the myosin heads are
which thin filaments of F-actin (see p. 204)
attached to actin. When ATP is bound, the
extend on each side. In the A bands, there
heads detach themselves from the actin (the
are also thick parallel filaments of myosin.
“plasticizing” effect of ATP).
The H bands in the middle of the A bands
[2 ] The myosin head hydrolyzes the bound
only contain myosin, while only actin is found
ATP to ADP and Pi, but initially withholds the
on each size of the Z lines.
two reaction products. ATP cleavage leads to
Myosin is quantitatively the most impor-
allosteric tension in the myosin head.
tant protein in the myofibrils, representing
[3 ] The myosin head now forms a new
65% of the total. It is shaped like a golf club
bond with a neighboring actin molecule.
(bottom right). The molecule is a hexamer
[4 ] The actin causes the release of the Pi,
consisting of two identical heavy chains
and shortly afterwards release of the ADP as
(2 × 223 kDa) and four light chains
(each
well. This converts the allosteric tension in
about 20 kDa). Each of the two heavy chains
the myosin head into a conformational
has a globular “head” at its amino end, which
change that acts like a rowing stroke.
extends into a “tail” about 150 nm long in
which the two chains are intertwined to
The cycle can be repeated for as long as ATP
form a superhelix. The small subunits are at-
is available, so that the thick filaments are
tached in the head area. Myosin is present as a
constantly moving along the thin filaments
bundle of several hundred stacked molecules
in the direction of the Z disk. Each rowing
in the form of a “thick myosin filament.” The
stroke of the 500 or so myosin heads in a thick
head portion of the molecule acts as an
filament produces a contraction of about
ATPase, the activity of which is modulated
10 nm. During strong contraction, the process
by the small subunits.
is repeated about five times per second. This
Actin (42 kDa) is the most important com-
leads to the whole complex of thin filaments
ponent of the “thin filaments.” It represents ca.
moving together; the H band becomes shorter
20-25% of the muscle proteins. F-actin is also
and the Z lines slide closer together.
an important component of the cytoskeleton
(see p. 204). This filamentous polymer is held
in equilibrium with its monomer, G-actin. The
other protein components of muscle include
tropomyosin and troponin. Tropomyosin
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Muscle
333
A. Organization of striated muscle
Muscle
Tendon
Bundle of muscle fibers
Nucleus
Sarcomere
Z
Z
Z
Muscle fiber
H zone
A band
I band
Z
H
Z line
Myofibril
Troponin
Actin
Tropomyosin
Thin actin filament
NH3
150 nm
Small subunits
OOC
OOC
Globular head
Thick myosin filament
Myosin molecule
NH3
B. Mechanism of muscle contraction
Thin
Center of the
filament
sarcomere
Thick
filament
P
5
Myosin head relaxed
1
A
P P
Actin
Power stroke
A
P P P
ATP
Myosin
H2O
4
2
Contact
Myosin head contracted
3
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
334
Tissues and organs
enter the sarcoplasm, where they lead to a
Control of muscle contraction
rapid increase in Ca2+ concentrations. This in
turn causes the myofibrils to contract (C).
A. Neuromuscular junction
Muscle contraction is triggered by motor
C. Regulation by calcium ions
neurons that release the neurotransmitter
acetylcholine (see p. 352). The transmitter dif-
In relaxed skeletal muscle, the complex con-
fuses through the narrow synaptic cleft and
sisting of troponin and tropomyosin blocks
binds to nicotinic acetylcholine receptors on
the access of the myosin heads to actin (see
the plasma membrane of the muscle cell
p. 332). Troponin consists of three different
(the sarcolemma), thereby opening the ion
subunits (T, C, and I). The rapid increase in
channels integrated into the receptors (see
cytoplasmic Ca2+ concentrations caused by
p. 222). This leads to an inflow of Na+, which
opening of the calcium channels in the SR
triggers an action potential (see p. 350) in the
leads to binding of Ca2+ to the C subunit of
sarcolemma. The action potential propagates
troponin, which closely resembles calmodulin
from the end plate in all directions and con-
(see p. 386). This produces a conformational
stantly stimulates the muscle fiber. With a
change in troponin that causes the whole tro-
delay of a few milliseconds, the contractile
ponin-tropomyosin complex to slip slightly
mechanism responds to this by contracting
and expose a binding site for myosin (red).
the muscle fiber.
This initiates the contraction cycle. After con-
traction, the sarcoplasmic Ca2+ concentration
is quickly reduced again by active transport
B. Sarcoplasmic reticulum (SR)
back into the SR. This results in troponin los-
The action potential (A) produced at the neu-
ing the bound Ca2+ ions and returning to the
romuscular junction is transferred in the
initial state, in which the binding site for my-
muscle cell into a transient increase in the
osin on actin is blocked. It is not yet clear
Ca2+ concentration in the cytoplasm of the
whether the mechanism described above is
muscle fiber (the sarcoplasm).
the only one that triggers binding of myosin
In the resting state, the Ca2+ level in the
to actin.
sarcoplasm is very low (less than 10-7 M). By
When triggering of contraction in striated
contrast, the sarcoplasmic reticulum
(SR),
muscle occurs, the following sequence of
which corresponds to the ER, contains Ca2+
processes thus takes place:
ions at a concentration of about 10-3 M. The
1. The sarcolemma is depolarized.
SR is a branched organelle that surrounds the
2. The action potential is signaled to Ca2+
myofibrils like a net stocking inside the
channels in the SR.
muscle fibers (illustrated at the top using
3. The Ca2+ channels open and the Ca2+ level
the example of a heart muscle cell). The high
in the sarcoplasm increases.
Ca2+ level in the SR is maintained by Ca2+-
4. Ca2+ binds to troponin C and triggers a
transporting ATPases (see p. 220). In addition,
conformational change.
the SR also contains calsequestrin, a protein
5. Troponin causes tropomyosin to slip, and
(55 kDa) that is able to bind numerous Ca2+ -
the myosin heads bind to actin.
ions via acidic amino acid residues.
6. The actin-myosin cycle takes place and the
The transfer of the action potential to the SR
muscle fibers contract.
is made possible by transverse tubules (T tu-
bules), which are open to the extracellular space
Conversely, at the end of contraction, the fol-
and establish a close connection with the SR.
lowing processes take place:
There is a structure involved in the contact be-
1. The Ca2+ level in the sarcoplasm declines
tween the T tubule and the SR that was formerly
due to transport of Ca2+ back into the SR.
known as the “SR foot” (it involves parts of the
2. Troponin C loses Ca2+ and tropomyosin re-
ryanodine receptor; see p. 386).
turns to its original position on the actin
At the point of contact with the SR, the
molecule.
action potential triggers the opening of the
3. The actin-myosin cycle stops and the
Ca2+ channels on the surface of the sarco-
muscle relaxes.
lemma. Calcium ions then leave the SR and
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Muscle
335
A. Neuromuscular junction
Motoneuron
Action potential
End plate
Nucleus
Muscle fiber
B. Sarcoplasmatic reticulum (SR)
Transverse tubule
Sarcoplasmic reticulum
Extracellular
Depolarization
space
of membrane
causes
Ca2
A
A
opening of
Cytoplasm
P P
P P P
P
Ca2
-channels
SR
T tubule
Ca2
=10-3M
SR foot
Calsequestrin
A
A
P
A
A
P
Ca2
<10-7M
Ca2 channel
P P P
P P
P P P
P P
C. Regulation by calcium ions
Troponin
Ca2
Ca2
Myosin-
exposes
binding
Ca2
myosin-
T
T
site on
binding
actin
C
site on
I
blocked
I
actin
Actin
Tropomyosin
Myosin head
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
336
Tissues and organs
maintain glucose degradation and thus ATP
Muscle metabolism I
formation. If there is a lack of O2, this is
Muscle contraction is associated with a high
achieved by the formation of lactate, which
level of ATP consumption (see p. 332). With-
is released into the blood and is resynthesized
out constant resynthesis, the amount of ATP
into glucose in the liver
(Cori cycle; see
available in the resting state would be used up
p. 338).
in less than 1 s of contraction.
Muscle-specific auxiliary reactions for ATP
synthesis exist in order to provide additional
ATP in case of emergency. Creatine phosphate
A. Energy metabolism in the white and red
(see B) acts as a buffer for the ATP level.
muscle fibers
Another ATP-supplying reaction is catalyzed
Muscles contain two types of fibers, the pro-
by adenylate kinase [1] (see also p. 72). This
portions of which vary from one type of
disproportionates two molecules of ADP into
muscle to another. Red fibers (type I fibers)
ATP and AMP. The AMP is deaminated into
are suitable for prolonged effort. Their metab-
IMP in a subsequent reaction [2] in order to
olism is mainly aerobic and therefore depends
shift the balance of the reversible reaction [1]
on an adequate supply of O2. White fibers
in the direction of ATP formation.
(type II fibers) are better suited for fast, strong
contractions. These fibers are able to form
B. Creatine metabolism
suf cient ATP even when there is little O2
available. With appropriate training, athletes
Creatine (N-methylguanidoacetic acid) and its
and sports participants are able to change the
phosphorylated form creatine phosphate
proportions of the two fiber types in the mus-
(a guanidophosphate) serve as an ATP buffer
culature and thereby prepare themselves for
in muscle metabolism. In creatine phosphate,
the physiological demands of their disciplines
the phosphate residue is at a similarly high
in a targeted fashion. The expression of func-
chemical potential as in ATP and is therefore
tional muscle proteins can also change during
easily transferred to ADP. Conversely, when
the course of training.
there is an excess of ATP, creatine phosphate
Red fibers provide for their ATP require-
can arise from ATP and creatine. Both proce-
ments mainly (but not exclusively) from fatty
sses are catalyzed by creatine kinase [5].
acids, which are broken down via β-oxidation,
In resting muscle, creatine phosphate
the tricarboxylic acid cycle, and the respira-
forms due to the high level of ATP. If there is
tory chain (right part of the illustration). The
a risk of a severe drop in the ATP level during
red color in these fibers is due to the mono-
contraction, the level can be maintained for a
meric heme protein myoglobin, which they
short time by synthesis of ATP from creatine
use as an O2 reserve. Myoglobin has a much
phosphate and ADP. In a nonenzymatic reac-
higher af nity for O2 than hemoglobin and
tion [6], small amounts of creatine and crea-
therefore only releases its O2 when there is
tine phosphate cyclize constantly to form cre-
a severe drop in O2
partial pressure
atinine, which can no longer be phosphory-
(cf. p. 282).
lated and is therefore excreted with the urine
At a high level of muscular effort—e. g.,
(see p. 324).
during weightlifting or in very fast contrac-
Creatine does not derive from the muscles
tions such as those carried out by the eye
themselves, but is synthesized in two steps in
muscles—the O2
supply from the blood
the kidneys and liver (left part of the illustra-
quickly becomes inadequate to maintain the
tion). Initially, the guanidino group of argi-
aerobic metabolism. White fibers (left part of
nine is transferred to glycine in the kidneys,
the illustration) therefore mainly obtain ATP
yielding guanidino acetate [3]. In the liver,
from anaerobic glycolysis. They have supplies
N-methylation of guanidino acetate leads to
of glycogen from which they can quickly re-
the formation of creatine from this [4]. The
lease glucose-1-phosphate when needed (see
coenzyme in this reaction is S-adenosyl methi-
p. 156). By isomerization, this gives rise to
onine (SAM; see p. 110).
glucose-6-phosphate, the substrate for glycol-
ysis. The NADH+H+ formed during glycolysis
has to be reoxidized into NAD+ in order to
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Muscle
337
A. Energy metabolism in the white and red muscle fibers
White (slow) fibers, anaerobic
Red (fast) fibers, aerobic
Creatine
Glycogen
ADP
ADP
Myoglobin
Fatty acids
phosphate
2+
Glucose
Glucose
O2
β-Oxidation
Creatine
NAD+
Ketone
N
A
bodies
A
ATP
2+
H2O
P P P
Cori cycle
Glycolysis
Acetyl CoA
(liver)
Contraction
NADH + H+
O2
N
A
ADP
A
P
P
Respiratory
chain
N
A
Tricarboxylic
Lactate
Pyruvate
acid cycle
1
NADH + H+
Adenylate kinase
1
A
I
2.7.4.3
2
P
P
2
AMP deaminase
CO2
3.5.4.6
AMP
NH3
IMP
CO2
B. Creatine metabolism
Muscle
S-adenosyl
homocysteine
A
Liver
A
P
P
P
P
P
S-adenosyl
NH2
methionine
H3C
C
N
NH2
H2C
O
O
4
5
C
O
P
OH
O
HN
Creatine
H
3
C
C
N
NH
Guanidino
Creatine
Creatine
Creatine
acetate
phosphate
H2C
O
C
O
Blood
6
6
Creatinine
Glycine
Guanidino-
3
amidinotransferase
acetate
2.1.4.1
Orn
H3C
NH2
NH2
N
C
Guanidinoacetate
3
4
methyltransferase
H2C
N
H
C
2.1.1.2
NH
C
N
2
Arg
Creatine
H2C
O
O
5
kinase 2.7.3.2
C
Glycine
Not-
O
6
enzymatic
Kidney
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
338
Tissues and organs
B. Protein and amino acid metabolism
Muscle metabolism II
The skeletal muscle is the most important site
A. Cori and alanine cycle
for degradation of the branched-chain amino
acids (Val, Leu, Ile; see p. 414), but other amino
White muscle fibers (see p. 336) mainly ob-
acids are also broken down in the muscles.
tain ATP from anaerobic glycolysis—i. e., they
Alanine and glutamine are resynthesized
convert glucose into lactate. The lactate aris-
from the components and released into the
ing in muscle and, in smaller quantities, its
blood. They transport the nitrogen that arises
precursor pyruvate are released into the
during amino acid breakdown to the liver
blood and transported to the liver, where lac-
(alanine cycle; see above) and to the kidneys
tate and pyruvate are resynthesized into glu-
(see p. 328).
cose again via gluconeogenesis, with ATP being
During periods of hunger, muscle proteins
consumed in the process (see p. 154). The
serve as an energy reserve for the body. They
glucose newly formed by the liver returns
are broken down into amino acids, which are
via the blood to the muscles, where it can be
transported to the liver. In the liver, the car-
used as an energy source again. This circula-
bon skeletons of the amino acids are con-
tion system is called the Cori cycle, after the
verted into intermediates in the tricarboxylic
researchers who first discovered it. There is
acid cycle or into acetoacetyl-CoA (see p. 175).
also a very similar cycle for erythrocytes,
These amphibolic metabolites are then avail-
which do not have mitochondria and there-
able to the energy metabolism and for gluco-
fore produce ATP by anaerobic glycolysis (see
neogenesis. After prolonged starvation, the
p. 284).
brain switches to using ketone bodies in order
The muscles themselves are not capable of
to save muscle protein (see p. 356).
gluconeogenesis. Nor would this be useful, as
The synthesis and degradation of muscle
gluconeogenesis requires much more ATP
proteins are regulated by hormones. Cortisol
than is supplied by glycolysis. As O2 deficien-
leads to muscle degradation, while testo-
cies do not arise in the liver even during in-
sterone stimulates protein formation. Syn-
tensive muscle work, there is always suf -
thetic anabolics with a testosterone-like ef-
cient energy there available for gluconeogen-
fect have repeatedly been used for doping
esis.
purposes or for intensive muscle-building.
There is also a corresponding circulation
system for the amino acid alanine. The alanine
cycle in the liver not only provides alanine as
Further information
a precursor for gluconeogenesis, but also
transports to the liver the amino nitrogen
Smooth muscle differs from skeletal muscle in
arising in muscles during protein degrada-
various ways. Smooth muscles—which are
tion. In the liver, it is incorporated into urea
found, for example, in blood vessel walls
for excretion.
and in the walls of the intestines—do not
Most of the amino acids that arise in
contain any muscle fibers. In smooth-muscle
muscle during proteolysis are converted into
cells, which are usually spindle-shaped, the
glutamate and 2-oxo acids by transamination
contractile proteins are arranged in a less reg-
(not shown; cf. p. 180). Again by transamina-
ular pattern than in striated muscle. Contrac-
tion, glutamate and pyruvate give rise to ala-
tion in this type of muscle is usually not
nine, which after glutamine is the second im-
stimulated by nerve impulses, but occurs in
portant form of transport for amino nitrogen
a largely spontaneous way. Ca2+ (in the form
in the blood. In the liver, alanine and 2-oxo-
of Ca2+-calmodulin; see p. 386) also activates
glutarate are resynthesized into pyruvate and
contraction in smooth muscle; in this case,
glutamate (see p. 178). Glutamate supplies
however, it does not affect troponin, but acti-
the urea cycle (see p. 182), while pyruvate is
vates a protein kinase that phosphorylates the
available for gluconeogenesis.
light chains in myosin and thereby increases
myosin’s ATPase activity. Hormones such as
epinephrine and angiotensin II (see p. 330)
are able to influence vascular tonicity in this
way, for example.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Muscle
339
A. Cori and alanine cycle
Liver
Blood
Muscle
Storage
Glucose
Glucose
Glucose
3.70-5.18
mM
Glycogen
Glucose 6
P
Glucose 6
P
Glycogen
4
1
6 ATP
2 ATP
Pyruvate
Pyruvate
Pyruvate
Protein
0.02-0.07
N
A
N
A
mM
3
3
N
A
N
A
Amino
acids
Lactate
Lactate
Lactate
0.74-2.40
Urea
[NH2]
mM
2
2
Alanine
Alanine
Alanine
0.33-0.61
mM
NADH/NAD+
NADH/NAD+
quotient low
quotient high
1
Glycolysis
Alanine cycle
Plasma concentration
2
Transamination
in adults
Cori cycle
Lactate dehydrogenase
3
1.1.1.27
4
Gluconeogenesis
B. Protein and amino acid metabolism
Blood
Muscle
Muscle breaks
Energy
down
reserve in
branched-chain
prolonged
amino acids
Protein
hunger
Testosterone
Anabolic
Cortisol
agents
Val
Leu
Ile
0.25
0.16
0.08
Amino acids
mM
mM
mM
Ala
Gln
Amino acids
[NH2]
0.42
0.65
Ala, Gln
mM
mM
Synthesis
Degradation
Tri-
carboxylic
Energy gain
acid
Plasma
cycle
concentration
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
340
Tissues and organs
B. Teeth
Bone and teeth
The illustration shows a longitudinal section
The family of connective-tissue cells includes
through an incisor, one of the 32 permanent
fibroblasts, chondrocytes (cartilage cells), and
teeth in humans. The majority of the tooth
osteoblasts (bone-forming cells). They are spe-
consists of dentine. The crown of the tooth
cialized to secrete extracellular proteins, par-
extends beyond the gums, and it is covered in
ticularly collagens, and mineral substances,
enamel. By contrast, the root of the tooth is
which they use to build up the extracellular
coated in dental cement.
matrix (see p. 346). By contrast, osteoclasts
Cement, dentin, and enamel are bone-like
dissolve bone matter again by secreting H+
substances. The high proportion of inorganic
and collagenases (see p. 342).
matter they contain (about 97% in the dental
enamel) gives them their characteristic hard-
A. Bone
ness. The organic components of cement,
dentin, and enamel mainly consist of collagens
Bone is an extremely dense, specialized form
and proteoglycans; their most important min-
of connective tissue. In addition to its suppor-
eral component is apatite, as in bone (see
tive function, it serves to store calcium and
above).
phosphate ions. In addition, blood cells are
A widespread form of dental disease, ca-
formed in the bone marrow. The most impor-
ries, is caused by acids that dissolve the min-
tant mineral component of bone is apatite, a
eral part of the teeth by neutralizing the neg-
form of crystalline calcium phosphate.
atively charged counter-ions in apatite (see
Apatites are complexes of cationic Ca2+
A). Acids occur in food, or are produced by
matched by HPO42-, CO32-, OH-, or F- as
microorganisms that live on the surfaces of
anions. Depending on the counter-ion, apatite
the teeth (e. g., Streptococcus mutans).
can occur in the forms carbonate apatite
The main product of anaerobic degradation
Ca10(PO4)6CO3, as hydroxyapatite Ca10(PO4)6
of sugars by these organisms is lactic acid.
(OH)2, or fluoroapatite Ca10(PO4)6F2. In addi-
Other products of bacterial carbohydrate me-
tion, alkaline earth carbonates also occur in
tabolism include extracellular dextrans (see
bone. In adults, more than 1 kg calcium is
p. 40)—insoluble polymers of glucose that
stored in bone.
help bacteria to protect themselves from their
Osteoblast and osteoclast activity is con-
environment. Bacteria and dextrans are com-
stantly incorporating Ca2+ into bone and re-
ponents of dental plaque, which forms on in-
moving it again. There are various hormones
adequately cleaned teeth. When Ca2+ salts
that regulate these processes: calcitonin in-
and other minerals are deposited in plaque
creases deposition of Ca2+ in the bone matrix,
as well, tartar is formed.
while parathyroid hormone (PTH) promotes
The most important form of protection
the mobilization of Ca2+, and calcitriol im-
against caries involves avoiding sweet sub-
proves mineralization (for details, see p. 342).
stances (foods containing saccharose, glucose,
The most important organic components of
and fructose). Small children in particular
bone are collagens (mainly type I; see p. 344)
should not have very sweet drinks freely
and proteoglycans (see p. 346). These form
available to them. Regular removal of plaque
the extracellular matrix into which the apa-
by cleaning the teeth and hardening of the
tite crystals are deposited (biomineralization).
dental enamel by fluoridization are also im-
Various proteins are involved in this not yet
portant. Fluoride has a protective effect be-
fully understood process of bone formation,
cause fluoroapatite (see A) is particularly re-
including collagens and phosphatases. Alka-
sistant to acids.
line phosphatase is found in osteoblasts and
acid phosphatase in osteoclasts. Both of these
enzymes serve as marker enzymes for bone
cells.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Connective tissue
341
A. Bone
Functions:
Composition:
Mechanical support
Inorganic
Organic
Storage for Ca2
Apatite
Type I collagen
and phosphate
Carbonate
Proteoglycans
Synthesis of blood cells
Water
Phosphatases
Maturation of B cells
Hormones
Hydroxyapatite Ca10(PO4)6(OH)2
of the calcium
metabolism
Parathyroid hormone
Ca2
in plasma
Ca2
Calcitonin
O
Ca
Absence leads
to rickets
Calcitriol
P
OH
B. Teeth
Dental enamel
97%
Hardest substance
Dentin
70%
Dextrans
in the body
Cement
65%
Jawbone
45%
Plaque
Crown
Dental
Inorganic
components
plaque
Dental
Bacterium
enamel
Neck
Sugar
Attack
Dental
Protect
apatite
enamel
bacteria
Saccharose
Glucose
Fructose
Anaerobic
Dextrans
Acids
metabolism
Lactic acid
Propionic acid
Acetic acid
Bacterium
Root
Butyric acid
E.g.,
Streptococcus
De-
Calcium
mineral-
mutans
salts
ization
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
342
Tissues and organs
tion in estrogen levels. Estrogens normally
Calcium metabolism
inhibit the stimulation of osteoclast differen-
tiation by osteoblasts. If the effects of estrogen
A. Functions of calcium
decline, the osteoclasts predominate and ex-
The human body contains 1-1.5 kg Ca2+, most
cess bone removal occurs.
of which (about 98%) is located in the mineral
The effects of the steroid hormone calcitriol
substance of bone (see p. 362).
(see p. 330) in bone are complex. On the one
In addition to its role as a bone component,
hand, it promotes bone formation by stimu-
calcium functions as a signaling substance.
lating osteoblast differentiation (top). This is
Ca2+ ions act as second messengers in signal
particularly important in small children, in
transduction pathways
(see p. 386), they
whom calcitriol deficiency can lead to miner-
trigger exocytosis
(see p. 228) and muscle
alization disturbances (rickets; see p. 364). On
contraction (see p. 334), and they are indis-
the other hand, calcitriol increases blood Ca2+
pensable as cofactors in blood coagulation (see
levels through increased Ca2+ mobilization
p. 290). Many enzymes also require Ca2+ for
from bone. An overdose of vitamin D (chole-
their activity. The intracellular and extracel-
calciferol), the precursor of calcitriol, can
lular concentrations of Ca2+ are strictly regu-
therefore have unfavorable effects on the
lated in order to make these functions possi-
skeleton similar to those of vitamin deficiency
ble (see B, C, and p. 388).
(hypervitaminosis; see p. 364).
Proteins bind Ca2+ via oxygen ligands, par-
ticularly carboxylate groups and carbonyl
C. Calcium homeostasis
groups of peptide bonds. This also applies to
the structure illustrated here, in which a Ca2+
Ca2+ metabolism is balanced in healthy adults.
ion is coordinated by the oxygen atoms of
Approximately 1g Ca2+ is taken up per day,
carboxylate and acid amide groups.
about 300 mg of which is resorbed. The same
amount is also excreted again. The amounts of
Ca2+ released from bone and deposited in it
B. Bone remodeling
per day are much smaller. Milk and milk prod-
Deposition of Ca2+ in bone (mineralization)
ucts, especially cheese, are particularly rich in
and Ca2+ mobilization from bone are regu-
calcium.
lated by at least 15 hormones and hormone-
Calcitriol and parathyroid hormone, on the
like signaling substances. These mainly influ-
one hand, and calcitonin on the other, ensure
ence the maturation and activity of bone cells.
a more or less constant level of Ca2+ in the
Osteoblasts (top) deposit collagen, as well
blood plasma and in the extracellular space
as Ca2+ and phosphate, and thereby create
(80-110 mg
2.0-2.6 mM). The peptide para-
new bone matter, while osteoclasts (bottom)
thyroid hormone (PTH; 84 AA) and the steroid
secrete H+ ions and collagenases that locally
calcitriol (see p. 374) promote direct or indi-
dissolve bone (bone remodeling). Osteoblasts
rect processes that raise the Ca2+ level in
and osteoclasts mutually activate each other
blood. Calcitriol increases Ca2+ resorption in
by releasing cytokines
(see p. 392) and
the intestines and kidneys by inducing trans-
growth factors. This helps keep bone forma-
porters. Parathyroid hormone supports these
tion and bone breakdown in balance.
processes by stimulating calcitriol biosynthe-
The Ca2+-selective hormones calcitriol, par-
sis in the kidneys (see p. 330). In addition, it
athyroid hormone, and calcitonin influence
directly promotes resorption of Ca2+ in the
this interaction in the bone cells. Parathyroid
kidneys (see p. 328) and Ca2+ release from
hormone promotes Ca2+ release by promoting
bone (see B). The PTH antagonist calcitonin
the release of cytokines by osteoblasts. In
(32 AA) counteracts these processes.
turn, the cytokines stimulate the develop-
ment of mature osteoclasts from precursor
cells (bottom). Calcitonin inhibits this process.
At the same time, it promotes the develop-
ment of osteoblasts (top). Osteoporosis, which
mainly occurs in women following the men-
opause, is based (at least in part) on a reduc-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Connective tissue
343
A. Functions of calcium
B. Bone remodeling
Osteoblast
Calcitriol
Calcitonin
Mineralization
Ca2
Muscle
Precursor
con-
cells
traction
Parathyroid
Signal
Hydroxy-
Collagen I
hormone
apatite
transduction
Ca2
Growth
Estrogens
factors
Nerve
H
conduction
Cyto-
Exocytosis
kines
H
Precursor
Protein
cells
function
HCO3
H2O
CO2
Ca2
complex
Calcitonin
Osteoclast
C. Calcium homeostasis
Calci-
600 - 900mg
1000000 mg
Stock
Calcitriol
tonin
1000 mg · d-1
oder
Hydroxy-
Parathyroid hormone
500 mg · d-1
apatite
500 mg · d-1
Calcitriol
Bone
Calci-
Parathyroid
tonin
hormone
Filtration
Kidney
300mg · d-1
till 150mg · d-1
10 000 mg · d-1
9 850 mg · d-1
Resorption
Calci-
Parathyroid
850mg · d-1
triol
hormone
Intestine
Blood
150
mg·d-1
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
344
Tissues and organs
are seen to have a characteristic banding pat-
Collagens
tern of elements that are repeated every
Collagens are quantitatively the most abun-
64-67 nm.
dant of animal proteins, representing 25 % of
Tropocollagen molecules are firmly linked
the total. They form insoluble tensile fibers
together, particularly at their ends, by cova-
that occur as structural elements of the ex-
lent networks of altered lysine side chains.
tracellular matrix and connective tissue
The number of these links increases with
throughout the body. Their name (which lit-
age. Type IV collagens form networks with a
erally means
“glue-producers”) is derived
defined mesh size. The size-selective filtering
from the gelatins that appear as a decompo-
effect of the basal membranes in the renal
sition product when collagen is boiled.
glomeruli is based on this type of structure
(see p. 322).
A. Structure of collagens
B. Biosynthesis
Nineteen different collagens are now known,
and they are distinguished using roman nu-
The precursor molecule of collagen (prepro-
merals. They mostly consist of a dextro-
collagen), formed in the rER, is subject to
rotatory triple helix made up of three poly-
extensive post-translational modifications
peptides (α-chains) (see p. 70).
(see p. 232) in the ER and Golgi apparatus.
The triplet Gly-X-Y is constantly repeated in
Cleavage of the signal peptide gives rise to
the sequence of the triple-helical regions—
procollagen, which still carries large propep-
i. e., every third amino acid in such sequences
tides at each end [1]. During this phase, most
is a glycine. Proline (Pro) is frequently found in
proline residues and some lysine residues of
positions X or Y; the Y position is often occu-
procollagen are hydroxylated [2]. The procol-
pied by 4-hydroxyproline (4Hyp), although
lagen is then glycosylated at hydroxylysine
3-hydroxyproline (3Hyp) and 5-hydroxylysine
residues [3]. Intramolecular and intermolecu-
(5Hyl) also occur. These hydroxylated amino
lar disulfide bonds form in the propeptides
acids are characteristic components of colla-
[4], allowing correct positioning of the pep-
gen. They are only produced after protein
tide strands to form a triple helix [5]. It is only
biosynthesis by hydroxylation of the amino
after these steps have been completed that
acids in the peptide chain (see p. 62).
procollagen is secreted into the extracellular
The formation of Hyp and Hyl residues in
space by exocytosis. This is where the N- and
procollagen is catalyzed by iron-containing
C-terminal propeptides are removed proteo-
oxygenases
(“proline and lysine hydrox-
lytically [6], allowing the staggered aggrega-
ylase,” EC 1.14.11.1/2). Ascorbate is required
tion of the tropocollagen molecules to form
to maintain their function. Most of the symp-
fibrils [7]. Finally, several ε-amino groups in
toms of the vitamin C deficiency disease
lysine residues are oxidatively converted into
scurvy (see p. 368) are explained by disturbed
aldehyde groups [8]. Covalent links between
collagen biosynthesis.
the molecules then form as a result of con-
The hydroxyproline residues stabilize the
densation [9]. In this way, the fibrils reach
triple helix by forming hydrogen bonds be-
their final structure, which is characterized
tween the α-chains, while the hydroxyl
by its high tensile strength and proteinase re-
groups of hydroxylysine are partly glycosy-
sistance.
lated with a disaccharide (-Glc-Gal).
The various types of collagen consist of
different combinations of α-chains (α1 to α3
and other subtypes). Types I, II, and III repre-
sent 90% of collagens. The type I collagen
shown here has the structure [α1(I)]2α2(1).
Numerous tropocollagen molecules (mass
285 kDa, length 400 nm) aggregate extracell-
ularly into a defined arrangement, forming
cylindrical fibrils
(20-500 nm in diameter).
Under the electron microscope, these fibrils
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Connective tissue
345
A. Structure of collagens
B. Biosynthesis
Special amino acids:
Gene
4-Hydroxyproline (4Hyp)
3-Hydroxyproline (3Hyp)
Transcription
5-Hydroxylysine (5Hyl)
– Gly - X - Y - Gly
- X
-
Y - Gly - X - Y -
Basic unit
Translation
Primary structure
Pre-procollagen
O
C
OH
1
Hyl
C CH
CH2
C CH2
NH3
2
2
1
2
HN
H
3
C O
4
Intracellular
H2
HO C
Hyp = Y
5
protein
CH
modification
C
N
H
C
H2
C O
Exocytosis
H2
C
Pro = X
CH
H2C
N
C
Procollagen
H2
C O
6
Gly
H2C
7
NH
8
3
Extracellular
protein
9
modification
Type I tropocollagen
~285 kDa
Collagen fibril
1 Removal of the prepeptide
Overlap
Gap
Diameter
2
Hydroxylation of Pro and Lys residues
D=64-67 nm
40 nm
1.5 nm
3 Glycosylation of 5Hyl and Asn
4
Oxidation of Cys in propeptides
5 Assemblage to form triple helix
6
Removal of the propeptide
7
Staggered deposition to form fibrils
Length 400 nm
8 Oxidation of Lys and 5Hyl to aldehydes
9 Cross-linking to form supramolecules
Procollagen-proline 4-dioxygenase 1.14.11.2
1
[ascorbate, Fe]
Procollagen-lysine 5-dioxygenase 1.14.11.4
2
[ascorbate, Fe]
Collagen fibril (section)
3
Protein-lysine 6-oxidase 1.4.3.13 [Cu]
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
346
Tissues and organs
B. Fibronectins
Extracellular matrix
Fibronectins are typical representatives of
A. Extracellular matrix
adhesive proteins. They are filamentous
dimers consisting of two related peptide
The space between the cells (the interstitium)
chains (each with a mass of 250 kDa) linked
is occupied by a substance with a complex
to each other by disulfide bonds. The fibro-
composition known as the extracellular
nectin molecules are divided into different
matrix (ECM). In many types of tissue—e. g.,
domains, which bind to cell-surface receptors,
muscle and liver—the ECM is only a narrow
collagens, fibrin, and various proteoglycans.
border between the cells, while in others it
This is what gives fibronectins their “molec-
forms a larger space. In connective tissue,
ular glue” characteristics.
cartilage, and bone, the ECM is particularly
The domain structure in fibronectins is
strongly marked and is actually the functional
made up of a few types of peptide module
part of the tissue (see p. 340). The illustration
that are repeated numerous times. Each of
shows the three main constituents of the ex-
the more than 50 modules is coded for by
tracellular matrix in a highly schematic way:
one exon in the fibronectin gene. Alternative
collagen fibers, network-forming adhesive
splicing (see p. 246) of the hnRNA transcript of
proteins, and space-filling proteoglycans.
the fibronectin gene leads to fibronectins with
The ECM has a very wide variety of func-
different compositions. The module that cau-
tions: it establishes mechanical connections
ses adhesion to cells contains the characteristic
between cells; it creates structures with spe-
amino acid sequence -Arg-Gly-Asp-Ser-. It is
cial mechanical properties (as in bone, carti-
these residues that enable fibronectin to bind
lage, tendons, and joints); it creates filters
to cell-surface receptors, known as integrins.
(e. g., in the basal membrane in the renal cor-
puscles; see p. 322); it separates cells and tis-
sues from each other (e. g., to allow the joints
C. Proteoglycans
to move freely); and it provides pathways to
guide migratory cells (important for embry-
Proteoglycans are giant molecule complexes
onic development). The chemical composi-
consisting of carbohydrates (95%) and pro-
tion of the ECM is just as diverse as its func-
teins (5%), with masses of up to 2
106 Da.
tions.
Their bottlebrush-shaped structure is pro-
Collagens (see p. 344), of which there are
duced by an axis consisting of hyaluronate.
at least 19 different varieties, form fibers, fi-
This thread-like polysaccharide
(see p. 44)
brils, networks, and ligaments. Their charac-
has proteins attached to it, from which in
teristic properties are tensile strength and
turn long polysaccharide chains emerge. Like
flexibility. Elastin is a fiber protein with a
the central hyaluronate, these terminal poly-
high degree of elasticity.
saccharides belong to the glycosaminoglycan
Adhesive proteins provide the connections
group (see p. 44).
between the various components of the ex-
The glycosaminoglycans are made up of
tracellular matrix. Important representatives
repeating disaccharide units, each of which
include laminin and fibronectin (see B). These
consists of one uronic acid (glucuronic acid
multifunctional proteins simultaneously bind
or iduronic acid) and one amino sugar (N-
to several other types of matrix component.
acetylglucosamine or N-acetylgalactosamine)
Cells attach to the cell surface receptors in the
(see p. 38). Many of the amino sugars are also
ECM with the help of the adhesive proteins.
esterified with sulfuric acid (sulfated), further
Due to their polarity and negative charge,
increasing their polarity. The proteoglycans
proteoglycans (see C) bind water molecules
bind large amounts of water and fill the
and cations. As a homogeneous “cement,”
gaps between the fibrillar components of
they fill the gaps between the ECM fibers.
the ECM in the form of a hydrated gel. This
inhibits the spread of pathogens in the ECM,
for example.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Connective tissue
347
A. Extracellular matrix
Cell membrane
Collagens
Adhesive
(at least
proteins:
19 different
Elastin, laminin,
types)
fibronectin
Cell surface
Proteoglycans
receptors
and
(Integrins)
hyaluronic acid
B. Fibronectins
Heparan sulfate
Fibrin
Collagens
Integrins
Heparane sulfate Fibrin
Domains
N
C
ss
ss
N
C
-Arg-Gly-Asp-Ser-
Peptide modules
Variable peptides
C. Proteoglycans
COO
CH2OH
Disaccharide units
100 nm
H
O
H
O
O H
O
- Uronic acid - Amino sugar -
OH H
H
H
CH2OH
O3S
H HO
H
H
O
O
O
COO
O H
O
H
OH
H
HN
C
CH3
OH
H
H
Dermatan
Hyaluronate
O
H
H
H
sulfate
H
OH
H
HN
C CH3
O
IduUA
GalNAc
Core protein
COO
H2C
O SO3
Glycosamino-
H
O
H
H
O
H
glycans
OH
H
OH H
Heparin
O
O
H
O
SO
3
H
HN
SO
3
GlcUA
GlcNAc
COO
H2C
O SO3
HO
O
H
O
Keratan
H
O
OH
H
O
20 - 40
sulfate
H
H
H
Disaccharide
H
OH
H
HN
C CH3
units
O
GalUA
GlcNAc
IduUA
= Iduronate
COO
H2C
O
SO3
GlcUA
= Glucuronate
H
O
HO
O
O H
O
Chondroitin
Ribosome
GalNAc
= N-Acetyl-
OH H
H
6-sulfate
for
= galactosamine
H
H
H
comparison
H
OH
H
HN
C CH3
GlcNAc
= N-Acetyl-
= glucosamine
GlcUA
GalNAcO
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
348
Tissues and organs
only by a narrow synaptic cleft. When an ac-
Signal transmission in the CNS
tion potential (see p. 350) reaches the presyn-
aptic membrane, voltage-gated Ca2+ channels
A. Structure of nerve cells
integrated into the membrane open and
Nerve cells (neurons) are easily excitable cells
trigger exocytosis of the neurotransmitter
that produce electrical signals and can react to
stored in the presynaptic cell (for details, see
such signals as well. Their structure is mark-
p. 228).
edly different from that of other types of cell.
Each neuron usually releases only one type
Numerous branching processes project from
of neurotransmitter. Neurons that release
their cell body (soma). Neurons are able to
dopamine are referred to as “dopaminergic,”
receive signals via dendrites and to pass
for example, while those that release acetyl-
them on via axons. The axons, which can be
choline are “cholinergic,” etc. The transmit-
up to 1 m long, are usually surrounded by
ters that are released diffuse through the
Schwann cells, which cover them with a
synaptic cleft and bind on the other side to
lipid-rich myelin sheath to improve their
receptors on the postsynaptic membrane.
electrical insulation.
These receptors are integral membrane pro-
The transfer of stimuli occurs at the
teins that have binding sites for neurotrans-
synapses, which link the individual neurons
mitters on their exterior (see p. 224).
to each other as well as linking neurons func-
The receptors for neurotransmitters are
tionally to muscle fibers. Neurotransmitters
divided into two large groups according to
(see p. 352) are stored in the axonal nerve
the effect produced by binding of the trans-
endings. These signaling substances are re-
mitter (for details, see p. 354).
leased in response to electrical signals in or-
Ionotropic receptors
(bottom left) are
der to excite neighboring neurons (or muscle
ligand-gated ion channels. When they open
cells). It is estimated that each neuron in the
as a result of the transmitter’s influence,
brain is in contact via synapses with approx-
ions flow in due to the membrane potential
imately 10 000 other neurons.
(see p. 126). If the inflowing ions are cations
There is a noticeably high proportion of
(Na+, K+, Ca2+), depolarization of the mem-
lipids in the composition of nerve cells, rep-
brane occurs and an action potential is trig-
resenting about 50% of their dry weight. In
gered on the surface of the postsynaptic cell.
particular, there is a very wide variety of
This is the way in which stimulatory trans-
phospholipids, glycolipids, and sphingolipids
mitters work (e. g., acetylcholine and gluta-
(see p. 216).
mate). By contrast, if anions flow in (mainly
Cl-), the result is hyperpolarization of the
postsynaptic membrane, which makes the
B. Neurotransmitters and neurohormones
production of a postsynaptic action potential
Neurosecretions are classed into two groups:
more dif cult. The action of inhibitory trans-
neurotransmitters are released into the syn-
mitters such as glycine and GABA is based on
aptic cleft in order to influence neighboring
this effect.
cells (C). They have a short range and a short
A completely different type of effect is ob-
lifespan. By contrast, neurohormones are re-
served in metabotropic receptors (bottom
leased into the blood, allowing them to cover
right). After binding of the transmitter, these
larger distances. However, the distinction be-
interact on the inside of the postsynaptic
tween the two groups is a fluid one; some
membrane with G proteins (see p. 384), which
neurotransmitters simultaneously function
in turn activate or inhibit the synthesis of
as neurohormones.
second messengers. Finally, second messen-
gers activate or inhibit protein kinases, which
phosphorylate cellular proteins and thereby
C. Synaptic signal transmission
alter the behavior of the postsynaptic cells
All chemical synapses function according to a
(signal transduction; see p. 386).
similar principle. In the area of the synapse,
the surface of the signaling cell (presynaptic
membrane) is separated from the surface of
the receiving cell
(postsynaptic membrane)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Brain and sensory organs
349
A. Structure of nerve cells
Dendrite
Axon
terminals
Cell body
(Soma)
Nucleus
Synapses
Axon
Myelin sheath
Schwann cell
Node of Ranvier
A. Neurotransmitters and neurohormones
Neuronal axon
Synaptic
Neurotransmitter
Neurohormone
cleft
Receptor
Postsynaptic cell
Erythrocyte
Blood capillary
C. Synaptic signal transmission
1
Action potential
Opens voltage-
gated calcium
channels
Presynaptic
Voltage-gated
membrane
Ca2 channel
Ca2
Ca2
Synaptic vesicle
with transmitter
2
Action
2
Ca2
Postsynaptic
potential
membrane
3
3
4
Enzyme
Na
Second
6
messenger
Ionotropic
Metabotropic
De-
receptor
receptor
Pre-
SM
polarization
cursor
5
Hyper-
Cellular
polarization
Cl
response
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
350
Tissues and organs
uli (or more rarely electrical stimuli). Binding
Resting potential and action
of a neurotransmitter to an ionotropic recep-
potential
tor results in a brief local increase in the
membrane potential from -60 mV to about
A. Resting potential
+30 mV. Although the membrane potential
A characteristic property of living cells is the
quickly returns to the initial value within a
uneven distribution of positively and nega-
few milliseconds (ms) at its site of origin, the
tively charged ions on the inside and outside
depolarization is propagated because neigh-
of the plasma membrane. This gives rise to a
boring membrane areas are activated during
membrane potential (see p. 126)—i. e., there is
this time period.
electrical voltage between the two sides of
[1] The process starts with the opening of
the membrane, which can only balance out
voltage-gated Na+ channels (see p. 222). Due
when ion channels allow the unevenly distrib-
to their high equilibrium potential (see A), Na+
uted ions to move.
ions flow into the cell and reverse the local
At rest, the membrane potential in most
membrane potential (depolarization).
cells is -60 to -90 mV. It mainly arises from
[2] The Na+ channels immediately close
the activity of Na+/K+ transporting ATPase
again, so that the inflow of positive charges
(“Na+/K+ ATPase”), which occurs on practically
is only very brief.
all animal cells. Using up ATP, this P-type
[3] Due to the increase in the membrane
enzyme (see p. 220) “pumps” three Na+ ions
potential, voltage-dependent K+ channels
out of the cell in exchange for two K+ ions.
open and K+ ions flow out. In addition, Na+/
Some of the K+ ions, following the concentra-
K+ ATPase (see A) pumps the Na+ ions that
tion gradient, leave the cell again through
have entered back out again. This leads to
potassium channels. As the protein anions
repolarization of the membrane.
that predominate inside the cell cannot fol-
[4] The two processes briefly lead to the
low them, and inflow of Cl- ions from the
charge even falling below the resting poten-
outside is not possible, the result is an excess
tial (hyperpolarization). The K+ channels also
of positive charges outside the cell, while
close after a few milliseconds. The nerve cell
anions predominate inside it.
is then ready for re-stimulation.
An equilibrium potential exists for each of
Generally, it is always only a very small
the ions involved. This is the value of the
part of the membrane that is depolarized dur-
membrane potential at which there is no net
ing an action potential. The process can there-
inflow or outflow of the ions concerned. For
fore be repeated again after a short refractory
K+ ions, the resting potential lies in the range
period, when the nerve cell is stimulated
of the membrane potential, while for Na+ ions
again. Conduction of the action potential on
it is much higher at +70 mV. At the first op-
the surface of the nerve cell is based on the
portunity, Na+ ions will therefore spontane-
fact that the local increase in the membrane
ously flow into the cell. The occurrence of
potential causes neighboring voltage-gated
action potentials is based on this (see B).
ion channels to open, so that the membrane
Nerve cell membranes contain ion chan-
stimulation spreads over the whole cell in the
nels for Na+, K+, Cl-, and Ca2+. These channels
form of a depolarization wave.
are usually closed and open only briefly to let
ions pass through. They can be divided into
channels that are regulated by membrane po-
tentials (“voltage-gated”—e. g., fast Na+ chan-
nels; see p. 222) and those regulated by
ligands (“ligand-gated”—e. g., nicotinic acetyl-
choline receptors; see p. 222).
B. Action potential
Action potentials are special signals that are
used to transmit information in the nervous
system. They are triggered by chemical stim-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Brain and sensory organs
351
A. Resting potential
Extracellular space
Na
Cl
K
Organic
anions
-60 mV
1
A
A
Sodium
P P P
P
P
Potassium
Chloride
channel
P
channel
channel
Cytoplasm
Na
Cl
K
Organic
Usually
Often
1
Na /K ATPase
anions
closed
open
3.6.1.37
B. Action potential
(mV)
Action potential
+40
2
+20
2
Depolarization
0
Na
-20
1
3
-40
Resting potential
1
3
K
-60
4
-80
Hyperpolarization
4
1
2
3
1
2
3
Time (ms)
Time (ms)
Na
Na
K
K
*
1
1
Na
Na
1
2
K
K
Na
Na
K
K
*
1
1
*Charge reversal
Na
at membrane
Na
3
4
K
K
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
352
Tissues and organs
This provides better protection against break-
Neurotransmitters
down by peptidases.
Neurotransmitters in the strict sense are sub-
Endorphins, dynorphins, and enkephalins
stances that are produced by neurons, stored
are a particularly interesting group of neuro-
in the synapses, and released into the synap-
peptides. They act as “endogenous opiates” by
tic cleft in response to a stimulus. At the post-
producing analgetic, sedative, and euphoriant
synaptic membrane, they bind to special re-
effects in extreme situations. Drugs such as
ceptors and affect their activity.
morphine and heroin activate the receptors
for these peptides (see p. 354).
Purine derivatives with neurotransmitter
A. Important neurotransmitters
function are all derived from adenine-con-
Neurotransmitters can be classified into sev-
taining nucleotides or nucleosides. ATP is re-
eral groups according to their chemical struc-
leased along with acetylcholine and other
ture. The table lists the most important rep-
transmitters, and among other functions it
resentatives of this family, which has more
regulates the emission of transmitters from
than 100 members.
its synapse of origin. The stimulatory effect
Acetylcholine, the acetic acid ester of the
of caffeine is mainly based on the fact that it
cationic alcohol choline (see p. 50) acts at
binds to adenosine receptors.
neuromuscular junctions, where it triggers
muscle contraction (see p. 334), and in certain
B. Biosynthesis of catecholamines
parts of the brain and in the autonomous
nervous system.
The catecholamines are biogenic amines that
Several proteinogenic amino acids
(see
have a catechol group. Their biosynthesis in
p. 60) have neurotransmitter effects. A partic-
the adrenal cortex and CNS starts from tyro-
ularly important one is glutamate, which acts
sine.
as a stimulatory transmitter in the CNS. More
[1] Hydroxylation of the aromatic ring ini-
than half of the synapses in the brain are
tially produces dopa (3,4-dihydroxyphenyl-
glutaminergic. The metabolism of glutamate
alanine). This reaction uses the unusual
and that of the amine GABA synthesized from
coenzyme tetrahydrobiopterin
(THB). Dopa
it (see below) are discussed in more detail on
(cf. p. 6) is also used in the treatment of Par-
p. 356. Glycine is an inhibitory neurotransmit-
kinson’s disease.
ter with effects in the spinal cord and in parts
[2] Decarboxylation of dopa yields dopa-
of the brain.
mine, an important transmitter in the CNS. In
Biogenic amines arise from amino acids by
dopaminergic neurons, catecholamine syn-
decarboxylation (see p. 62). This group in-
thesis stops at this point.
cludes
4-aminobutyrate
(γ-aminobutyric
[3] The adrenal gland and adrenergic neu-
acid, GABA), which is formed from glutamate
rons continue the synthesis by hydroxylating
and is the most important inhibitory trans-
dopamine into norepinephrine (noradrena-
mitter in the CNS. The catecholamines norepi-
line). Ascorbic acid
(vitamin C; see p. 368)
nephrine and epinephrine (see B), serotonin,
acts as a hydrogen-transferring coenzyme
which is derived from tryptophan, and hista-
here.
mine also belong to the biogenic amine group.
[4] Finally, N-methylation of norepineph-
All of them additionally act as hormones or
rine yields epinephrine
(adrenaline). The
mediators (see p. 380).
coenzyme for this reaction is S-adenosylme-
Peptides make up the largest group among
thionine (SAM; see p. 110).
the neurosecretions. Many peptide hormo-
nes—e. g., thyroliberin (TRH) and angiotensin
The physiological effects of the catechol-
II—simultaneously act as transmitters. Most
amines are mediated by a large number of
neuropeptides are small (3-15 AA). At their
different receptors that are of particular inter-
N-terminus, many of them have a glutamate
est in pharmacology. Norepinephrine acts in
residue that has been cyclized to form
the autonomic nervous system and certain
pyroglutamate (5-oxoproline, <G), while the
areas of the brain. Epinephrine is also used
C-terminus is often an acid amide (-NH2).
as a transmitter by some neurons.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Brain and sensory organs
353
A. Important neurotransmitters
O
CH3
Acetyl-
H3C C O
CH2
CH2
N CH3
choline
CH3
COO
Amino
Glutamate
H
N
C H
COO
Glycine
3
acids:
Dopa
(CH
)
H3N
CH2
2 2
COO
Glutamate
Glycine
H3N
CH2
Biogenic
γ-Aminobutyrate (GABA)
H3N
CH2
H3N
CH2
amines:
Dopa
CH2
CH2
CH2
Dopamine
Norepinephrine
HO
CH2
N
Epinephrine
Serotonine
COO
N
Histamine
N
H
H
GABA
Serotonin
Histamine
Peptides:
β-Endorphin
YGGFMTSEKSQTPLVTLFKNAITKNAYKKGE
NH
Met- and Leu-enkephalin
YGGFM und YGGFL
N
Thyroliberin (TRH)
<GHP-NH2
Gonadoliberin (GnRH)
<GHWSYGLRPG-NH
O CH2
2
Substance P
RPKPQQFFGLM
H
N
C
C
N
Somatostatin
AGCKNFFWKTFTSC
N
H
C
Angiotensin II
DRVYIHPF
O
C
H
Cholecystokinin (CCK-4)
WMDF-NH2
C
O
and many others
NH
2
Pyroglutamate (<G)
Thyroliberin
O
Purine
ATP
A
A
A
A
ADP
P
P P
derivatives:
P P
P
AMP
Adenosine
B. Biosynthesis of the catecholamines
1
3
Tyrosine 3-monooxygenase [Fe2 ,THB] 1.14.16.2
Dopamine β-monooxygenase[Cu] 1.14.17.1
Aromatic-L-amino-acid decarboxylase
2
4
Phenylethanolamine N-methyltransferase 2.1.1.28
(Dopa decarboxylase) [PLP] 4.1.1.28
Dehydro-
S-Adenosyl-
S-Adenosyl-
ascorbate
methionine homocysteine
N
A
N
A
P
P
COO
COO
CO2
CH3
Ascorbate
H
N
CH
THB
H3N CH
H3N
CH2
H3N
CH2
H2N
CH2
3
CH2
CH2
CH2
HO
CH
HO
CH
1
2
3
4
O2
H2O
O2
H2O
OH
OH
OH
OH
OH
OH
OH
OH
OH
Tyrosine
Dopa
Dopamine
Norepinephrine
Epinephrine
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
354
Tissues and organs
The nicotinic ACh receptor responds to the
Receptors for neurotransmitters
alkaloid nicotine contained in tobacco (many
Like all signaling substances, neurotrans-
of the physiological effects of nicotine are
mitters (see p. 352) act via receptor proteins.
based on this). The nicotinic receptor is iono-
The receptors for neurotransmitters are inte-
tropic. Its properties are discussed in greater
grated into the membrane of the postsynaptic
detail on p. 222.
cell, where they trigger ion inflow or signal
The muscarinic ACh receptors (of which
transduction processes (see p. 348).
there are at least five subtypes) are metabo-
tropic. Their name is derived from the
alkaloid muscarine, which is found in the fly
A. Receptors for neurotransmitters
agaric mushroom (Amanita muscaria), for ex-
A considerable number of receptors for neu-
ample. Like ACh, muscarine is bound at the
rotransmitters are already known and new
receptor, but in contrast to ACh (see C), it is
ones are continuing to be discovered. The
not broken down and therefore causes per-
table only lists the most important examples.
manent stimulation of muscle.
They are classified into two large groups ac-
The muscarinic ACh receptors influence the
cording to their mode of action.
cAMP level in the postsynaptic cells (M1, M3
Ionotropic receptors are ligand-gated ion
and M5 increase it, while subtypes M2 and M4
channels (left half of the table). The receptors
reduce it).
for stimulatory transmitters (indicated in the
table by a ⊕) mediate the inflow of cations
C. Metabolism of acetylcholine
(mainly Na+). When these open after binding
of the transmitter, local depolarization of the
Acetylcholine is synthesized from acetyl-CoA
postsynaptic membrane occurs. By contrast,
and choline in the cytoplasm of the presynap-
inhibitory neurotransmitters (GABA and gly-
tic axon [1] and is stored in synaptic vesicles,
cine) allow Cl- to flow in. This increases the
each of which contains around 1000-10 000
membrane’s negative resting potential and
ACh molecules. After it is released by exocy-
hinders the action of stimulatory transmitters
tosis (see p. 228), the transmitter travels by
(hyperpolarization,
-).
diffusion to the receptors on the postsynaptic
Metabotropic receptors (right half of the
membrane. Catalyzed by acetylcholinesterase,
table) are coupled to G proteins (see p. 386),
hydrolysis of ACh to acetate and choline im-
through which they influence the synthesis
mediately starts in the synaptic cleft [2], and
of second messengers. Receptors that work
within a few milliseconds, the ACh released
with type Gs proteins (see p. 386) increase
has been eliminated again. The cleavage
the cAMP level in the postsynaptic cell
products choline and acetate are taken up
([cAMP]
), while those that activate Gi pro-
again by the presynaptic neuron and reused
teins reduce it ([cAMP]
). Via type Gq pro-
for acetylcholine synthesis [3].
teins, other receptors increase the intracellu-
Substances that block the serine residue in
lar Ca2+ concentration ([Ca2+]
).
the active center of acetylcholinesterase
There are several receptor subtypes for
[2]—e. g., the neurotoxin E605
and other
most neurotransmitters. These are distin-
organophosphates—prevent ACh degradation
guished numerically (e. g., D1 to D5) or are
and thus cause prolonged stimulation of the
named after their agonists—i. e., after mole-
postsynaptic cell. This impairs nerve conduc-
cules experimentally found to activate the
tion and muscle contraction. Curare, a para-
receptor. For example, one specific subtype
lyzing arrow-poison used by South American
of glutamate receptors reacts to NMDA
Indians, competitively inhibits binding of ACh
(N-methyl-D-aspartate), while another sub-
to its receptor.
type reacts to the compound AMPA, etc.
B. Acetylcholine receptors
Acetylcholine (ACh) was the neurotransmitter
first discovered, at the beginning of the last
century. It binds to two types of receptor.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Brain and sensory organs
355
A. Receptors for neurotransmitters
Ionotropic
Metabotropic
Receptor
Transmitter
Ion(s)
Effect
Receptor
Transmitter
Effect
Acetyl-
Acetyl-
Acetylcholine
Acetylcholine
[
]
Ca2
choline
choline
Na
(muscarinic)
[cAMP]
(nicotinic)
M1, M3, M5, M2, M4
5HT3
Serotonin
5HT1
Serotonin
[
Ca2
]
Na
5HT2
„
[cAMP]
5HT4
„
[cAMP]
GABAa
GABA
CI
α1
Norepinephrine
[
Ca2
]
α2
„
[cAMP]
„
β1, β2, β3
[cAMP]
Glycine
Glycine
CI
[cAMP]
D1, D5
Dopamine
AMPA
Glutamate
Na K
D2, D3, D4
„
[cAMP]
NMDA
Glutamate
Na K Ca2
Kainate
Glutamate
Opioids
[cAMP]
Na K
δ,κ,µ
B. Acetylcholine receptors
Acetylcholine
Na
Effector enzyme
Receptor G protein
PL-C
α
β
Adenylate cyclase
γ
α
GTP
1. Nicotinic receptor
2. Muscarinic receptors
C. Metabolism of acetylcholine
S
A
Acetyl-
S
1
Acetylcholine
Choline acetyltransferase
A
1
A
2.3.1.6
P
Acetylcholinesterase
2
P P
3.1.1.7
Choline
Packing
Acetate-CoA ligase
3
3
6.2.1.1
A
S
P P P
A
Ca2
Acetate
Synaptic
Storage
vesicle
Re-uptake
Release
Presynaptic membrane
Choline
H2O
Acetate
2
Acetylcholine
Synaptic
cleft
Curare
Receptor
Neurotoxins
Postsynaptic membrane
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
356
Tissues and organs
B. Glutamate, glutamine, and GABA
Metabolism
The proteinogenic amino acid glutamate
The brain and other areas of the central ner-
(Glu) and the biogenic amine 4-aminobuty-
vous system (CNS) have high ATP require-
rate derived from it are among the most im-
ments. Although the brain only represents
portant neurotransmitters in the brain (see
about 2% of the body’s mass, it consumes
p. 352). They are both synthesized in the
around 20% of the metabolized oxygen and
brain itself. In addition to the neurons, which
ca. 60% of the glucose. The neurons’ high en-
use Glu or GABA as transmitters, neuroglia are
ergy requirements are mainly due to ATP-de-
also involved in the metabolism of these sub-
pendent ion pumps (particularly Na+/K+ AT-
stances.
Pase) and other active transport processes
Since glutamate and GABA as transmitters
that are needed for nerve conduction (see
must not appear in the extracellular space in
p. 350).
an unregulated way, the cells of the neuroglia
(center) supply “glutaminergic” and “GABAer-
A. Energy metabolism of the brain
gic” neurons with the precursor glutamine
(Gln), which they produce from glutamate
Glucose is normally the only metabolite from
with the help of glutamine synthetase [1].
which the brain is able to obtain adequate
GABA neurons (left) and glutamate neu-
amounts of ATP through aerobic glycolysis
rons
(right) initially hydrolyze glutamine
and subsequent terminal oxidation to CO2
with the help of glutaminase [1] to form glu-
and H2O. Lipids are unable to pass the blood-
tamate again. The glutamate neurons store
brain barrier, and amino acids are also only
this in vesicles and release it when stimu-
available in the brain in limited quantities
lated. The GABA neurons continue the degra-
(see B). As neurons only have minor glycogen
dation process by using glutamate decarbox-
reserves, they are dependent on a constant
ylase [3] to convert glutamate into the trans-
supply of glucose from the blood. A severe
mitter GABA.
drop in the blood glucose level—as can occur
Both types of neuron take up their trans-
after insulin overdosage in diabetics, for ex-
mitter again. Some of it also returns to the
ample—rapidly leads to a drop in the ATP level
neuroglia, where glutamate is amidated back
in the brain. This results in loss of conscious-
into glutamine.
ness and neurological deficits that can lead to
Glutamate can also be produced again from
death. Oxygen deficiency (hypoxia) also fint
GABA. The reaction sequence needed for this,
affects the brain. The effects of a brief period
known as the GABA shunt, is characteristic of
of hypoxia are still reversible, but as time
the CNS. A transaminase
[4] first converts
progresses irreversible damage increasingly
GABA and 2-oxoglutarate into glutamate and
occurs and finally complete loss of function
succinate semialdehyde
(-OOC-CH2-CH2-
(“brain death”).
CHO). In an NAD+-dependent reaction, the
During periods of starvation, the brain after
aldehyde is oxidized to succinic acid
[5],
a certain time acquires the ability to use ke-
from which 2-oxoglutarate can be regener-
tone bodies (see p. 312) in addition to glucose
ated again via tricarboxylic acid cycle reac-
to form ATP. In the first weeks of a starvation
tions.
period, there is a strong increase in the activ-
The function of glutamate as a stimulatory
ities of the enzymes required for this in the
transmitter in the brain is the cause of what is
brain. The degradation of ketone bodies in the
known as the “Chinese restaurant syndrome.”
CNS saves glucose and thereby reduces the
In sensitive individuals, the monosodium glu-
breakdown of muscle protein that maintains
tamate used as a flavor enhancer in Chinese
gluconeogenesis in the liver during starva-
cooking can raise the glutamate level in the
tion. After a few weeks, the extent of muscle
brain to such an extent that transient mild
breakdown therefore declines to one-third of
neurological disturbances can occur (dizzi-
the initial value.
ness, etc.).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Brain and sensory organs
357
A. Energy metabolism of the brain
Energy suppliers
Constantly
Small amount
Glycogen
during prolonged
required
starvation
Blood glucose
Glucose
Amino acids
Ketone bodies
A
O2
O2
P P
P
Glycolysis
3 Na
Tricarboxylic acid cycle
1
Respiratory chain
2K
A
P P P
CO2 + H2O
Na /K
-exchanging
1
ATPase 3.6.1.37
B. Glutamate, glutamine, and GABA
Glutamate
GABA
Glia cell
neuron
neuron
Glutamine
Glutamine
Glutamine
A
P P
P
H2O
H2O
2
A
1
NH
P P P
1
4
NH4
NH4
Glutamate
Glutamate
Glutamate
GABA
4
2-Oxoglutarate
3
Succinate
CO2
semialdehyde
Tricarboxylic
5
acid cycle
GABA
Exo-
N
A
cytosis
N
A
Succinic acid
GABA
GABA
Glutamate
Glutamate
1
Glutaminase 3.5.1.2
2
Glutamine synthetase 6.3.1.2
3
Glutamate decarboxylase 4.1.1.15
Glutamat-
GABA receptor
4
4-Aminobutyrate transaminase 2.6.1.19
Glutamate receptor
Rezeptor
Rezeptor
5
Succinic semialdehyde dehydrogenase
1.2.1.24
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
358
Tissues and organs
contain in such a way that colors can also be
Sight
perceived.
Two types of photoreceptor cell are found in
the human retina—rods and cones. Rods are
B. Signal cascade
sensitive to low levels of light, while the cones
are responsible for color vision at higher light
Dark (bottom left). Rod cells that are not
intensities.
exposed to light contain relatively high con-
Signaling substances and many proteins
centrations (70 µM) of the cyclic nucleotide
are involved in visual processes. Initially, a
cGMP
(3 ,5 -cycloGMP; cf. cAMP, p. 386),
light-induced cis-trans isomerization of the
which is synthesized by a guanylate cyclase
pigment retinal triggers a conformational
([2], see p. 388). The cGMP binds to an ion
change in the membrane protein rhodopsin.
channel in the rod membrane (bottom left)
Via the G protein transducin, which is associ-
and thus keeps it open. The inflow of cations
ated with rhodopsin, an enzyme is activated
(Na+, Ca2+) depolarizes the membrane and
that breaks down the second messenger
leads to release of the neurotransmitter glu-
cGMP. Finally, the cGMP deficiency leads to
tamate at the synapse (see p. 356).
hyperpolarization of the light-sensitive cell,
Light (bottom right). When the G protein
which is registered by subsequent neurons
transducin binds to light-activated rhodop-
as reduced neurotransmitter release.
sin* (see A, on the structure of the complex;
see p. 224), it leads to the GDP that is bound to
the transducin being exchanged for GTP. In
A. Photoreceptor
transducin* that has been activated in this
The cell illustrated opposite, a rod, has a
way, the GTP-containing α-subunit breaks
structure divided by membrane discs into
off from the rest of the molecule and in turn
which the 7-helix receptor rhodopsin is inte-
activates a membrane cGMP phosphodiester-
grated (see p. 224). In contrast to other recep-
ase [1]. This hydrolyzes cGMP to GMP and
tors in the 7-helix class (see p. 384), rhodop-
thus reduces the level of free cGMP within
sin is a light-sensitive chromoprotein. Its pro-
milliseconds. As a consequence, the cGMP
tein part, opsin, contains the aldehyde retinal
bound at the ion channel dissociates off and
(see p. 364)—an isoprenoid which is bound to
the channel closes. As cations are constantly
the ε-amino group of a lysine residue as an
being pumped out of the cell, the membrane
aldimine.
potential falls and hyperpolarization of the
The light absorption of rhodopsin is in the
cell occurs, which interrupts glutamate re-
visible range, with a maximum at about
lease.
500 nm. The absorption properties of the vis-
Regeneration. After exposure to light, several
ual pigment are thus optimally adjusted to
processes restore the initial conditions:
the spectral distribution of sunlight.
1. The α-subunit of transducin* inactivates
Absorption of a photon triggers isomeriza-
itself by GTP hydrolysis and thus termi-
tion from the 11-cis form of retinal to all-
nates the activation of cGMP esterase.
trans-retinal (top right). Within milliseconds,
2. The reduced Ca2+ concentration causes ac-
this photochemical process leads to an allos-
tivation of guanylate cyclase, which in-
teric conformational change in rhodopsin.
creases the cGMP level until the cation
The active conformation (rhodopsin*) binds
channels reopen.
and activates the G protein transducin. The
3. An isomerase [3] transfers all-trans -retinal
signal cascade (B) that now follows causes
to the 11-cis -form, in which it is available
the rod cells to release less neurotransmitter
for the next cycle. A dehydrogenase [4] can
(glutamate) at their synapses. The adjoining
also allow retinal to be supplied from vita-
bipolar neurons register this change and
min A (retinol).
transmit it to the brain as a signal for light.
There are several different rhodopsins in
the cones. All of them contain retinal mole-
cules as light-sensitive components, the ab-
sorption properties of which are modulated
by the different proportions of opsin they
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Brain and sensory organs
359
A. Photoreceptor
Light (h · ν)
CH3
H3C
CH3
11
Cytoplasm
Rhodopsin
12
Membrane
CH3
H3C
discs
HC
NH
11-cis-Retinal
Opsin
Interlamellar
space
The double bond
Cilium
between C-11 and C-
12 is isomerized from
Mitochon-
cis to trans by light
drion
B. Signal cascade
Nucleus
Light
CH3
CH3
H3C
CH3
H
Synapse
C
NH
Rhodopsin
Rhodopsin
*
CH3
Decreased release
all-trans-Retinal
of neurotransmitter
Rhodopsin*
11-cis-
Transducin
Retinal
GTP
Transducin
N
A
GDP
GDP
N
A
Opsin
3
GDP
all-trans
4
Retinal
Retinol
Transducin*
GTP
β
Vitamin A
γ
cGMP esterase 3.1.4.35
*activates
P
1
*
α
α
2
Guanylate cyclase 4.6.1.2
GTP
GDP
3
Retinal isomerase 5.2.1.3
4
Retinol dehydrogenase 1.1.1.105
P P
H2O
GTP
2
cGMP
1
GMP
Light
2
Ca
Na
Dark
Na
Ca2
ATP
ADP P
cGMP
Cytoplasm
Cation channel
Cation pump
Cation channel
Extracellular space
Na
Ca2
Na
Ca2
Closes due to
lack of cGMP
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
360
Nutrition
The minimum daily requirement of protein
Organic substances
is 37 g for men and 29 g for women, but the
A balanced human diet needs to contain a
recommended amounts are about twice these
large number of different components. These
values. Requirements in pregnant and breast-
include proteins, carbohydrates, fats, minerals
feeding women are even higher. Not only the
(including water), and vitamins. These sub-
quantity, but also the quality of protein is
stances can occur in widely varying amounts
important. Proteins that lack several essential
and proportions, depending on the type of
amino acids or only contain small quantities
diet. As several components of the diet are
of them are considered to be of low value, and
essential for life, they have to be regularly
larger quantities of them are therefore
ingested with food. Recommended daily min-
needed. For example, pulses only contain
imums for nutrients have been published by
small amounts of methionine, while wheat
the World Health Organization (WHO) and a
and corn proteins are poor in lysine. In con-
number of national expert committees.
trast to vegetable proteins, most animal pro-
teins are high-value (with exceptions such as
collagen and gelatin).
A. Energy requirement
Carbohydrates serve as a general and easily
The amount of energy required by a human is
available energy source. In the diet, they are
expressed in kJ d-1 (kilojoule per day). An
present as monosaccharides in honey and
older unit is the kilocalorie (kcal; 1 kcal =
fruit, or as disaccharides in milk and in all
4.187 kJ). The figures given are recommended
foods sweetened with sugar (sucrose). Meta-
values for adults with a normal body weight.
bolically usable polysaccharides are found in
However, actual requirements are based on
vegetable products (starch) and animal prod-
age, sex, body weight, and in particular on
ucts (glycogen). Carbohydrates represent a
physical activity. In those involved in compet-
substantial proportion of the body’s energy
itive sports, for example, requirements can
supply, but they are not essential.
increase from 12 000 to 17 000 kJ d-1.
Fats are primarily important energy suppli-
It is recommended that about half of the
ers in the diet. Per gram, they provide more
energy intake should be in the form of carbo-
than twice as much energy as proteins and
hydrates, a third at most in the form of fat, and
carbohydrates. Fats are essential as suppliers
the rest as protein. The fact that alcoholic bever-
of fat-soluble vitamins (see p. 364) and as
ages can make a major contribution to daily
sources of polyunsaturated fatty acids, which
energy intake is often overlooked. Ethanol has
are needed to biosynthesize eicosanoids (see
a caloric value of about 30 kJ g-1 (see p. 320).
pp. 48, 390).
Mineral substances and trace elements, a
very heterogeneous group of essential nu-
B. Nutrients
trients, are discussed in more detail on
Proteins provide the body with amino acids,
p. 362. They are usually divided into macro-
which are used for endogenous protein bio-
minerals and microminerals.
synthesis. Excess amino acids are broken
Vitamins are also indispensable compo-
down to provide energy (see p. 174). Most
nents of the diet. The animal body requires
amino acids are glucogenic—i. e., they can be
them in very small quantities in order to syn-
converted into glucose (see p. 180).
thesize coenzymes and signaling substances
Proteins are essential components of the
(see pp. 364-369).
diet, as they provide essential amino acids
that the human body is not capable of pro-
ducing on its own (see the table). Some amino
acids, including cysteine and histidine, are not
absolutely essential, but promote growth in
children. Some amino acids are able to sub-
stitute for each other in the diet. For example,
humans can form tyrosine, which is actually
essential, by hydroxylation from phenylala-
nine, and cysteine from methionine.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nutrients
361
A. Energy requirement
Daily
Recommended proportion of energy supply:
requirement
(average)
Fats 30%
Proteins
Carbohydrates
15 - 20%
50 - 55%
9 200 kJ
12 600 kJ
(2200 kcal)
(3
000 kcal)
B. Nutrients
Quantity
Energy content
Daily
General function
Essential
in body
kJ · g
-1
requirement (g)
in metabolism
constituents
(kg)
(kcal · g-1)
a
b c
10
17
37
55
92
Supplier of amino acids
Essential
(4.1)
amino acids::
29
45
75
Energy source
Val
(14)
Leu
(16)
Ile
(12)
Lys
(12)
Daily require-
Phe
(16)
Cys and
ment in mg per
Trp
( 3)
His stimu-
Met
(10)
kg body weight
late growth
Thr
( 8)
1
17
0
390
240-
General source
Non-essential
(4.1)
310
of energy (glucose)
nutritional
Energy reserve
constituent
(glycogen)
Roughage (cellulose)
Supporting substances
(bones, cartilage, mucus)
10-15
39
10
80
130
General energy source
Poly-
(9.3)
unsaturated
Most important energy
fatty acids:
reserve
Linoleic acid
Solvent for vitamins
Linolenic acid
Arachidonic acid
Supplier of
essential fatty acids
(together
10 g/day)
35-40
0
2 400
-
-
Solvent
Cellular building block
Dielectric
Reaction partner
Temperature regulator
Building blocks
Macrominerals
3
Electrolytes
Microminerals
0
Cofactors of enzymes
(trace elements)
Often precursors
Lipid-soluble vitamins
–
–
of coenzymes
Water-soluble
vitamins
a: Minimum daily requirement
b: Recommended daily intake
c: Actual daily intake in
c: industrialized nations
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
362
Nutrition
Ca2+, and phosphate (see p. 328), and storage
Minerals and trace elements
of Fe2+ and I-.
Resorption of the required mineral sub-
A. Minerals
stances from food usually depends on the
Water is the most important essential inor-
body’s requirements, and in several cases
ganic nutrient in the diet. In adults, the body
also on the composition of the diet. One ex-
has a daily requirement of 2-3 L of water,
ample of dietary influence is calcium (see
which is supplied from drinks, water con-
p. 342). Its resorption as Ca2+ is promoted by
tained in solid foods, and from the oxidation
lactate and citrate, but phosphate, oxalic acid,
water produced in the respiratory chain (see
and phytol inhibit calcium uptake from food
p. 140). The special role of water for living
due to complex formation and the production
processes is discussed in more detail else-
of insoluble salts.
where (see p. 26).
Mineral deficiencies are not uncommon
The elements essential for life can be div-
and can have quite a variety of causes—e. g.,
ided into macroelements (daily requirement
an unbalanced diet, resorption disturbances,
> 100 mg) and microelements (daily require-
and diseases. Calcium deficiency can lead to
ment < 100 mg). The macroelements include
rickets, osteoporosis, and other disturbances.
the electrolytes sodium (Na), potassium (K),
Chloride deficiency is observed as a result of
calcium (Ca), and magnesium (Mg), and the
severe Cl- losses due to vomiting. Due to the
nonmetals chlorine (Cl), phosphorus (P), sul-
low content of iodine in food in many regions
fur (S), and iodine (I).
of central Europe, iodine deficiency is wides-
The essential microelements are only re-
pread there and can lead to goiter. Magnesium
quired in trace amounts (see also p. 2). This
deficiency can be caused by digestive disor-
group includes iron (Fe), zinc (Zn), manganese
ders or an unbalanced diet—e. g., in alco-
(Mn), copper (Cu), cobalt (Co), chromium (Cr),
holism. Trace element deficiencies often re-
selenium (Se), and molybdenum (Mo). Fluo-
sult in a disturbed blood picture—i. e., forms of
rine (F) is not essential for life, but does pro-
anemia.
mote healthy bones and teeth. It is still a
The last column in the table lists some of
matter of controversy whether vanadium,
the functions of minerals. It should be noted
nickel, tin, boron, and silicon also belong to
that almost all of the macroelements in the
the essential trace elements.
body function either as nutrients or electro-
The second column in the table lists the
lytes. Iodine (as a result of its incorporation
average amounts of mineral substances in
into iodothyronines) and calcium act as sig-
the body of an adult weighing 65 kg. The daily
naling substances. Most trace elements are
requirements listed in the fourth column also
cofactors for proteins, especially for enzymes.
apply to an adult, and are average values.
Particularly important in quantitative terms
Children, pregnant and breast-feeding wo-
are the iron proteins hemoglobin, myoglobin,
men, and those who are ill generally have
and the cytochromes (see p. 286), as well as
higher mineral requirements relative to
more than 300 different zinc proteins.
body weight than men.
As the human body is able to store many
minerals, deviations from the daily ration are
balanced out over a given period of time.
Minerals stored in the body include water,
which is distributed throughout the whole
body; calcium, stored in the form of apatite
in the bones (see p. 340); iodine, stored as
thyroglobulin in the thyroid; and iron, stored
in the form of ferritin and hemosiderin in the
bone marrow, spleen, and liver (see p. 286).
The storage site for many trace elements is
the liver. In many cases, the metabolism of
minerals is regulated by hormones—for exam-
ple, the uptake and excretion of H2O, Na+,
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Nutrients
363
A. Minerals
Mineral
Content
Major source
Daily requirement
Functions/Occurrence
*
(g)
(g)
Water
35 000-
Drinks
1200
Solvent, cellular buil-
40 000
Water in solid
ding block, dielectric,
900
foods
coolant, medium for
From metabolism 300g
transport, reaction partner
Macroelements (daily requirement >100 mg)
Osmoregulation,
Na
100
Table salt
1.1-3.3
membrane potential,
mineral metabolism
Vegetables, fruit,
Membrane potential,
K
150
1.9-5.6
cereals
mineral metabolism
Bone formation,
1 300
Ca
Milk, milk products
0.8
blood clotting,
signal molecule
Bone formation,
Mg
20
Green vegetables
0.35
cofactor for enzymes
Cl
100
Table salt
1.7-5.1
Mineral metabolism
Bone formation, energy
Meat, milk,
P
650
0.8
metabolism, nucleic
cereals, vegetables
acid metabolism
S-containing
Lipid and carbohydrate
S
200
amino acids
0.2
metabolism,
(Cys and Met)
conjugate formation
Microelements (trace elements)
(mg
Meat, liver, eggs,
Hemoglobin, myoglobin,
Fe
4-5
vegetables, potatoes,
10
cytochromes,
cereals
Fe/S clusters
Meat, liver,
Zn
2-3
15
Zinc enzymes
cereals
Found in many
Mn
0.02
2-5
Enzymes
foodstuffs
Meat, vegetables,
Cu
0.1-0.2
2-3
Oxidases
fruit, fish
Co
<0.01
Meat
Traces
Vitamin B12
Cr
<0.01
0.05-0.2
Not clear
Cereals, nuts,
Mo
0.02
0.15-0.5
Redox enzymes
legumes
Se
Vegetables, meat
0.05-0.2
Selenium enzymes
Seafood,
I
0.03
iodized salt,
0.15
Thyroxin
drinking water
Requirement not known
Metals
Non-metals
Drinking water
F
0.0015-0.004
Bones, dental enamel
(fluoridated), tea, milk
* Content in the body of a 65 kg adult
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
364
Nutrition
the chromoprotein rhodopsin
(see p. 358).
Lipid-soluble vitamins
Retinoic acid, like the steroid hormones, in-
Vitamins are essential organic compounds
fluences the transcription of genes in the cell
that the animal organism is not capable of
nucleus. It acts as a differentiation factor in
forming itself, although it requires them in
growth and development processes. Vitamin
small amounts for metabolism. Most vitamins
A deficiency can result in night blindness, vis-
are precursors of coenzymes; in some cases,
ual impairment, and growth disturbances.
they are also precursors of hormones or act as
Vitamin D (calciol, cholecalciferol) is the
antioxidants. Vitamin requirements vary from
precursor of the hormone calcitriol (1α,25-di-
species to species and are influenced by age,
hydroxycholecalciferol; see p. 320). Together
sex, and physiological conditions such as
with two other hormones (parathyrin and
pregnancy, breast-feeding, physical exercise,
calcitonin), calcitriol regulates the calcium
and nutrition.
metabolism (see p. 342). Calciol can be syn-
thesized in the skin from 7-dehydrocholes-
terol, an endogenous steroid, by a photo-
A. Vitamin supply
chemical reaction. Vitamin D deficiencies
A healthy diet usually covers average daily
only occur when the skin receives insuf cient
vitamin requirements. By contrast, malnutri-
exposure to ultraviolet light and vitamin D is
tion, malnourishment (e. g., an unbalanced
lacking in the diet. Deficiency is observed in
diet in older people, malnourishment in alco-
the form of rickets in children and osteomala-
holics, ready meals), or resorption disturban-
cia in adults. In both cases, bone mineraliza-
ces lead to an inadequate supply of vitamins
tion is disturbed.
from which hypovitaminosis, or in extreme
Vitamin E (tocopherol) and related com-
cases avitaminosis, can result. Medical treat-
pounds only occur in plants
(e. g., wheat
ments that kill the intestinal flora—e. g., anti-
germ). They contain what is known as a chro-
biotics—can also lead to vitamin deficiencies
man ring. In the lipid phase, vitamin E is
(K, B12, H) due to the absence of bacterial
mainly located in biological membranes,
vitamin synthesis.
where as an antioxidant it protects unsatu-
Since only a few vitamins can be stored (A,
rated lipids against ROS (see p. 284) and other
D, E, B12), a lack of vitamins quickly leads to
radicals.
deficiency diseases. These often affect the
Vitamin K (phylloquinone) and similar sub-
skin, blood cells, and nervous system. The
stances with modified side chains are in-
causes of vitamin deficiencies can be treated
volved in carboxylating glutamate residues
by improving nutrition and by administering
of coagulation factors in the liver
(see
vitamins in tablet form. An overdose of vita-
p. 290). The form that acts as a cofactor for
mins only leads to hypervitaminoses, with
carboxylase is derived from the vitamin by
toxic symptoms, in the case of vitamins A
enzymatic reduction. Vitamin K antagonists
and D. Normally, excess vitamins are rapidly
(e. g., coumarin derivatives) inhibit this reduc-
excreted with the urine.
tion and consequently carboxylation as well.
This fact is used to inhibit blood coagulation
in prophylactic treatment against thrombosis.
B. Lipid-soluble vitamins
Vitamin K deficiency occurs only rarely, as the
Vitamins are classified as either lipid-soluble
vitamin is formed by bacteria of the intestinal
or water-soluble. The lipid-soluble vitamins
flora.
include vitamins A, D, E, and K, all of which
belong to the isoprenoids (see p. 52).
Vitamin A (retinol) is the parent substance
of the retinoids, which include retinal and
retinoic acid. The retinoids also can be synthe-
sized by cleavage from the provitamin β-car-
otene. Retinoids are found in meat-containing
diets, whereas β-carotene occurs in fruits and
vegetables (particularly carrots). Retinal is in-
volved in visual processes as the pigment of
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Vitamins
365
A. Vitamin supply
Healthy
nutrition
Vitamin require-
ment satisfied
Vitamin poisoning
Poor nutrition
Vitamin
Malnutrition
A and D
Antibiotics
Vitamin intake
Disturbed
Hypervitaminosis
resorption
Excess
vitamins
Hypo-
Diseases
vitaminosis
Urine
B. Lipid-soluble vitamins
* Adult daily
Provitamin
Functional form
Important for
* requirement
β-Carotene
Retinal
Sight
A
Vegetables
CH3
CH3
Visual pigments
H3C
CH3
Fruit
CH2OH
CH3
Retinol
Retinol
Sugar
transport
1
Coenzyme
Milk
Liver
Egg yolk
Development
Retinoic acid
D
Differentiation
H3C
CH
3
Signal molecule
H3C
CH3
Cholesterol
uv
Calcium
CH
2
Calciol
Calcitriol
metabolism
0.01
Hormone
HO
Cod liver oil
Milk
Egg yolk
E
CH
3
HO
CH3
Anti-
Tocopherols
Tocopherols
H
oxidant
H3C
O
10
Reducing agent
CH
3
3
CH
3
Cereals
Liver
Eggs
Seed oil
O
K
Blood clotting
3
CH
Phyllohydro-
Phylloquinones
(carboxylation
quinones
of plasma
H
0.08
proteins)
O
CH3
Intestinal
4
bacteria
Vegetables
Liver
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
366
Nutrition
synthesis and the amino acid metabolism are
Water-soluble vitamins I
affected.
The B group of vitamins covers water-soluble
In contrast to animals, microorganisms are
vitamins, all of which serve as precursors for
able to synthesize folate from their own com-
coenzymes. Their numbering sequence is not
ponents. The growth of microorganisms can
continuous, as many substances that were
therefore be inhibited by sulfonamides, which
originally regarded as vitamins were not later
competitively inhibit the incorporation of 4-
confirmed as having vitamin characteristics.
aminobenzoate into folate (see p. 254). Since
folate is not synthesized in the animal organ-
ism, sulfonamides have no effect on animal
A. Water-soluble vitamins I
metabolism.
Vitamin B1 (thiamine) contains two heterocy-
Nicotinate and nicotinamide, together re-
clic rings—a pyrimidine ring (a six-membered
ferred to as “niacin,” are required for biosyn-
aromatic ring with two Ns) and a thiazole ring
thesis of the coenzymes nicotinamide ad-
(a five-membered aromatic ring with N and
enine dinucleotide (NAD+) and nicotinamide
S), which are joined by a methylene group.
adenine dinucleotide phosphate
(NADP+).
The active form of vitamin B1 is thiamine
These both serve in energy and nutrient me-
diphosphate (TPP), which contributes as a
tabolism as carriers of hydride ions
(see
coenzyme to the transfer of hydroxyalkyl res-
pp. 32, 104). The animal organism is able to
idues (active aldehyde groups). The most im-
convert tryptophan into nicotinate, but only
portant reactions of this type are oxidative
with a poor yield. Vitamin deficiency there-
decarboxylation of
2-oxoacids
(see p. 134)
fore only occurs when nicotinate, nicotin-
and the transketolase reaction in the pentose
amide, and tryptophan are all simultaneously
phosphate pathway (see p. 152). Thiamine
are lacking in the diet. It manifests in the form
was the first vitamin to be discovered, around
of skin damage (pellagra), digestive distur-
100 years ago. Vitamin B1 deficiency leads to
bances, and depression.
beriberi, a disease with symptoms that in-
Pantothenic acid is an acid amide consist-
clude neurological disturbances, cardiac in-
ing of β-alanine and 2,4-dihydroxy-3,3 -di-
suf ciency, and muscular atrophy.
methylbutyrate (pantoic acid). It is a precur-
Vitamin B2 is a complex of several vita-
sor of coenzyme A, which is required for acti-
mins: riboflavin, folate, nicotinate, and pan-
vation of acyl residues in the lipid metabolism
tothenic acid.
(see pp. 12, 106). Acyl carrier protein (ACP; see
Riboflavin (from the Latin flavus, yellow)
p. 168) also contains pantothenic acid as part
serves in the metabolism as a component of
of its prosthetic group. Due to the widespread
the redox coenzymes flavin mononucleotide
availability of pantothenic acid in food (Greek
(FMN) and flavin adenine dinucleotide (FAD;
pantothen = “from everywhere”), deficiency
see p. 104). As prosthetic groups, FMN and FAD
diseases are rare.
are cofactors for various oxidoreductases (see
p. 32). No specific disease due to a deficiency
Further information
of this vitamin is known.
Folate, the anion of folic acid, is made up of
The requirement for vitamins in humans and
three different components—a pteridine
other animals is the result of mutations in the
derivative,
4-aminobenzoate, and one or
enzymes involved in biosynthetic coenzymes.
more glutamate residues. After reduction to
As intermediates of coenzyme biosynthesis
tetrahydrofolate
(THF), folate serves as a
are available in suf cient amounts in the
coenzyme in the C1 metabolism (see p. 418).
diet of heterotrophic animals
(see p. 112),
Folate deficiency is relatively common, and
the lack of endogenous synthesis did not
leads to disturbances in nucleotide biosynthe-
have unfavorable effects for them. Microor-
sis and thus cell proliferation. As the precur-
ganisms and plants whose nutrition is mainly
sors for blood cells divide particularly rapidly,
autotrophic have to produce all of these com-
disturbances of the blood picture can occur,
pounds themselves in order to survive.
with increased amounts of abnormal precur-
sors for megalocytes (megaloblastic anemia).
Later, general damage ensues as phospholipid
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Vitamins
367
A. Water-soluble vitamins I
* Adult daily
Vitamin
Active form:
Function in
*requirement
coenzyme
metabolism
B1
CH
3
Transfer of
N
N
Thiamine
TPP
hydroxy-
CH2
CH2OH
alkyl residues
S
Thiamine
H3C
N
NH2
1.5 mg *
diphosphate
Grain
Yeast products
Pork
O
B2
N
CH3
HN
F
O
N
N
CH3
FMN
Hydrogen
CH2
Riboflavin
F
A
transfer
H
C
OH
FAD
1.8 mg*
H
C
OH
Milk
H
C
OH
Eggs
CH2OH
4-Aminobenzoate residue
O
C1-
COO
OH
HN
C
Folate
THF
metabolism
N
CH2
NH
CH
N
Tetrahydro-
0.2 mg*
CH2
folate
H2N
N
N
CH2
Fresh green
vegetables
COO
Liver
Pteridine derivative
Glu
N
A
O
P
Nicotinate
Hydride
COO
Nicotinamide
NADP
transfer
NH
2
N
A
20 mg*
N
N
(or 1.2 g tryptophan)
NAD
Nicotinate
Nicotinamide
Meat, yeast products
Fruit and vegetables
H3C
CH3O
C
C
COO
S
Activation
Pantothenate
N
A
of carboxy-
H
lic acids
HO OH
7 mg*
CoA
β-Alanine
Pantoinate
Widely
distributed
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
368
Nutrition
Vitamin C is L-ascorbic acid (chemically:
Water-soluble vitamins II
2-oxogulonolactone). The two hydroxyl
groups have acidic properties. By releasing a
A. Water-soluble vitamins II
proton, ascorbic acid therefore turns into its
Vitamin B6 consists of three substituted pyr-
anion, ascorbate. Humans, apes, and guinea
idines—pyridoxal, pyridoxol, and pyrid-
pigs require vitamin C because they lack the
oxamine. The illustration shows the structure
enzyme L-gulonolactone oxidase
(1.1.3.8),
of pyridoxal, which carries an aldehyde group
which catalyzes the final step in the conver-
(-CHO) at C-4. Pyridoxol is the corresponding
sion of glucose into ascorbate.
alcohol
(-CH2OH), and pyridoxamine the
Vitamin C is particularly abundant in fresh
amine (-CH2NH2).
fruit and vegetables. Many soft drinks and
The active form of vitamin B6, pyridoxal
foodstuffs also have synthetic ascorbic acid
phosphate, is the most important coenzyme
added to them as an antioxidant and flavor
in the amino acid metabolism (see p. 106).
enhancer. Boiling slowly destroys vitamin C.
Almost all conversion reactions involving
In the body, ascorbic acid serves as a reducing
amino acids require pyridoxal phosphate, in-
agent in variations reactions (usually hydrox-
cluding transaminations, decarboxylations,
ylations). Among the processes involved are
dehydrogenations, etc. Glycogen phosphory-
collagen synthesis, tyrosine degradation, cate-
lase, the enzyme for glycogen degradation,
cholamine synthesis, and bile acid biosynthesis.
also contains pyridoxal phosphate as a cofac-
The daily requirement for ascorbic acid is
tor. Vitamin B6 deficiency is rare.
about 60 mg, a comparatively large amount
Vitamin B12 (cobalamine) is one of the most
for a vitamin. Even higher doses of the vita-
complex low-molecular-weight substances
min have a protective effect against infec-
occurring in nature. The core of the molecule
tions. However, the biochemical basis for
consists of a tetrapyrrol system (corrin), with
this effect has not yet been explained. Vitamin
cobalt as the central atom (see p. 108). The
C deficiency only occurs rarely nowadays; it
vitamin is exclusively synthesized by micro-
becomes evident after a few months in the
organisms. It is abundant in liver, meat, eggs,
form of scurvy, with connective-tissue dam-
and milk, but not in plant products. As the
age, bleeding, and tooth loss.
intestinal flora synthesize vitamin B12, strict
Vitamin H (biotin) is present in liver, egg
vegetarians usually also have an adequate
yolk, and other foods; it is also synthesized by
supply of the vitamin.
the intestinal flora. In the body, biotin is co-
Cobalamine can only be resorbed in the
valently attached via a lysine side chain to
small intestine when the gastric mucosa se-
enzymes that catalyze carboxylation reac-
cretes what is known as intrinsic factor—a
tions. Biotin-dependent carboxylases include
glycoprotein that binds cobalamine (the ex-
pyruvate carboxylase (see p. 154) and acetyl-
trinsic factor) and thereby protects it from
CoA carboxylase (see p. 162). CO2 binds, using
degradation. In the blood, the vitamin is
up ATP, to one of the two N atoms of biotin,
bound to a special protein known as trans-
from which it is transferred to the acceptor
cobalamin. The liver is able to store vitamin
(see p. 108).
B12 in amounts suf cient to last for several
Biotin
binds with high af nity
months. Vitamin B12
deficiency is usually
(Kd = 10-15 M) and specificity to avidin, a pro-
due to an absence of intrinsic factor and the
tein found in egg white. Since boiling dena-
resulting resorption disturbance. This leads to
tures avidin, biotin deficiency only occurs
a disturbance in blood formation known as
when egg whites are eaten raw.
pernicious anemia.
In animal metabolism, derivatives of cobal-
amine are mainly involved in rearrangement
reactions. For example, they act as coenzymes
in the conversion of methylmalonyl-CoA to
succinyl-CoA (see p. 166), and in the formation
of methionine from homocysteine (see p. 418).
In prokaryotes, cobalamine derivatives also
play a part in the reduction of ribonucleotides.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Vitamins
369
A. Water-soluble vitamins II
* Adult daily
Vitamin
Active form:
Function in
*requirement
coenzyme
metabolism
B
O
H
6
C
Pyridoxal
Activation
HO
CH2OH
Pyridoxol
PLP
of amino
Pyridoxamine
acids
Pyridoxal
phosphate
H3C
N
Pyridoxal
2 mg*
Meat
Vegetables
Grain products
B12
H2N
H2N
C
O
C
O
5-Deoxy-
Isomerization
X
Cobalamine
H3C
H3C
adenosyl
e.g.
H3C
O
cobalamine
H2N
N
C
C
NH2
0.002 mg*
N
Co2
N
CH3
O
NH2
H
H
O
C
HN
C
H3C
N
Meat
S
O
H C C C
H2C
O
CH3
CH
Liver
A
3
H3C CH
CH3
Milk
N
CH3
H COO
O
Eggs
O
P
O
N
H3C
C
Methylmalonyl CoA
O NH2
O
HO
O
HO
CH2
O
H H
S
C C
C
H
A
H
COO
Succinyl CoA
C
HO
O
Stabilization
of enzyme
Ascorbic acid
Ascorbate
systems,
O
HO
coenzyme,
60 mg*
antioxidant
HO
C
H
CH2OH
Fruit
Vegetables
O
H
Transfer
of
Biotin
B
HN NH
carboxyl
groups
0.1 mg*
Biotin
Yeast products
S
COO
Legumes
Nuts
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
370
Hormones
subject to hormonal regulation are the up-
Basics
take and degradation of storage substances
Hormones are chemical signaling substances.
(glycogen, fat), metabolic pathways for bio-
They are synthesized in specialized cells that
synthesis and degradation of central me-
are often associated to form endocrine glands.
tabolites
(glucose, fatty acids, etc.), and
Hormones are released into the blood and
the supply of metabolic energy.
transported with the blood to their effector
•
Digestive processes
organs. In the organs, the hormones carry
Digestive processes are usually regulated
out physiological and biochemical regulatory
by locally acting peptides (paracrine; see
functions. In contrast to endocrine hormones,
p. 372), but mediators, biogenic amines,
tissue hormones are only active in the imme-
and neuropeptides are also involved (see
diate vicinity of the cells that secrete them.
p. 270).
The distinctions between hormones and
•
Maintenance of ion concentrations
(ho-
other signaling substances (mediators, neuro-
meostasis)
transmitters, and growth factors) are fluid.
Concentrations of Na+, K+, and Cl- in body
Mediators is the term used for signaling sub-
fluids, and the physiological variables de-
stances that do not derive from special hor-
pendent on these (e. g. blood pressure), are
mone-forming cells, but are form by many cell
subject to strict regulation. The principal
types. They have hormone-like effects in their
site of action of the hormones involved is
immediate surroundings. Histamine
(see
the kidneys, where hormones increase or
p. 352) and prostaglandins (see p. 390) are
reduce the resorption of ions and recovery
important examples of these substances.
of water (see pp. 326-331). The concentra-
Neurohormones and neurotransmitters are
tions of Ca2+ and phosphate, which form
signaling substances that are produced and
the mineral substance of bone and teeth,
released by nerve cells (see p. 348). Growth
are also precisely regulated.
factors and cytokines mainly promote cell
Many hormones influence the above pro-
proliferation and cell differentiation
(see
cesses only indirectly by regulating the syn-
p. 392).
thesis and release of other hormones (hormo-
nal hierarchy; see p. 372).
A. Hormones: overview
B. Hormonal regulation system
The animal organism contains more than 100
hormones and hormone-like substances,
Each hormone is the center of a hormonal
which can be classified either according to
regulation system. Specialized glandular cells
their structure or according to their function.
synthesize the hormone from precursors,
In chemical terms, most hormones are amino
store it in many cases, and release it into the
acid derivatives, peptides or proteins, or ste-
bloodstream when needed (biosynthesis). For
roids. Hormones regulate the following pro-
transport, the poorly water-soluble lipophilic
cesses:
hormones are bound to plasma proteins
• Growth and differentiation of cells, tissues,
known as hormone carriers. To stop the ef-
and organs
fects of the hormone again, it is inactivated by
These processes include cell proliferation,
enzymatic reactions, most of which take place
embryonic development, and sexual dif-
in the liver (metabolism). Finally, the hor-
ferentiation—i. e., processes that require a
mone and its metabolites are expelled via
prolonged time period and involve proteins
the excretory system, usually in the kidney
de novo synthesis. For this reason, mainly
(excretion). All of these processes affect the
steroid hormones which function via tran-
concentration of the hormone and thus con-
scription regulation are active in this field
tribute to regulation of the hormonal signal.
(see p. 244).
In the effector organs, target cells receive the
• Metabolic pathways
hormone’s message. These cells have hormone
Metabolic regulation requires rapidly act-
receptors for the purpose, which bind the hor-
ing mechanisms. Many of the hormones
mone. Binding of a hormone passes informa-
involved therefore regulate interconversion
tion to the cell and triggers a response (effect).
of enzymes (see p. 120). The main processes
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Hormonal system
371
A. Hormones: overview
Peptide
Proliferation
Protein
Growth
Sexual
Cytokines
Steroid
hormones
Development
Growth factors
Amino
Retinoic
Somatotropin
acid
acid
derivative
Hormones
Metabolism
Digestion
Insulin
Gastrin
Glucagon
Secretin
Homeostasis
Cortisol
CCK
(H2O, electrolytes,
P
)
etc.
Thyroxin
Epinephrine
Parathyroid hormone
Calcitonin
ANF
Aldosterone
Calcitriol
A. Hormonal regulation system
Gland cell
Circulatory system
M
Target cell
H
H
P
H
H
H
Response
H
Biosynthesis
Transport
Effect
H
M
H
H
M
H
H
Hormone
H
M
H
M
HM
H
P
Hormone precursor
M
Hormone metabolite
Hormone carrier
H
M
Metabolism
HM
Excretion
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
372
Hormones
mone synthesis, release, and degradation al-
Plasma levels and hormone
lows the blood concentrations of hormones to
hierarchy
be precisely adjusted. This is based either on
simple feedback control or on hierarchically
A. Endocrine, paracrine, and autocrine hor-
structured regulatory systems.
mone effects
Hormones transfer signals by migrating from
C. Closed-loop feedback control
their site of synthesis to their site of action.
The biosynthesis and release of insulin by the
They are usually transported in the blood. In
pancreatic B cells (see p. 160) is stimulated by
this case, they are said to have an endocrine
high blood glucose levels (> 5 mM). The in-
effect (1; example: insulin). By contrast, tissue
sulin released then stimulates increased up-
hormones, the target cells for which are in the
take and utilization of glucose by the cells of
immediate vicinity of the glandular cells that
the muscle and adipose tissues. As a result,
produce them, are said to have a paracrine
the blood glucose level falls back to its normal
effect (2; example: gastrointestinal tract hor-
value, and further release of insulin stops.
mones). When signal substances also pass ef-
fects back to the cells that synthesize them,
they are said to have an autocrine effect (3;
D. Hormone hierarchy
example: prostaglandins). Autocrine effects
Hormone systems are often linked to each
are often found in tumor cells (see p. 400),
another, giving rise in some cases to a hier-
which stimulate their own proliferation in
archy of higher-order and lower-order hor-
this way.
mones. A particularly important example is
Insulin, which is formed in the B cells of the
the pituitary-hypothalamic axis, which is con-
pancreas, has both endocrine and paracrine
trolled by the central nervous system (CNS).
effects. As a hormone with endocrine effects,
Nerve cells in the hypothalamus react to
it regulates glucose and fat metabolism. Via a
stimulatory or inhibitory signals from the CNS
paracrine mechanism, it inhibits the synthesis
by releasing activating or inhibiting factors,
and release of glucagon from the neighboring
which are known as liberins (“releasing hor-
A cells.
mones”) and statins (“inhibiting hormones”).
These neurohormones reach the adenohy-
pophysis by short routes through the blood-
B. Dynamics of the plasma level
stream. In the adenohypophysis, they stimu-
Hormones circulate as signaling substances in
late
(liberins) or inhibit
(statins) the bio-
the blood at very low concentrations (10-12 to
synthesis and release of tropines. Tropines
between 10-7 mol L-1). These values change
(glandotropic hormones) in turn stimulate pe-
periodically in rhythms that depend on the
ripheral glands to synthesize glandular hor-
time of day, month, or year, or on physiolog-
mones. Finally, the glandular hormone acts on
ical cycles.
its target cells in the organism. In addition, it
The first example shows the circadian
passes effects back to the higher-order hormone
rhythm of the cortisol level. As an activator
systems. This (usually negative) feedback influ-
of gluconeogenesis (see p. 158), cortisol is
ences the concentrations of the higher-order hor-
mainly released in the early morning, when
mones, creating a feedback loop.
the liver’s glycogen stores are declining. Dur-
Many steroid hormones are regulated by
ing the day, the plasma cortisol level declines.
this type of axis—e. g., thyroxin, cortisol, es-
Many hormones are released into the blood
tradiol, progesterone, and testosterone. In the
in a spasmodic and irregular manner. In this
case of the glucocorticoids, the hypothalamus
case, their concentrations change in an
releases corticotropin-releasing hormone
episodic or pulsatile fashion. This applies, for
(CRH or corticoliberin, a peptide consisting
instance, to luteinizing hormone (LH, lutropin).
of 41 amino acids), which in turn releases
Concentrations of other hormones are
corticotropin (ACTH, 39 AAs) in the pituitary
event-regulated. For example, the body re-
gland. Corticotropin stimulates synthesis and
sponds to increased blood sugar levels after
release of the glandular steroid hormone cor-
meals by releasing insulin. Regulation of hor-
tisol in the adrenal cortex.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Hormonal system
373
A. Endocrine, paracrine and autocrine hormone effects
Gland cell
Target
Hormone receptor
cell
V
H
V
H
H
H
H
H
Gland and target cell
H
H
H
H
H
H
V
H
H
H
H
H
Blood
Hormone
H
stream
H
H
1. Endocrine
2. Paracrine
3. Autocrine
B. Plasma level dynamics
D. Hormone hierarchy
mg · l-1
CNS
Periodic
200
Precursors
100
Cortisol
R
12
18
24
6
12
18
24
Hypo-
Time of day
mU · ml-1
thalamus
Lutropin
Episodic, pulsatile
30
Liberin
Statin
20
(releasing
H
H
(inhibiting
hormone)
hormone)
10
12
15
18
21
24
3
6
9
Time of day
Precursors
mU · ml-1
R
Event-dependent
30
Pituitary
20
Insulin
10
Meals
Negative
12
15
18
21
24
3
6
9
Tropin
H
feed-
Time of day
back
C. Regulatory circuit
Precursors
Pancreatic B cell
Peripheral
V
gland
H
Glandular
hormone
Glucose
H Insulin
Glucose
Metabolite
Muscle cell, fat cell
Response
Target cell
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
374
Hormones
Testosterone is the most important of the
Lipophilic hormones
male sexual steroids (androgens). It is synthe-
Classifying hormones into hydrophilic and
sized in the Leydig intersitial cells of the
lipophilic molecules indicates the chemical
testes, and controls the development and
properties of the two groups of hormones
functioning of the male gonads. It also deter-
and also reflects differences in their mode of
mines secondary sexual characteristics in
action (see p. 120).
men (muscles, hair, etc.).
Cortisol, the most important glucocorticoid,
is synthesized by the adrenal cortex. It is in-
A. Lipophilic hormones
volved in regulating protein and carbohydrate
Lipophilic hormones, which include steroid
metabolism by promoting protein degrada-
hormones, iodothyronines, and retinoic acid,
tion and the conversion of amino acids into
are relatively small molecules (300-800 Da)
glucose. As a result, the blood glucose level
that are poorly soluble in aqueous media.
rises
(see p. 152). Synthetic glucocorticoids
With the exception of the iodothyronines,
(e. g., dexamethasone) are used in drugs due
they are not stored by hormone-forming cells,
to their anti-inflammatory and immunosup-
but are released immediately after being syn-
pressant effects.
thesized. During transport in the blood, they
Aldosterone, a mineralocorticoid, is also
are bound to specific carriers. Via intracellular
synthesized in the adrenal gland. In the kid-
receptors, they mainly act on transcription
neys, it promotes Na+ resorption by inducing
(see p. 358). Other effects of steroid hor-
Na+/K+ ATPase and Na+ channels (see p. 328).
mones—e. g., on the immune system—are
At the same time, it leads to increased K+
not based on transcriptional control. Their
excretion. In this way, aldosterone indirectly
details have not yet been explained.
increases blood pressure.
Calcitriol is a derivative of vitamin D (see
p. 364). On exposure to ultraviolet light, a
Steroid hormones
precursor of the hormone can also arise in
The most important steroid hormones in ver-
the skin. Calcitriol itself is synthesized in the
tebrates are listed on p. 57. Calcitriol (vitamin
kidneys (see p. 330). Calcitriol promotes the
D hormone) is also included in this group,
resorption of calcium in the intestine and in-
although it has a modified steroid structure.
creases the Ca2+ level in the blood.
The most important steroid hormone in in-
vertebrates is ecdysone.
Iodothyronines
Progesterone is a female sexual steroid be-
longing to the progestin (gestagen) family. It is
The thyroid hormone thyroxine (tetraiodo-
synthesized in the corpus luteum of the ova-
thyronine, T4) and its active form triiodo-
ries. The blood level of progesterone varies
thyronine (T3) are derived from the amino
with the menstrual cycle. The hormone pre-
acid tyrosine. The iodine atoms at positions 3
pares the uterus for a possible pregnancy.
and 5 of the two phenol rings are character-
Following fertilization, the placenta also starts
istic of them. Post-translational synthesis of
to synthesize progesterone in order to main-
thyroxine takes place in the thyroid gland
tain the pregnant state. The development of
from tyrosine residues of the protein thyro-
the mammary glands is also stimulated by
globulin, from which it is proteolytically
progesterone.
cleaved before being released. Iodothyronines
Estradiol is the most important of the
are the only organic molecules in the animal
estrogens. Like progesterone, it is synthesized
organism that contain iodine. They increase
by the ovaries and, during pregnancy, by the
the basal metabolic rate, partly by regulating
placenta as well. Estradiol controls the men-
mitochondrial ATP synthesis. In addition, they
strual cycle. It promotes proliferation of the
promote embryonic development.
uterine mucosa, and is also responsible for the
development of the female secondary sexual
characteristics (breast, fat distribution, etc.).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipophilic hormones
375
A. Lipophilic hormones
Hormone
Site of formation
Sites of action
Actions
Prepares uterus
Maintenance of
for pregnancy
pregnancy
Ovaries
Promotes implantation
Development of
of fertilized egg
mammary glands
Progesterone
Uterus
Menstrual cycle
Bone development
Stimulates proliferation
of endometrium
Development of
secondary female
Ovaries
sex characteristics
e.g., fat distribution,
Estradiol
Uterus and other organs
breasts, body hair
Causes:
Development of
Testes
secondary male
Sexual differentiation
sex characteristics
to male phenotype
e.g., skeleton,
Formation of ejaculate
muscles, body hair
Testosterone
Spermatogenesis
Protein synthesis
Proteolysis
Amino
Protein synthesis
Proteins
Glucose
acids
Gluconeogenese
Adrenal glands
Blut-Glucose
(cortex)
Activity of the immune
Cortisol
system
ATP
3Na
3Na
Na retention
Adrenal glands
K excretion
(cortex)
2K
2K
Aldosterone
Kidneys
Blood pressure
ADP+Pi
Bones
Ca2
- and phosphate
Kidneys
resorption
Ca2 metabolism
Ca2
Calcitriol
Gut
of bones
Thyroid gland
O2
H2O
Fetal development,
growth, and maturation
S
CO2
Basal metabolic rate
ADP+Pi
ATP, Heat
Heat generation
Thyroxine
Embryo
O2 consumption
Intermediary
metabolism
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
376
Hormones
thesis of the androgen testosterone from pro-
Metabolism of steroid hormones
gesterone, the side chain is completely re-
moved. Aromatization of the A ring, as men-
A. Biosynthesis of steroid hormones
tioned above, finally leads to estradiol.
All steroid hormones are synthesized from
On the way to calcitriol (vitamin D hor-
cholesterol. The gonane core of cholesterol
mone; see p. 342), another double bond in
consists of
19 carbon atoms in four rings
the B ring of cholesterol is first introduced.
(A-D). The D ring carries a side chain of eight
Under the influence of UV light on the skin,
C atoms (see p. 54).
the B ring is then photochemically cleaved,
The cholesterol required for biosynthesis of
and the secosteroid cholecalciferol arises (vi-
the steroid hormones is obtained from vari-
tamin D3; see p. 364). Two Cyt P450-depen-
ous sources. It is either taken up as a constit-
dent hydroxylations in the liver and kidneys
uent of LDL lipoproteins (see p. 278) into the
produce the active vitamin D hormone (see
hormone-synthesizing glandular cells, or syn-
p. 330).
thesized by glandular cells themselves from
acetyl-CoA (see p. 172). Excess cholesterol is
B. Inactivation of steroid hormones
stored in the form of fatty acid esters in lipid
droplets. Hydrolysis allows rapid mobilization
The steroid hormones are mainly inactivated
of the cholesterol from this reserve again.
in the liver, where they are either reduced or
Biosynthetic pathways. Only an overview
further hydroxylated and then conjugated
of the synthesis pathways that lead to the
with glucuronic acid or sulfate for excretion
individual hormones is shown here. Further
(see p. 316). The reduction reactions attack
details are given on p. 410.
oxo groups and the double bond in ring A. A
Among the reactions involved, hydroxyla-
combination of several inactivation reactions
tions (H) are particularly numerous. These are
gives rise to many different steroid metabo-
catalyzed by specific monooxygenases (“hy-
lites that have lost most of their hormonal
droxylases”) of the cytochrome P450 family
activity. Finally, they are excreted with the
(see p. 318). In addition, there are NADPH-
urine and also partly via the bile. Evidence of
dependent and NADP+-dependent hydrogena-
steroids and steroid metabolites in the urine
tions (Y) and dehydrogenations (D), as well as
is used to investigate the hormone metabo-
cleavage and isomerization reactions (S, I). The
lism.
estrogens have a special place among the ste-
roid hormones, as they are the only ones that
Further information
contain an aromatic A ring. When this is
formed, catalyzed by aromatase, the angular
Congenital defects in the biosynthesis of ste-
methyl group (C-19) is lost.
roid hormones can lead to severe develop-
Pregnenolone is an important intermedi-
mental disturbances. In the adrenogenital
ate in the biosynthesis of most steroid hor-
syndrome (AGS), which is relatively common,
mones. It is identical to cholesterol with the
there is usually a defect in 21-hydroxylase,
exception of a shortened and oxidized side
which is needed for synthesis of cortisol and
chain. Pregnenolone is produced in three
aldosterone from progesterone. Reduced syn-
steps by hydroxylation and cleavage in the
thesis of this hormone leads to increased for-
side chain. Subsequent dehydrogenation of
mation of testosterone, resulting in masculin-
the hydroxyl group at C-3 (b) and shifting of
ization of female fetuses. With early diagno-
the double bond from C-5 to C-4 results in the
sis, this condition can be avoided by providing
gestagen progesterone.
the mother with hormone treatment before
With the exception of calcitriol, all steroid
birth.
hormones are derived from progesterone. Hy-
droxylations of progesterone at C atoms 17,
21, and 11 lead to the glucocorticoid cortisol.
Hydroxylation at C-17 is omitted during syn-
thesis of the mineralocorticoid aldosterone.
Instead, the angular methyl group (C-18) is
oxidized to the aldehyde group. During syn-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipophilic hormones
377
A. Biosynthesis of steroid hormones
OH
H
DSHH
C
27
HO
CH2
HO
OH
C
D
H
Cholesterol
Calcitriol
A
B
D
H
H
HO
H/S
CH3
CH
OH
2
D
I
CO
CO
1
Cholesterol
HO
OH
HYDHH
HO
O
Pregnenolone
Cortisol
CH3
H
D
C21
Y
H
CO
D
CH3
CH2OH
H
CO
HHH
S
OHCCO
C
D
HO
HHHD
A
B
O
2
O
O
Progesterone
Progesterone
Aldosterone
H
S
Y
OH
C19
H:
Hydroxylation
A
O
H
D:
Dehydrogenation
Y
Progesterone
O
I:
Isomerization
H
C
D
A
Y:
Hydrogenation
A
B
OH
S:
Cleavage
O
Aromatization
A:
3
Androstenedione
C18
HO
Estradiol
B. Inactivation of steroid hormones
Oxidative cleavage
Conjugate
Oxidation
formation
OH
CH2OH
Conjugate
Oxidation
formation
Hydroxylation
CO
Reduction
HO
OH
Conjugate
formation
HO
O
Conjugate formation
Reduction
Cortisol
Estradiol
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
378
Hormones
“n” stands for any nucleotide). Each hormone
Mechanism of action
receptor only recognizes its “own” HRE and
therefore only influences the transcription of
A. Mechanism of action of lipophilic
genes containing that HRE. Recognition be-
hormones
tween the receptor and HRE is based on in-
Lipophilic signaling substances include the
teraction between the amino acid residues in
steroid hormones, calcitriol, the iodothy-
the DNA-binding domain (B) and the relevant
ronines (T3 and T4), and retinoic acid. These
bases in the HRE (emphasized in color in the
hormones mainly act in the nucleus of the
structure illustrated).
target cells, where they regulate gene tran-
As discussed on p.244, the hormone recep-
scription in collaboration with their receptors
tor does not interact directly with the RNA
and with the support of additional proteins
polymerase, but rather—along with other
(known as coactivators and mediators; see
transcription factors—with a coactivator/me-
p. 244). There are several effects of steroid
diator complex that processes all of the sig-
hormones that are not mediated by transcrip-
nals and passes them on to the polymerase. In
tion control. These alternative pathways for
this way, hormonal effects lead within a pe-
steroid effects have not yet been fully ex-
riod of minutes to hours to altered levels of
plained.
mRNAs for key proteins in cellular processes
In the blood, there are a number of trans-
(“cellular response”).
port proteins for lipophilic hormones (see
p. 276). Only the free hormone is able to pen-
B. Steroid receptors
etrate the membrane and enter the cell. The
hormone encounters its receptor in the nu-
The receptors for lipophilic signaling substan-
cleus (and sometimes also in the cytoplasm).
ces all belong to one protein superfamily. They
The receptors for lipophilic hormones are
are constructed in a modular fashion from
rare proteins. They occur in small numbers
domains with various lengths and functions.
(103-104
molecules per cell) and show
Starting from the N terminal, these are: the
marked specificity and high affinity for the
regulatory domain, the DNA-binding domain, a
hormone (Kd = 10-8-10-10 M). After binding
nuclear localization sequence (see p. 228), and
to the hormone, the steroid receptors are able
the hormone-binding domain (see p. 73D).
to bind as homodimers or heterodimers to
The homology among receptors is particu-
control elements in the promoters of specific
larly great in the area of the DNA-binding
genes, from where they can influence the
domain. The proteins have cysteine-rich se-
transcription of the affected genes—i. e., they
quences here that coordinatively bind zinc
act as transcription factors.
ions (A, Cys shown in yellow, Zn2+ in light
The illustration shows the particularly
blue). These centers, known as “zinc fingers”
well-investigated mechanism of action for
or “zinc clusters,” stabilize the domains and
cortisol, which is unusual to the extent that
support their dimerization, but do not take
the hormone-receptor complex already
part in DNA binding directly. As in other tran-
arises in the cytoplasm. The free receptor is
scription factors
(see p. 118),
“recognition
present in the cytoplasm as a monomer in
helices” are responsible for that.
complex with the chaperone hsp90
(see
In addition to the receptors mentioned in
p. 232). Binding of cortisol to the complex
A, the family of steroid receptors also includes
leads to an allosteric conformational change
the product of the oncogene erb-A
(see
in the receptor, which is then released from
p. 398), the receptor for the environmental
the hsp90 and becomes capable of DNA bind-
toxin dioxin, and other proteins for which a
ing as a result of dimerization.
distinct hormone ligand has not been identi-
In the nucleus, the hormone-receptor
fied (known as “orphan receptors”). Several
complex binds to nucleotide sequences
steroid receptors—e. g., the retinoic acid re-
known as hormone response elements
ceptor—form functional heterodimers with
(HREs). These are short palindromic DNA seg-
orphan receptors.
ments that usually promote transcription as
enhancer elements (see p. 244). The illustra-
tion shows the HRE for glucocorticoids (GRE;
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Lipophilic hormones
379
A. Mechanism of action of lipophilic hormones
Hormone
Target cell
Hormone
receptor
hsp
90
hsp
90
Heat-shock
protein
Nucleus
2x
Glucocorticoid receptor/DNA complex
DNA-binding
Hormone-receptor
domain (dimer)
bound to DNA
dimer
RNA
polymerase Gene
DNA
mRNA
Steroid
hormone
DNA
T3, T4
A
G
A
A
C
A
n
n
n
T
G
T
T
C
T
Calcitriol
Translation
Retinoic acid
T
C
T
T
G
T
n
n
n
A
C
A
A
G
A
Hormone response element (HRE)
Protein
Cell response
B. Receptors of lipophilic hormones
Variable length
Domains
Receptor
A/B
C
D
E
gene
Regulatory
DNA-
Nuclear-
Hormone-
domain
binding
targeting
binding
domain
sequence
domain
Binds ligand
Interaction with other
nuclear components
Domain E
Receptor protein
with a total of
~250 aa
Domain A/B
400 - 1000 aa
100 - 600 aa
Domain C
Binds to DNA
Domain D
~70 aa
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
380
Hormones
B. Examples of peptide hormones and
Hydrophilic hormones
proteohormones
The hydrophilic hormones are derived from
Numerically the largest group of signaling
amino acids, or are peptides and proteins
substances, these arise by protein biosynthe-
composed of amino acids. Hormones with
sis
(see p. 382). The smallest peptide hor-
endocrine effects are synthesized in glandular
mone, thyroliberin (362 Da), is a tripeptide.
cells and stored there in vesicles until they are
Proteohormones can reach masses of more
released. As they are easily soluble, they do
than 20 kDa—e. g., thyrotropin (28 kDa). Sim-
not need carrier proteins for transport in the
ilarities in the primary structures of many
blood. They bind on the plasma membrane of
peptide hormones and proteohormones
the target cells to receptors that pass the hor-
show that they are related to one another.
monal signal on
(signal transduction; see
They probably arose from common predeces-
p. 384). Several hormones in this group have
sors in the course of evolution.
paracrine effects—i. e., they only act in the
Thyroliberin
(thyrotropin-releasing hor-
immediate vicinity of their site of synthesis
mone, TRH) is one of the neurohormones of
(see p. 372).
the hypothalamus (see p. 330). It stimulates
pituitary gland cells to secrete thyrotropin
A. Signaling substances derived from amino
(TSH). TRH consists of three amino acids,
acids
which are modified in characteristic ways
(see p. 353).
Histamine, serotonin, melatonin, and the cat-
Thyrotropin
(thyroid-stimulating hor-
echolamines dopa, dopamine, norepineph-
mone, TSH) and the related hormones
rine, and epinephrine are known as “biogenic
lutropin
(luteinizing hormone, LH) and
amines.” They are produced from amino acids
follitropin
(follicle-stimulating hormone,
by decarboxylation and usually act not only as
FSH) originate in the adenohypophysis. They
hormones, but also as neurotransmitters.
are all dimeric glycoproteins with masses of
Histamine, an important mediator (local
around 28 kDa. Thyrotropin stimulates the
signaling substance) and neurotransmitter, is
synthesis and secretion of thyroxin by the
mainly stored in tissue mast cells and baso-
thyroid gland.
philic granulocytes in the blood. It is involved
Insulin (for the structure, see p. 70) is pro-
in inflammatory and allergic reactions. “His-
duced and released by the B cells of the pan-
tamine liberators” such as tissue hormones,
creas and is released when the glucose level
type E immunoglobulins (see p. 300), and
rises. Insulin reduces the blood sugar level by
drugs can release it. Histamine acts via vari-
promoting processes that consume glu-
ous types of receptor. Binding to H1 receptors
cose—e. g., glycolysis, glycogen synthesis,
promotes contraction of smooth muscle in the
and conversion of glucose into fatty acids. By
bronchia, and dilates the capillary vessels and
contrast, it inhibits gluconeogenesis and gly-
increases their permeability. Via H2 receptors,
cogen degradation. The transmission of the
histamine slows down the heart rate and pro-
insulin signal in the target cells is discussed
motes the formation of HCl in the gastric mu-
in greater detail on p. 388.
cosa. In the brain, histamine acts as a neuro-
Glucagon, a peptide of 29 amino acids, is a
transmitter.
product of the A cells of the pancreas. It is the
Epinephrine is a hormone synthesized in
antagonist of insulin and, like insulin, mainly
the adrenal glands from tyrosine
(see
influences carbohydrate and lipid metabo-
p. 352). Its release is subject to neuronal con-
lism. Its effects are each opposite to those of
trol. This “emergency hormone” mainly acts
insulin. Glucagon mainly acts via the second
on the blood vessels, heart, and metabolism. It
messenger cAMP (see p. 384).
constricts the blood vessels and thereby in-
creases blood pressure (via α1 and α2 recep-
tors); it increases cardiac function (via β2 re-
ceptors); it promotes the degradation of gly-
cogen into glucose in the liver and muscles
(via β2 receptors); and it dilates the bronchia
(also via β2 receptors).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Hydrophilic hormones
381
A. Signaling substances derived from amino acids
Hormone
Sites of formation
Sites of action
Actions
Width of bronchi
Histamine
H3N
CH2
stores
Capillaries:
CH2
width
Mast cell
Lungs
permeability
N
HCl
N
Gastric acid
H
secretion
Basophilic
by parietal cells
Histamine
granulocyte
Stomach
Cardiac output
CH3
H2N
CH2
Width of
Adrenal glands
Heart
blood vessels
HO CH
(medulla)
Blood pressure
Metabolism:
OH
Glycogenolysis
OH
Liver
Blood glucose
Adipose
Epinephrine
tissue
Muscle
Lipolysis
B. Examples of peptide hormones and proteohormones
Hypothalamus
Pituitary
Thyrotropin
secretion
Thyroliberin
(TRH)
Neurotransmitter
3 AA
action
Brain
TSH
Thyrotropin
(TSH)
Synthesis and
secretion of thyroxine
α chain 92 AA
Thyroid
β chain 112 AAAdeno-
hypophysis
gland
Thyroxine
Insulin
Glucose
Glucose
A chain 21 AA
uptake by cells
B chain 30 AA
Blood glucose
B cells
Glycogen
Proteins
Fats
Pancreas
Storage compounds:
formation
Glucose
Amino
Fatty
degradation
acids
acids
Glycogen
Fats
Glycogenolysis
Glucagon
Gluconeogenesis
29 AA
Glucose
Amino
Fatty
Blood glucose
A cells
acids
acids
Pancreas
Ketone body
Ketone
formation
bodies
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
382
Hormones
arises, which is modified at both ends (see
Metabolism of peptide hormones
p. 246). This mRNA codes for prepro-POMC—
Hydrophilic hormones and other water-solu-
i. e., a POMC protein that still has a signal
ble signaling substances have a variety of bio-
peptide for the ER at the N terminus (see
synthetic pathways. Amino acid derivatives
p. 230).
arise in special metabolic pathways
(see
[2] Prepro-POMC arises through translation
p. 352) or through post-translational modi-
in the rough endoplasmic reticulum (rER).
fication (see p. 374). Proteohormones, like all
The growing peptide chain is introduced
proteins, result from translation in the ribo-
into the ER with the help of a signal peptide.
some (see p. 250). Small peptide hormones
[3] Cleavage of the signal peptide and other
and neuropeptides, most of which only con-
modifications in the ER (formation of disulfide
sist of 3-30 amino acids, are released from
bonds, glycosylation, phosphorylation) give
precursor proteins by proteolytic degradation.
rise to the mature prohormone (“pro-POMC”).
[4] The neuropeptides and hormones men-
tioned are now formed by limited proteolysis
A. Biosynthesis
and stored in vesicles. Release from these
The illustration shows the synthesis and pro-
vesicles takes place by exocytosis when
cessing of the precursor protein proopiome-
needed.
lanocortin (POMC) as an example of the bio-
The biosynthesis of peptide hormones and
synthesis of small peptides with signaling
proteohormones, as well as their secretion, is
functions. POMC arises in cells of the adeno-
controlled by higher-order regulatory sys-
hypophysis, and after processing in the rER
tems (see p. 372). Calcium ions are among
and Golgi apparatus, it supplies the opiate-
the substances involved in this regulation as
like peptides met-enkephalin and E-endor-
second messengers; an increase in calcium
phin
(implying
“opio-”; see p. 352), three
ions stimulates synthesis and secretion.
melanocyte-stimulating hormones (α-, β- and
γ-MSH, implying “melano-”), and the glan-
B. Degradation and inactivation
dotropic hormone corticotropin (ACTH, im-
plying
“-cortin”). Additional products of
Degradation of peptide hormones often starts
POMC degradation include two lipotropins
in the blood plasma or on the vascular walls;
with catabolic effects in the adipose tissue
it is particularly intensive in the kidneys.
(β- and γ-LPH).
Several peptides that contain disulfide
Some of the peptides mentioned are over-
bonds (e. g., insulin) can be inactivated by
lapping in the POMC sequence. For example,
reductive cleavage of the disulfide bonds (1).
additional cleavage of ACTH gives rise to α-
Peptides and proteins are also cleaved by
MSH and corticotropin-like intermediary
peptidases, starting from one end of the pep-
peptide (CLIP). Proteolytic degradation of β-
tide by exopeptidases (2), or in the middle of it
LPH provides γ-LPH and β-endorphin. The
by proteinases (endopeptidases, 3). Proteoly-
latter can be further broken down to yield
sis gives rise to a variety of hormone frag-
met-enkephalin, while γ-LPH can still give
ments, several of which are still biologically
rise to β-MSH (not shown). Due to the numer-
active. Some peptide hormones and proteo-
ous derivative products with biological activ-
hormones are removed from the blood by
ity that it has, POMC is also known as a poly-
binding to their receptors with subsequent
protein. Which end product is formed and in
endocytosis of the hormone-receptor com-
what amounts depends on the activity of the
plex (4). They are then broken down in the
proteinases in the ER that catalyze the indi-
lysosomes. All of the degradation reactions
vidual cleavages.
lead to amino acids, which become available
The principles underlying protein synthe-
to the metabolism again.
sis and protein maturation (see pp. 230-233)
can be summed up once again using the ex-
ample of POMC:
[1] As a result of transcription of the POMC
gene and maturation of the hnRNA, a mature
mRNA consisting of some 1100 nucleotides
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Hydrophilic hormones
383
A. Biosynthesis
Chromosome
DNA
Human
chromosome
with
1
POMC gene
Transcription
87~3 900
151
~2 800
834 Base pairs
5`
3`
Splicing
Intron
Intron
Exon
TATA-
Box
~1100 Nucleotides
POMC-
mRNA
meGppp
AAAA---
mRNA
Kappe
Poly(A)
Translation
Translation
sequence
2
Translation
start
stop
γ-MSH
ACTH
β-LPH
Peptides
encoded
by
50-6
112-150
153-236
POMC gene
α-MSH CLIP
γ-LPH
β-Endorphin
Cleavage of
3
signal peptide
112-124
129-150
153-207
220-236
β-MSH
Met-Enke-
phalin
190-207
210-214
Signal peptide
for secretion
Prohormone
Pro-ACTH
P
Limited
proteolysis
ACTH
β-LPH
Protein
4
modification
Storage
P
Secretion
ACTH
Hormone
ACTH
(Cortico-
P
tropin)
β-En-
γ-MSH
α-MSH-CLIP
γ-LPH
dorphin
β-Endorphin
and other
peptides
P
B. Degradation and inactivation
Extracellular degradation:
Intracellular degradation:
1. Cleavage of
4. Binding to
1. disulfide bonds
4. membrane
1. by reductases
receptors
4. Endocytosis
4. Degradation
2. Degradation by
in lysosomes
2. exopeptidases
3. Degradation by
3. proteinases
Disulfide bond
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
384
Hormones
B. Signal transduction by G proteins
Mechanisms of action
G proteins transfer signals from 7-helix re-
The messages transmitted by hydrophilic sig-
ceptors to effector proteins (see above). G
naling substances (see p. 380) are sent to the
protein are heterotrimers consisting of three
interior of the cell by membrane receptors.
different types of subunit (α, β, and γ; see
These bind the hormone on the outside of
p. 224). The α-subunit can bind GDP or GTP
the cell and trigger a new second signal on
(hence the name “G protein”) and has GTPase
the inside by altering their conformation. In
activity. Receptor-coupled G proteins are re-
the interior of the cell, this secondary signal
lated to other GTP-binding proteins such as
influences the activity of enzymes or ion
Ras (see pp. 388, 398) and EF-Tu (see p. 252).
channels. Via further steps, switching of the
G proteins are divided into several types,
metabolism, changes in the cytoskeleton, and
depending on their effects. Stimulatory G pro-
activation or inhibition of transcription factors
teins (Gs) are widespread. They activate ad-
can occur (“signal transduction”) can occur.
enylate cyclases (see below) or influence ion
channels. Inhibitory G proteins (Gi) inhibit ad-
A. Mechanisms of action
enylate cyclase. G proteins in the Gq family
activate another effector enzyme—phospholi-
Receptors are classified into three different
pase c (see p. 386).
types according to their structure (see also
Binding of the signaling substance to a 7-
p. 224):
helix receptor alters the receptor conforma-
1. 1-Helix receptors (left) are proteins that
tion in such a way that the corresponding G
span the membrane with only one α-helix. On
protein can attach on the inside of the cell.
their inner (cytoplasmic) side, they have do-
This causes the α-subunit of the G protein to
mains with allosterically activatable enzyme
exchange bound GDP for GTP (1). The G pro-
activity. In most cases, these are tyrosine
tein then separates from the receptor and
kinases.
dissociates into an α-subunit and a βγ-unit.
Insulins (see p. 388), growth factors, and
Both of these components bind to other mem-
cytokines (see p. 392), for example, act via 1-
brane proteins and alter their activity; ion
helix receptors. Binding of the signaling sub-
channels are opened or closed, and enzymes
stance leads to activation of internal kinase
are activated or inactivated.
activity (in some cases, dimerization of the
In the case of the β2-catecholamine recep-
receptor is needed for this). The activated
tor (illustrated here), the α-subunit of the Gs
kinase phosphorylates itself using ATP (auto-
protein, by binding to adenylate cyclase, leads
phosphorylation), and also phosphorylates ty-
to the synthesis of the second messenger
rosine residues of other proteins (known as
cAMP. cAMP activates protein kinase A, which
receptor substrates). Adaptor proteins that
in turn activates or inhibits other proteins (2;
recognize the phosphotyrosine residues bind
see p. 120).
to the phosphorylated proteins (see pp. 388,
The βγ-unit of the G protein stimulates a
392). They pass the signal on to other protein
kinase (βARK, not shown), which phosphory-
kinases.
lates the receptor. This reduces its af nity for
2. Ion channels (center). These receptors
the hormone and leads to binding of the
contain ligand-gated ion channels. Binding of
blocking protein arrestin. The internal GTPase
the signaling substance opens the channels
activity of the α-subunit hydrolyzes the
for ions such as Na+, K+, Ca2+, and Cl-. This
bound GTP to GDP within a period of seconds
mechanism is mainly used by neurotrans-
to minutes, and thereby terminates the action
mitters such as acetylcholine (nicotinic recep-
of the G protein on the adenylate cyclase (3).
tor; see p. 224) and GABA (A receptor; see
p. 354).
3. 7-Helix receptors (serpentine receptors,
right) represent a large group of membrane pro-
teins that transfer the hormone or transmitter
signal, with the help of G proteins (see below), to
effector proteins that alter the concentrations
of ions and second messengers (see B).
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Hydrophilic hormones
385
A. Mechanisms of action
1. 1-Helix receptor
2. Ion channel
3. 7-Helix
Effector enzyme
receptor
Hydro-
Adenylate cyclase
philic
Pore for
Phospholipase C
signaling
ions
and A2
substance
Guanylate cyclase
Cytoplasm
G protein
7 Trans-
G protein
membrane
Tyrosine
increases
helices
forms
kinase
P
Ion concentration
Second messenger
Receptor
Ca2
cAMP
Ca2
substrate
Na
cGMP
NO
K
DAG
u.a.
ATP ADP
Cl
InsP3
Protein kinases (PKs)
Other
phosphorylate
Protein phosphatases
modify
enzymes
Enzymes
PK-A
and many
PK-G
others
PK-C
Transcription factors
modulate
Nucleus
Transcription
Metabolism
Cytoskeleton
B. Signal transduction by G proteins
Signaling
Signaling
Signaling
G protein
Active
Inactive
substance
substance
substance
(Gs)
α-unit
α-unit
1
1
1
P
GDP
ATP
GTP
GDP
PP
+
GDP GTP
β, γ-unit
cAMP
Arrestin
P
Activated 7-helix receptor
Second
messenger
1
Adenylate cyclase 4.6.1.1
1.
2.
3.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
386
Hormones
phosphate (InsP3), which is soluble, and diac-
Second messengers
ylglycerol (DAG). InsP3 migrates to the endo-
Second messengers are intracellular chemical
plasmic reticulum (ER), where it opens Ca2+
signals, the concentration of which is regu-
channels that allow Ca2+ to flow into the cy-
lated by hormones, neurotransmitters, and
toplasm (see C). By contrast, DAG, which is
other extracellular signals (see p. 384). They
lipophilic, remains in the membrane, where
arise from easily available substrates and only
it activates type C protein kinases, which
have a short half-life. The most important
phosphorylate proteins in the presence of
second messengers are cAMP, cGMP, Ca2+, in-
Ca2+ ions and thereby pass the signal on.
ositol triphosphate
(InsP3), diacylglycerol
(DAG), and nitrogen monoxide (NO).
C. Calcium ions
Calcium level. Ca2+ (see p. 342) is a signaling
A. Cyclic AMP
substance. The concentration of Ca2+ ions
Metabolism. The nucleotide cAMP (adenosine
in the cytoplasm is normally very low
3 ,5 -cyclic monophosphate) is synthesized
(10-100 nM), as it is kept down by ATP-
by membrane-bound adenylate cyclases [1]
driven Ca2+ pumps and Na+/Ca2+ exchangers.
on the inside of the plasma membrane. The
In addition, many proteins in the cytoplasm
adenylate cyclases are a family of enzymes
and organelles bind calcium and thus act as
that cyclize ATP to cAMP by cleaving diphos-
Ca2+ buffers.
phate (PPi). The degradation of cAMP to AMP
Specific signals (e. g., an action potential or
is catalyzed by phosphodiesterases [2], which
second messenger such as InsP3 or cAMP) can
are inhibited by methylxanthines such as caf-
trigger a sudden increase in the cytoplasmic
feine, for example. By contrast, insulin acti-
Ca2+ level to
500-1000 nM by opening
vates the esterase and thereby reduces the
Ca2+ channels in the plasma membrane or in
cAMP level (see p. 388).
the membranes of the endoplasmic or sarco-
Adenylate cyclase activity is regulated by G
plasmic reticulum. Ryanodine, a plant sub-
proteins (Gs and Gi), which in turn are con-
stance, acts in this way on a specific channel
trolled by extracellular signals via 7-helix re-
in the ER. In the cytoplasm, the Ca2+ level
ceptors (see p. 384). Ca2+-calmodulin (see be-
always only rises very briefly (Ca2+ “spikes”),
low) also activates specific adenylate cyclases.
as prolonged high concentrations in the cyto-
Action. cAMP is an allosteric effector of
plasm have cytotoxic effects.
protein kinase A (PK-A, [3]). In the inactive
Calcium effects. The biochemical effects of
state, PK-A is a heterotetramer (C2R2), the
Ca2+ in the cytoplasm are mediated by special
catalytic subunits of which (C) are blocked
Ca2+-binding proteins
(“calcium sensors”).
by regulatory units
(R; autoinhibition).
These include the annexins, calmodulin, and
When cAMP binds to the regulatory units,
troponin C in muscle (see p. 334). Calmodulin
the C units separate from the R units and
is a relatively small protein (17 kDa) that oc-
become enzymatically active. Active PK-A
curs in all animal cells. Binding of four Ca2+
phosphorylates serine and threonine residues
ions (light blue) converts it into a regulatory
of more than 100 different proteins, enzymes,
element. Via a dramatic conformational
and transcription factors. In addition to cAMP,
change (cf. 2a and 2b), Ca2+-calmodulin enters
cGMP also acts as a second messenger. It is
into interaction with other proteins and mod-
involved in sight (see p. 358) and in the signal
ulates their properties. Using this mechanism,
transduction of NO (see p. 388).
Ca2+ ions regulate the activity of enzymes, ion
pumps, and components of the cytoskeleton.
B. Inositol 1,4,5-trisphosphate and
diacylglycerol
Type Gq G proteins activate phospholipase C
[4]. This enzyme creates two second messen-
gers from the double-phosphorylated mem-
brane phospholipid phosphatidylinositol
bisphosphate (PInsP2), i. e., inositol 1,4,5-tris-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Hydrophilic hormones
387
A. Cyclic AMP
NH
2
7 Helix receptors
1
Adenylate cyclase 4.6.1.1
cAMP
C
N
C
N
2
Phosphodiesterase 3.1.4.17
HC
5'
C
CH
N
CH2
N
3
Protein kinase A 2.7.1.37
O
O
H H
O
G proteins
P
H
H
Caffeine
3'
G
s
Gi
O
O OH
Enzymes
ATP
1
cAMP
2
AMP
Transcription
ATP
factors
PPi
H2O
Ion channels
3
Protein kinase A
ADP
P
B. Inositol 1,4,5-trisphosphate and diacylglycerol
7 Helix
DAG (Diacylglycerol)
receptors
Phospholipid
Protein
kinase C
G protein (Gq)
Acyl residue 1
H2O
OH
OH
Acyl residue 2
4
H
O
P
H
H
Intracellular
P
Inositol
O
P
H
Ca2
P P
P
O
H
release
PlnsP2
H
OH
4
Phospholipase C 3.1.4.3
InsP3
C. Calcium ions
Ca
2
Na
Ca2
ATP
ATP
ER/SR
Ca2
Ca2
Ca2
10-100 nM
ADP
ADP
Pi
Pi
a
Ca2
InsP3
Ryanodine
Ca2
Calcium-
cAMP
500-1000 nM
binding
protein
Depolarization
b
Glutamate
Ca2
1. Calcium transport
ATP
2. Calmodulin
ca. 2 500 000 nM
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
388
Hormones
pids from the phosphatidylinositol group (see
Signal cascades
p. 50) at position
3. Protein kinase PDK-1
The signal transduction pathways that medi-
binds to these reaction products, becoming
ate the effects of the metabolic hormone in-
activated itself and in turn activating protein
sulin are of particular medical interest (see A).
kinase B (PK-B).
The mediator nitrogen monoxide (NO) is also
This has several effects. In a manner not yet
clinically important, as it regulates vascular
fully understood, PK-B leads to the fusion
caliber and thus the body’s perfusion with
with the plasma membrane of vesicles that
blood (see B).
contain the glucose transporter Glut-4. This
results in inclusion of Glut-4 in the membrane
and thus to increased glucose uptake into the
A. Insulin: signal transduction
muscles and adipose tissue (see p. 160). In
The diverse effects of insulin (see p. 160) are
addition, PK-B inhibits glycogen synthase kin-
mediated by protein kinases that mutually
ase 3 (GSK-3) by phosphorylation. As GSK-3 in
activate each other in the form of enzyme
turn inhibits glycogen synthase by phosphor-
cascades. At the end of this chain there are
ylation
(see p. 120), its inhibition by PK-B
kinases that influence gene transcription in
leads to increased glycogen synthesis. Protein
the nucleus by phosphorylating target pro-
phosphatase-1 (PP-1) converts glycogen syn-
teins, or promote the uptake of glucose and
thase into its active form by dephosphoryla-
its conversion into glycogen. The signal trans-
tion (see p. 120). PP-1 is also activated by
duction pathways involved have not yet been
insulin.
fully explained. They are presented here in a
simplified form.
B. Nitrogen monoxide (NO) as a mediator
The insulin receptor (top) is a dimer with
subunits that have activatable tyrosine kinase
Nitrogen monoxide (NO) is a short-lived rad-
domains in the interior of the cell (see p. 224).
ical that functions as a locally acting mediator
Binding of the hormone increases the tyrosine
(see p. 370).
kinase activity of the receptor, which then
In a complex reaction, NO arises from argi-
phosphorylates itself and other proteins
nine in the endothelial cells of the blood ves-
(receptor substrates) at various tyrosine re-
sels [1]. The trigger for this is Ca2+-calmodulin
sidues. Adaptor proteins, which conduct the
(see p. 386), which forms when there is an
signal further, bind to the phosphotyrosine
increase in the cytoplasmic Ca2+ level.
residues.
NO diffuses from the endothelium into the
The effects of insulin on transcription are
underlying vascular muscle cells, where it
shown on the left of the illustration. Adaptor
leads, as a result of activation of guanylate
proteins Grb-2 and SOS (“son of sevenless”)
cyclase [2], to the formation of the second
bind to the phosphorylated IRS (insulin-re-
messenger cGMP (see pp. 358, 384). Finally,
ceptor substrate) and activate the G protein
by activating a special protein kinase (PK-G),
Ras (named after its gene, the oncogene ras;
cGMP triggers relaxation of the smooth
see p. 398). Ras activates the protein kinase
muscle and thus dilation of the vessels. The
Raf (another oncogene product). Raf sets in
effects of atrionatriuretic peptide (ANP; see
motion a phosphorylation cascade that leads
p. 328) in reducing blood pressure are also
via the kinases MEK and ERK (also known as
mediated by cGMP-induced vasodilation. In
MAPK, “mitogen-activated protein kinase”) to
this case, cGMP is formed by the guanylate
the phosphorylation of transcription factors
cyclase activity of the ANP receptor.
in the nucleus.
Some of the effects of insulin on the carbo-
Further information
hydrate metabolism (right part of the illustra-
tion) are possible without protein synthesis.
The drug nitroglycerin
(glyceryl trinitrate),
In addition to Grb-2, another dimeric adaptor
which is used in the treatment of angina
protein can also bind to phosphorylated IRS.
pectoris, releases NO in the bloodstream and
This adaptor protein thereby acquires phos-
thereby leads to better perfusion of cardiac
phatidylinositol-3-kinase activity (PI3K) and,
muscle.
in the membrane, phosphorylates phospholi-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Hydrophilic hormones
389
A. Insulin: signal transduction
Insulin
Insulin
Phosphatidyl-
Glucose
receptor
α
inositols
α
Phosphatidyl-
inositol
3-phosphate
β
β
PI-3-
P
P
Kinase
SOS
P
P
Ras
Glut-4
Grb-2
IRS-1
PDK-1
?
Glucose
uptake
Raf
P
MEK
PK-B
(MAPKK)
Glut-4
PP-1
MAPK
P
(ERK)
GSK-3
Intracellular
Transcription
vesicle
factors
Glycogen synthase
Glycogen
(active)
synthesis
Activated
P
transcription
DNA
factors
Protein kinase
Inactive
Nucleus
Protein phosphatase
Active
Promoter
Transcription
B. Nitrogen monoxide (NO) as a mediator
Ca2
Physiological effects
PK-G
Signaling
substance
ER
5'
Guanine
ANP
CH
2O
G
InsP3
O
P
P P
H H
Arginine
O
GTP
P
H
H
3'
2 O2
O
O OH
cGMP
3
N
A
Ca2
P
3/2
Ca2
1
P
P
3/2
2
Calmodulin
N
A
P
2 H2O
Citrulline
NO·
NO·
G
P P P
GTP
Endothelial cell
Vascular muscle cell
1
NO synthase 1.14.13.39
2
Guanylate cyclase 4.6.1.2
3
ANF receptor 4.6.1.2
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
390
Hormones
The eicosanoids have a very wide range of
Eicosanoids
physiological effects. As they can stimulate or
The eicosanoids are a group of signaling sub-
inhibit smooth-muscle contraction, depend-
stances that arise from the C-20 fatty acid
ing on the substance concerned, they affect
arachidonic acid and therefore usually contain
blood pressure, respiration, and intestinal and
20 C atoms (Greek eicosa = 20). As mediators,
uterine activity, among other properties. In
they influence a large number of physiological
the stomach, prostaglandins inhibit HCl se-
processes (see below). Eicosanoid metabolism
cretion via Gi proteins (see p. 270). At the
is therefore an important drug target. As
same time, they promote mucus secretion,
short-lived substances, eicosanoids only act
which protects the gastric mucosa against
in the vicinity of their site of synthesis (para-
the acid. In addition, prostaglandins are in-
crine effect; see p. 372).
volved in bone metabolism and in the activity
of the sympathetic nervous system. In the
immune system, prostaglandins are impor-
A. Eicosanoids
tant in the inflammatory reaction. Among
Biosynthesis. Almost all of the body’s cells
other things, they attract leukocytes to the
form eicosanoids. Membrane phospholipids
site of infection. Eicosanoids are also deci-
that contain the polyunsaturated fatty acid
sively involved in the development of pain
arachidonic acid (20:4; see p. 48) provide the
and fever. The thromboxanes promote throm-
starting material.
bocyte aggregation and other processes in-
Initially, phospholipase A2 [1] releases the
volved in hemostasis (see p. 290).
arachidonate moiety from these phospholi-
Metabolism. Eicosanoids are inactivated
pids. The activity of phospholipase A2 is strictly
within a period of seconds to minutes. This
regulated. It is activated by hormones and
takes place by enzymatic reduction of double
other signals via G proteins. The arachidonate
bonds and dehydrogenation of hydroxyl
released is a signaling substance itself. How-
groups. As a result of this rapid degradation,
ever, its metabolites are even more important.
their range is very limited.
Two different pathways lead from arachi-
donate to prostaglandins, prostacyclins, and
Further information
thromboxanes, on the one hand, or leuko-
trienes on the other. The key enzyme for the
Acetylsalicylic acid and related non-steroidal
first pathway is prostaglandin synthase [2].
anti-inflammatory drugs (NSAIDs) selectively
Using up O2, it catalyzes in a two-step reac-
inhibit the cyclooxygenase activity of prosta-
tion the cyclization of arachidonate to prosta-
glandin synthase [2] and consequently the
glandin H2, the parent substance for the pros-
synthesis of most eicosanoids. This explains
taglandins, prostacyclins, and thromboxanes.
their analgesic, antipyretic, and antirheumatic
Acetylsalicylic acid (aspirin) irreversibly ace-
effects. Frequent side effects of NSAIDs also
tylates a serine residue near the active center
result from inhibition of eicosanoid synthesis.
of prostaglandin synthase, so that access for
For example, they impair hemostasis because
substrates is blocked (see below).
the synthesis of thromboxanes by thrombo-
As a result of the action of lipoxygenases [3],
cytes is inhibited. In the stomach, NSAIDs in-
hydroxyfatty acids and hydroperoxyfatty acids
crease HCl secretion and at the same time
are formed from arachidonate, from which
inhibit the formation of protective mucus.
elimination of water and various conversion
Long-term NSAID use can therefore damage
reactions give rise to the leukotrienes. The for-
the gastric mucosa.
mulae only show one representative from each
of the various groups of eicosanoids.
Effects. Eicosanoids act via membrane re-
ceptors in the immediate vicinity of their site
of synthesis, both on the synthesizing cell
itself (autocrine action) and on neighboring
cells (paracrine action). Many of their effects
are mediated by the second messengers cAMP
and cGMP.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Other signaling substances
391
A. Eicosanoids
Essential
Phospholipid
fatty acids:
Linoleic acid
Hormones and
Inclusion in
1
other signals
Linolenic acid
Lysophospholipid
Arachidonic acid
COO
Prostaglandin
Peroxidase
Arachidonate
CH3
synthase
center
Heme
2
Prosta-
3
Lipoxygenase
glandin
Cyclooxy-
Ser
synthase
genase
Acetylsalicylic
channel
acid
Hydroxy- and
Hydroperoxy fatty acids
Arachidonate
Prostaglandin H2
Leukotrienes
H
S Cys
Gly
O
COO
COO
CH3
OH
O
H
CH3
H OH
Prostaglandin H2
Leukotriene D4
Prostacyclins
Prostaglandins
Thromboxanes
COO
O
HO
HO
H
H
COO
COO
CH3
CH3
CH3
HO O
H
H
H
HO
H OH
HO
H OH
H OH
Prostacyclin I2
Prostaglandin F2α
Thromboxane B2
Effects: Stimulation of
Hormone-controlled lipases
Contraction of smooth muscle
Thrombocyte aggregation
Biosynthesis of steroid hormones
Pain production
Gastric juice secretion
Inflammatory response
Prostaglandin H-synthase [heme]
1
Phospholipase A2 3.1.1.4
2
(dioxygenase + peroxidase) 1.14.99.1
3
Arachidonate lipoxygenases 1.13.11. n
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
392
Hormones
STPs (2). Class I cytokine receptors interact
Cytokines
with three different STPs (gp130, βc, and γc).
The STPs themselves do not bind cytokines,
A. Cytokines
but conduct the signal to tyrosine kinases (3).
Cytokines are hormone-like peptides and pro-
The fact that different cytokines can activate
teins with signaling functions, which are syn-
the same STP via their receptors explains the
thesized and released by cells of the immune
overlapping biological activity of some cyto-
system and other cell types. Their numerous
kines.
biological functions operate in three areas:
As an example of the signal transduction
they regulate the development and homeosta-
pathway in cytokines, the illustration shows
sis of the immune system; they control the
the way in which the IL-6 receptor, after bind-
hematopoietic system; and they are involved
ing its ligand IL-6 (1), induces the dimeriza-
in non-specific defense, influencing inflamma-
tion of the STP gp130 (2). The dimeric gp130
tory processes, blood coagulation, and blood
binds cytoplasmic tyrosine kinases from the
pressure. In general, cytokines regulate the
Jak family (“Janus kinases,” with two kinase
growth, differentiation, and survival of cells.
centers) and activates them (3). The Janus
They are also involved in regulating apoptosis
kinases phosphorylate cytokine receptors,
(see p. 396).
STPs, and various cytoplasmic proteins that
There is an extremely large number of cy-
conduct the signal further. In addition, they
tokines; only the most important representa-
phosphorylate transcription factors known as
tives are listed opposite. The cytokines
STATs (“signal transducers and activators of
include interleukins (IL), lymphokines, mono-
transcription”). STATs are among the proteins
kines, chemokines, interferons (IFN), and col-
that have an SH2 domain and are able to bind
ony-stimulating factors (CSF). Via interleukins,
phosphotyrosine residues (see p. 388). They
immune cells stimulate the proliferation and
therefore bind to cytokine receptors that have
activity of other immune cells (see p. 294).
been phosphorylated by Janus kinases. When
Interferons are used medically in the treat-
STATs are then also phosphorylated them-
ment of viral infections and other diseases.
selves (4), they are converted into their active
Although cytokines rarely show structural
form and become dimers (5). After transfer to
homologies with each other, their effects are
the nucleus, they bind—along with auxiliary
often very similar. The cytokines differ from
proteins as transcription factors—to the pro-
hormones (see p. 370) only in certain re-
moters of inducible genes and in this way
spects: they are released by many different
regulate their transcription (6).
cells, rather than being secreted by defined
The activity of the cytokine receptors is
glands, and they regulate a wider variety of
terminated by protein phosphatases, which
target cells than the hormones.
hydrolytically cleave the phosphotyrosine
residues. Several cytokine receptors are able
to lose their ligand-binding extracellular do-
B. Signal transduction in the cytokines
main by proteolysis (not shown). The extra-
As peptides or proteins, the cytokines are hy-
cellular domain then appears in the blood,
drophilic signaling substances that act by
where it competes for cytokines. This reduces
binding to receptors on the cell surface (see
the effective cytokine concentration.
p. 380). Binding of a cytokine to its receptor
(1) leads via several intermediate steps (2 -5)
to the activation of transcription of specific
genes (6).
In contrast to the receptors for insulin and
growth factors (see p. 388), the cytokine re-
ceptors (with a few exceptions) have no ty-
rosine kinase activity. After binding of cyto-
kine (1), they associate with one another to
form homodimers, join together with other
signal transduction proteins (STPs) to form
dimers, or promote dimerization of other
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Other signaling substances
393
A. Cytokines
IL-1
Interleukin 1
G-CSF
Granulocyte colony-stimulating factor
IL-2
Interleukin 2
GM-CSF
Granulocyte/macrophage colony-stimulating
IL-3
Interleukin 3
factor
IL-4
Interleukin 4
MIF
Macrophage migration inhibitory factor
IL-5
Interleukin 5
M-CSF
Monocyte colony-stimulating factor
IL-6
Interleukin 6
TNFα
Tumor necrosis factor- α
IFN-α
Interferon α
TNFβ
Tumor necrosis factor- β
IFN-β
Interferon β
IFN-γ
Interferon γ
and others
Immune system
Hematopoietic system
Cytokine
Nonspecific defense
Secretion by
Signal peptide
Effects on many
individual cells
or signal protein
cell types
B. Signal transduction in the cytokines
gp 130
Phosphorylation of STAT
ATP
ATP
3
4
5
ADP
Janus
kinase
STAT
ADP
IL-6
gp 130
1
Dimerization of STAT
STAT
SH2 domain
can bind
Tyr
phospho-
tyrosine
Janus kinase
residues
2
4
IL-6
3
4
Phosphorylation
Dimerization
receptor
5
STAT
5
STAT dimer
Phospho-
STAT
tyrosine
dimer
residue
6
Nucleus
Transcriptional control
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
394
Growth and development
The above outline of cell cycle progression
Cell cycle
can be examined here in more detail using the
G2-M transition as an example.
A. Cell cycle
Entry of animal cells into mitosis is based
Proliferating cells undergo a cycle of division
on the “mitosis-promoting factor” (MPF). MPF
(the cell cycle), which lasts approximately
consists of CDK1 (cdc2) and cyclin B. The in-
24 hours in mammalian cells in cell culture.
tracellular concentration of cyclin B increases
The cycle is divided into four different phases
constantly until mitosis starts, and then de-
(G1, S, G2, and M—in that sequence).
clines again rapidly (top left). MPF is initially
Fully differentiated animal cells only divide
inactive, because CDK1 is phosphorylated and
rarely. These cells are in the so-called G0
cyclin B is dephosphorylated (top center). The
phase, in which they can remain permanently.
M phase is triggered when a protein phos-
Some G0 cells return to the G1 phase again
phatase [1] dephosphorylates the CDK while
under the influence of mitogenic signals
cyclin B is phosphorylated by a kinase [2]. In
(growth factors, cytokines, tumor viruses,
its active form, MPF phosphorylates various
etc.), and after crossing a control point (G1 to
proteins that have functions in mitosis—e. g.,
S), enter a new cycle. DNA is replicated (see
histone H1 (see p. 238), components of the
p. 240) during the S phase, and new chroma-
cytoskeleton such as the laminins in the nu-
tin is formed. Particularly remarkable in mor-
clear membrane, transcription factors, mitotic
phological terms is the actual mitosis
(M
spindle proteins, and various enzymes.
phase), in which the chromosomes separate
When mitosis has been completed, cyclin B
and two daughter cells are formed. The M and
is marked with ubiquitin and broken down
S phases are separated by two segments
proteolytically by proteasomes (see p. 176).
known as the G1 and G2 phases (the G stands
Protein phosphatases then regain control
for “gap”). In the G1 phase, the duration of
and dephosphorylate the proteins involved
which can vary, the cell grows by de novo
in mitosis. This returns the cell to the inter-
synthesis of cell components. Together, the
phase.
G1, G0, S, and G2 phases are referred to as
the interphase, which alternates in the cell
Further information
cycle with the short M phase.
The G1-S transition (not shown) is particu-
larly important for initiating the cell cycle. It
B. Control of the cell cycle
is triggered by the CDK4-cyclin D complex,
The progression of the cell cycle is regulated
which by phosphorylating the protein pRb
by interconversion processes. In each phase,
releases the transcription factor E2F previ-
special Ser/Thr-specific protein kinases are
ously bound to pRb. This activates the tran-
formed, which are known as cyclin-depen-
scription of genes needed for DNA replication.
dent kinases (CDKs). This term is used be-
If the DNA is damaged by mutagens or ion-
cause they have to bind an activator protein
izing radiation, the protein p53 initially delays
(cyclin) in order to become active. At each
entry into the S phase. If the DNA repair sys-
control point in the cycle, specific CDKs asso-
tem (see p. 256) does not succeed in remov-
ciate with equally phase-specific cyclins. If
ing the DNA damage, p53 forces the cell into
there are no problems (e. g., DNA damage),
apoptosis (see p. 396). The genes coding for
the CDK-cyclin complex is activated by phos-
pRb and p53 belong to the tumor-suppressor
phorylation and/or dephosphorylation. The
genes
(see p. 398). In many tumors
(see
activated complex in turn phosphorylates
p. 400), these genes are in fact damaged by
transcription factors, which finally lead to
mutation.
the formation of the proteins that are re-
quired in the cell cycle phase concerned (en-
zymes, cytoskeleton components, other CDKs,
and cyclins). The activity of the CDK-cyclin
complex is then terminated again by pro-
teolytic cyclin degradation.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Cell proliferation
395
A. Cell cycle
M phase
G1 phase
0 h
Mitosis
RNA and protein
Chromosome
synthesis
separation
Cell growth
Cell division
G2 phase
G0 phase
Preparation
for mitosis
12h
4 h
No cell
division
S phase
DNA replication
Restriction
Histone synthesis
point
Centrosome formed
Chromosome
8 h
duplication
B. Control of the cell cycle
Reactions early
Cyclin B concentration
M = Mitosis
in mitosis
Reactions late
in mitosis
M
M
M
M
Time
Cyclin-dependent
Cyclin B
protein kinase (CDK1)
Cyclin-
dependent
Catalytic
Regulatory
subunit
protein
P
subunit
kinases
Cyclin
A
CDK 1 - 6
P P
P
fragments
Inactive
P
protein
Cyclin
1
kinase
2
Proteolysis
A
A
Cyclins A - E
P P P
P P
Interphase
P
Mitosis
P
MPF
P
Active protein kinase
Protein
Histone H1
Spindle formation
Laminin
1
Chromosome
Protein kinases
condensation
Transcription
P
Disappearance of
factors
Phosphoprotein
nuclear membrane
1
phosphatase
Transcription stop
Other proteins
Cyclin degradation
2
Protein kinase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
396
Growth and development
p. 290). Other enzymes in this group, known
Apoptosis
as effector caspases, cleave cell components
after being activated—e. g., laminin in the nu-
A. Cell proliferation and apoptosis
clear membrane and snRP proteins
(see
The number of cells in any tissue is mainly
p. 246)—or activate special DNases which
regulated by two processes—cell proliferation
then fragment the nuclear DNA.
and physiological cell death, apoptosis. Both of
An important trigger for apoptosis is
these processes are regulated by stimulatory
known as the Fas system. This is used by
and inhibitory factors that act in solute form
cytotoxic T cells, for example, which eliminate
(growth factors and cytokines) or are pre-
infected cells in this way (top left). Most of the
sented in bound form on the surface of neigh-
body’s cells have Fas receptors (CD 95) on
boring cells (see below).
their plasma membrane. If a T cell is activated
Apoptosis is genetically programmed cell
by contact with an MHC presenting a viral
death, which leads to “tidy” breakdown and
peptide
(see p. 296), binding of its Fas
disposal of cells. Morphologically, apoptosis is
ligands occurs on the target cell’s Fas recep-
characterized by changes in the cell mem-
tors. Via the mediator protein FADD (“Fas-
brane (with the formation of small blebs
associated death domain”), this activates cas-
known as “apoptotic bodies”), shrinking of
pase-8 inside the cell, setting in motion the
the nucleus, chromatin condensation, and
apoptotic process.
fragmentation of DNA. Macrophages and
Another trigger is provided by tumor ne-
other phagocytic cells recognize apoptotic
crosis factor-D (TNF-α), which acts via a sim-
cells and remove them by phagocytosis with-
ilar protein (TRADD) and supports the endog-
out inflammatory phenomena developing.
enous defense system against tumors by in-
Cell necrosis (not shown) should be distin-
ducing apoptosis.
guished from apoptosis. In cell necrosis, cell
Caspase-8 activates the effector caspases
death is usually due to physical or chemical
either directly, or indirectly by promoting
damage. Necrosis leads to swelling and burst-
the cytochrome c (see p. 140) from mitochon-
ing of the damaged cells and often triggers an
dria. Once in the cytoplasm, cytochrome c
inflammatory response.
binds to and activates the protein Apaf-1
The growth of tissue (or, more precisely,
(not shown) and thus triggers the caspase
the number of cells) is actually regulated by
cascade. Apoptotic signals can also come
apoptosis. In addition, apoptosis allows the
from the cell nucleus. If irreparable DNA dam-
elimination of unwanted or superfluous
age is present, the p53
protein
(see
cells—e. g., during embryonic development
p. 394)—the product of a tumor suppressor
or in the immune system. The contraction of
gene—promotes apoptosis and thus helps
the uterus after birth is also based on apop-
eliminate the defective cell.
tosis. Diseased cells are also eliminated by
There are also inhibitory factors that op-
apoptosis—e. g., tumor cells, virus-infected
pose the signals that activate apoptosis. These
cells, and cells with irreparably damaged
include bcl-2 and related proteins. The ge-
DNA. An everyday example of this is the peel-
nomes of several viruses include genes for
ing of the skin after sunburn.
this type of protein. The genes are expressed
by the host cell and (to the benefit of the
virus) prevent the host cell from being pre-
B. Regulation of apoptosis
maturely eliminated by apoptosis.
Apoptosis can be triggered by a number of
different signals that use various transmission
pathways. Other signaling pathways prevent
apoptosis.
At the center of the apoptotic process lies a
group of specialized cysteine-containing as-
partate proteinases (see p. 176), known as cas-
pases. These mutually activate one another,
creating an enzyme cascade resembling the
cascade involved in blood coagulation (see
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Cell proliferation
397
A. Cell proliferation and apoptosis
Factors
Factors
Phagocytic
macrophage
Cell proliferation
Apoptosis
Constant cell number
Changes in
Dissolution
membranes
of nuclear structure
Shrinking of
Condensation
cytoplasm
of chromatin
Fragmentation
of DNA
Apoptotic cell
B. Regulation of apoptosis
Cytotoxic T cell
Fas ligand
TNF-α
TNF
receptor
Fas
type I
receptor
Caspase 8
Mitochondrion
FADD
Cytochrome c
TRADD
Effector
caspases
p53 protein
bcl-2 protein
cleave
snRNA
Laminin
Caspase-activated
Other proteins
proteins
DNAase
DNA with
irradiation
damage
Apoptosis
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
398
Growth and development
usually indicate the origin of the viral gene
Oncogenes
and are printed in italics (e. g., myc for mye-
Oncogenes are cellular genes that can trigger
locytomatosis, a viral disease in birds). Onco-
uncontrolled cell proliferation if their se-
gene products can be classified into the fol-
quence is altered or their expression is incor-
lowing groups according to their functions.
rectly regulated. They were first discovered as
1. Ligands such as growth factors and
viral (v-) oncogenes in retroviruses that cause
cytokines, which promote cell proliferation.
tumors (tumor viruses). Viruses of this type
2. Membrane receptors of the 1-helix type
(see p. 404) sometimes incorporate genes
with tyrosine kinase activity, which can
from the host cell into their own genome. If
bind growth factors and hormones (see
these genes are reincorporated into the host
p. 394).
DNA again during later infection, tumors can
3. GTP-binding proteins. This group includes
then be caused in rare cases. Although virus-
the G proteins in the strict sense and re-
related tumors are rare, research into them
lated proteins such as Ras (see p. 388), the
has made a decisive contribution to our
product of the oncogene c-ras.
understanding of oncogenes and their func-
4. Receptors for lipophilic hormones mediate
tioning.
the effects of steroid hormones and related
signaling substances. They regulate the
A. Proto-oncogenes: biological role
transcription of specific genes
(see
p. 378). The products of several oncogenes
The cellular form of oncogenes (known as c-
(e. g., erbA) belong to this superfamily of
oncogenes or proto-oncogenes) code for pro-
ligand-controlled transcription factors.
teins involved in controlling growth and dif-
5. Nuclear tumor suppressors inhibit return
ferentiation processes. They only become on-
to the cell cycle in fully differentiated cells.
cogenes if their sequence has been altered by
The genes that code for these proteins are
mutations (see p. 256), deletions, and other
referred to as anti-oncogenes due to this
processes, or when excessive amounts of the
function. On the role of p53 and pRb, see
gene products have been produced as a result
p.394.
of overexpression.
6. DNA-binding proteins. A whole series of
Overexpression can occur when amplifica-
oncogenes code for transcription factors.
tion leads to numerous functional copies of
Particularly important for cell proliferation
the respective gene, or when the gene falls
are myc, as well as fos and jun. The protein
under the influence of a highly active pro-
products of the latter two genes form the
moter (see p. 244). If the control of oncogene
transcription factor AP-1 as a heterodimer
expression by tumor suppressor genes (see
(see p. 244).
p. 394) is also disturbed, transformation and
7.
Protein kinases play a central role in intra-
unregulated proliferation of the cells can oc-
cellular signal transduction. By phosphor-
cur. A single activated oncogene does not
ylating proteins, they bring about altera-
usually lead to a loss of growth control. It
tions in biological activity that can only
only occurs when over the course of time
be reversed again by the effects of protein
mutations and regulation defects accumulate
phosphatases. The interplay between pro-
in one and the same cell. If the immune sys-
tein phosphorylation by protein kinases
tem does not succeed in eliminating the
and dephosphorylation by protein phos-
transformed cell, it can over the course of
phatases (interconversion) serves to regu-
months or years grow into a macroscopically
late the cell cycle (see p. 394) and other
visible tumor.
important processes. The protein kinase
Raf is also involved in the signal transduc-
B. Oncogene products:
tion of insulin (see p. 388).
biochemical functions
A feature common to all oncogenes is the fact
that they code for proteins involved in signal
transduction processes. The genes are desig-
nated using three-letter abbreviations that
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Cell proliferation
399
A. Proto-oncogenes: biological role
Transformation
Tumor
Altered
formation
Defective
proteins
control
Tumor virus
Altered
control
protein
Oncogene
Defective
suppressor gene
Reverse
transcription
Mutation, deletion,
Tumor
incorporation
amplification,
initiation
altered control
v-Oncogene
Tumor
Proto-oncogene
suppressor
gene
Control protein
e.g., p53 protein
Normal
Normal
Rb protein
development
growth and
differentiation
B. Oncogene products: biochemical functions
Oncogene products
1
(examples)
Effector
Receptor
3
enzyme
2
Gene
1
Ligands
GTP
P
G protein
sis, hst, int-2, wnt-1
Second
messenger
2
Receptors
Voltage-
fms, trk, trkB, ros, kit,
Other
Protein
mas, neu, erbB
gated
effects
kinase
ion channel
Ca2
3
GTP-binding proteins
7
Ha-ras, Ki-ras, N-ras
Nuclear hormone
4
7
P
receptors
Calmodulin
erbA, NGF1-B
P
Phospho-
Protein
rylated
Ligand-
Transcription
phospha-
protein
5
Nuclear tumor
gated
factor
tase
ion channel
5
6
suppressors
Rb, p53, wt1, DCC, APC
4
6
DNA-binding proteins
jun, fos, myc, N-myc,
Intra-
Hormone
GTP
myb, fra1, egr-1, rel
receptor
cellular
Transcription
G protein
3
7
Protein kinases
src, yes, fps, abl, met,
mos, raf
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
400
Growth and development
tumor cells (group 1) or are induced by them
Tumors
in other cells
(group
2). Group
1 tumor
markers include tumor-associated antigens,
A. Division behavior of cells
secreted hormones, and enzymes. The table
The body’s cells are normally subject to strict
lists a few examples.
“social” control. They only divide until they
The transition from a normal to a trans-
come into contact with neighboring cells; cell
formed state is a process involving several
division then ceases due to contact inhibition.
steps.
Exceptions to this rule include embryonic
1. Tumor initiation. Almost every tumor be-
cells, cells of the intestinal epithelium (where
gins with damage to the DNA of an individual
the cells are constantly being replaced), cells
cell. The genetic defect is almost always caused
in the bone marrow (where formation of
by environmental factors. These can include
blood cells takes place), and tumor cells. Un-
tumor-inducing chemicals (carcinogens—e. g.,
controlled cell proliferation is an important
components of tar from tobacco), physical
indicator of the presence of a tumor. While
processes (e. g., UV light, X-ray radiation; see
normal cells in cell culture only divide 20-60
p. 256), or in rare cases tumor viruses (see
times, tumor cells are potentially immortal
p. 398). Most of the approximately 1014 cells
and are not subject to contact inhibition.
in the human body probably suffer this type
In medicine, a distinction is made between
of DNA damage during the average lifespan,
benign and malignant tumors. Benign tumors
but it is usually repaired again (see p. 256). It
consist of slowly growing, largely differenti-
is mainly defects in proto-oncogenes
(see
ated cells. By contrast, malignant tumors
p. 398) that are relevant to tumor initiation;
show rapid, invasive growth and tend to
these are the decisive cause of transformation.
form metastases (dissemination of daughter
Loss of an anti-oncogene (a tumor-suppressor
lesions). The approximately
100
different
gene) can also contribute to tumor initiation.
types of tumor that exist are responsible for
2. Tumor promotion is preferential prolif-
more than 20% of deaths in Europe and North
eration of a cell damaged by transformation. It
America.
is a very slow process that can take many
years. Certain substances are able to strongly
accelerate it—e. g., phorbol esters. These occur
B. Transformation
in plants (e. g., Euphorbia species) and act as
The transition of a normal cell into a tumor
activators of protein kinase C (see p. 386).
cell is referred to as transformation.
3. Tumor progression finally leads to a
Normal cells have all the characteristics of
macroscopically visible tumor as a result of
fully differentiated cells specialized for a par-
growth. When solid tumors of this type ex-
ticular function. Their division is inhibited
ceed a certain size, they form their own vas-
and they are usually in the G0 phase of the
cular network that supplies them with blood
cell cycle (see p. 394). Their external shape is
(angiogenesis). Collagenases (matrix metallo-
variable and is determined by a strongly
proteinases, MMPs) play a special role in the
structured cytoskeleton.
metastatic process, by loosening surrounding
In contrast, tumor cells divide without in-
connective tissue and thereby allowing tumor
hibition and are often de-differentiated—i. e.,
cells to disseminate and enter the blood-
they have acquired some of the properties of
stream. New approaches to combating tumors
embryonic cells. The surface of these cells is
have been aimed at influencing tumor angio-
altered, and this is particularly evident in a
genesis and metastatic processes.
disturbance of contact inhibition by neighbor-
ing cells. The cytoskeleton of tumor cells is
also restructured and often reduced, giving
them a rounded shape. The nuclei of tumor
cells can be atypical in terms of shape, num-
ber, and size.
Tumor markers are clinically important for
detecting certain tumors. These are proteins
that are formed with increasing frequency by
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Cell proliferation
401
A. Division behavior of cells
Growth inhibition
Uncontrolled
due to contacts
cell proliferation
with adjacent cells
Nutrient
medium
Normal cells
Tumor cells
B. Transformation
Normal cell
Indicators:
Differentiated
Tumor initiators
Non-dividing
H3C
Defined form
Viruses
Carcinogenic
chemicals
Tumor initiation:
1
Genetic damage
UV
Physical processes
OH
Tumor promoters
OH
Tumor progression:
H3C
CH3
2
Preferential propagation
e.g. Esters
H
CH3
of phorbol
H3C
OH
H
Indicators:
O HO
Hormones
CH2OH
De-differentiated
Phorbol
Tumor cell
Uncontrolled cell division
Altered cell surface
Altered cytoskeleton
Tumor markers (examples)
and nucleus
Tumor-associated antigens
CEA
Carcinoembryonic
antigen
Tumor progression:
3
Acquisition of malignancy
AFP
α1-Fetoprotein
Hormones Calcitonin
ACTH
Enzymes Acid phosphatase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
402
Growth and development
The cytostatic drugs administered (indi-
Cytostatic drugs
cated by a syringe in the illustration) are often
Tumors (see p. 400) arise from degenerated
not active themselves but are only converted
(transformed) cells that grow in an uncon-
into the actual active agent in the metabolism.
trolled way as a result of genetic defects.
This also applies to the adenine analogue 6-
Most transformed cells are recognized by
mercaptopurine, which is initially converted
the immune system and eliminated
(see
to the mononucleotide tIMP
(thioinosine
p. 294). If endogenous defense is not suf -
monophosphate). Via several intermediate
ciently effective, rapid tumor growth can oc-
steps, tIMP gives rise to tdGTP, which is in-
cur. Attempts are then made to inhibit growth
corporated into the DNA and leads to cross-
by physical or chemical treatment.
links and other anomalies in it. The second
A frequently used procedure is targeted
effective metabolite of 6-mercaptopurine is
irradiation with γ-rays, which block cell re-
S-methylated tIMP, an inhibitor of amidophos-
production due their mutagenic effect (see
phoribosyl transferase (see p. 188).
p. 256). Another approach is to inhibit cell
Hydroxyurea selectively inhibits ribonu-
growth by chemotherapy. The growth-inhib-
cleotide reductase (see p. 190). As a radical
iting substances used are known as cytostatic
scavenger, it removes the tyrosine radicals
drugs. Unfortunately, neither radiotherapy
that are indispensable for the functioning of
nor chemotherapy act selectively—i. e., they
the reductase.
damage normal cells as well, and are there-
Two other important cytostatic agents tar-
fore often associated with severe side effects.
get the synthesis of DNA-typical thymine,
Most cytostatic agents directly or indirectly
which takes place at the level of the deoxy-
inhibit DNA replication in the S phase of the
mononucleotide (see p. 190). The deoxymo-
cell cycle (see p. 394). The first group (A) lead
nonucleotide formed by 5-fluorouracil or the
to chemical changes in cellular DNA that im-
corresponding nucleoside inhibits thymidy-
pede transcription and replication. A second
late synthase. This inhibition is based on the
group of cytostatic agents (B) inhibit the syn-
fact that the fluorine atom in the pyrimidine
thesis of DNA precursors.
ring cannot be substituted by a methyl group.
In addition, the fluorine analogue is also in-
corporated into the DNA.
A. Alkylating agents, anthracyclines
Dihydrofolate reductase acts as an auxiliary
Alkylating agents are compounds capable of
enzyme for thymidylate synthase. It is in-
reacting covalently with DNA bases. If a com-
volved in the regeneration of the coenzyme
pound of this type contains two reactive
N5,N10-methylene-THF, initially reducing DHF
groups, intramolecular or intermolecular
to THF with NADPH as the reductant (see
crosslinking of the DNA double helix and
p. 418). The folic acid analogue methotrexate,
“bending” of the double strand occurs. Exam-
a frequently used cytostatic agent, is an ex-
ples of this type shown here are cyclophos-
tremely effective competitive inhibitor of di-
phamide and the inorganic complex cisplatin.
hydrofolate reductase. It leads to the deple-
Anthracyclines such as doxorubicin (adriamy-
tion of N5,N10-methylene-THF in the cells and
cin) insert themselves non-covalently be-
thus to cessation of DNA synthesis.
tween the bases and thus lead to local alter-
ations in the DNA structure (see p. 254 B).
Further information
To reduce the side effects of cytostatic agents,
B. Antimetabolites
new approaches are currently being devel-
Antimetabolites are enzyme inhibitors
(see
oped on the basis of gene therapy
(see
p. 96) that selectively block metabolic pathways.
p. 264). Attempts are being made, for exam-
The majority of clinically important cytostatic
ple, to administer drugs in the form of pre-
drugs act on nucleotide biosynthesis. Many of
cursors (known as prodrugs), which only be-
these are modified nucleobases or nucleotides
come active in the tumor itself (“tumor tar-
that competitively inhibit their target enzymes
geting”).
(see p. 96). Many are also incorporated into the
DNA, thereby preventing replication.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Cell proliferation
403
A. Alkylating agents, anthracyclines
Cross-linking
“Bending”
of DNA
of the DNA
components
double helix
O
OH
O
H
N
C
H3N
Cl
CH
CH2
Cl
2
CH2OH
O
P
OH
Pt
CH2
CH2
Cl
H3N Cl
H O
H3CO O OH
Cyclophosphamide
Adriamycin
R
Cisplatin
B. Antimetabolites
S
CH3
SH
SH
C
C
C
N
N
N
N C
N
C
N
C
CH
2
CH
1
CH
HC
C
HC
C
HC
C
N
N
N
N
N
N
SAH
SAM
P
Rib
P
Rib
me-tIMP
tIMP
6-Mercaptopurine
GIn
Glu
+
P P
tIMP
tGMP
tGDP
tdGDP
Purine
DNA
Phospho-
synthesis
ribosyl
IMP
GMP
GDP
4
dGDP
3
PRPP
amine
O
Hypoxanthine phospho-
Amidophosphoribosyl
1
3
C
OH
ribosyltransferase 2.4.2.8
transferase 2.4.2.14
H2N
N
Thiopurine methyl-
Ribonucleoside diphosphate
2
4
H
transferase2.1.1.67
reductase 1.17.4.1
Hydroxyurea
Precursors
O
H2N
N
N
N5,N10
-methylene-THF
C
C
CH
C
F
HN
C
N
C
C
H
H
C
C
N
CH2
dUMP
C
CH
N
A
O
N
H3C
N
THF
NH2
P
P
Rib
5
6
5-fluoro-
N
A
Methotrexate
deoxyuridine
DHF
P
(amethopterin)
monophosphate
C O
dTMP
HN
Dihydrofolate
OOC
CH
CH2
C
COO
2
dTTP
H
5-fluorouracil
5-fluoro-
deoxyuridine
5
Thymidylate synthase 2.1.1.45
DNA
6
Dihydrofolate reductase 1.5.1.3
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
404
Growth and development
other to form pentamers and hexamers. In all,
Viruses
60 protein molecules are involved in the
Viruses are parasitic nucleoprotein complexes.
structure of the capsid.
They often consist of only a single nucleic acid
molecule (DNA or RNA, never both) and a
C. Life cycle of HIV
protein coat. Viruses have no metabolism of
their own, and can therefore only replicate
The human immunodeficiency virus
(HIV)
themselves with the help of host cells. They
causes the immunodeficiency disease known
are therefore not regarded as independent
as AIDS
(acquired immune deficiency syn-
organisms. Viruses that damage the host cell
drome). The structure of this virus is similar
when they replicate are pathogens. Diseases
to that of the influenza virus (A).
caused by viruses include AIDS, rabies, polio-
The HIV genome consists of two molecules
myelitis, measles, German measles, smallpox,
of ssRNA (each 9.2 kb). It is enclosed by a
influenza, and the common cold.
double-layered capsid and a protein-contain-
ing coating membrane. HIV mainly infects T
helper cells (see p. 294) and can thereby lead
A. Viruses: examples
to failure of the immune system in the longer
Only a few examples from the large number
term.
of known viruses are illustrated here. They are
During infection
(1), the virus’s coating
all shown on the same scale.
membrane fuses with the target cell’s plasma
Viruses that only replicate in bacteria are
membrane, and the core of the nucleocapsid
known as bacteriophages (or “phages” for
enters the cytoplasm (2). In the cytoplasm,
short). An example of a phage with a simple
the viral RNA is initially transcribed into an
structure is M13. It consists of a single-
RNA/DNA hybrid (3) and then into dsDNA (4).
stranded DNA molecule (ssDNA) of about
Both of these reactions are catalyzed by re-
7000 bp with a coat made up of 2700 helically
verse transcriptase, an enzyme deriving from
arranged protein subunits. The coat of a virus
the virus. The dsDNA formed is integrated
is referred to as a capsid, and the complete
into the host cell genome (5), where it can
structure as a nucleocapsid. In genetic engi-
remain in an inactive state for a long time.
neering, M13 is important as a vector for for-
When viral replication occurs, the DNA
eign DNA (see p. 258).
segment corresponding to the viral genome
The phage T4 (bottom left), one of the larg-
is first transcribed by host cell enzymes (6).
est viruses known, has a much more complex
This gives rise not only to viral ssRNA, but also
structure with around 170 000 base pairs (bp)
to transcription of mRNAs for precursors of
of double-stranded DNA (dsDNA) contained
the viral proteins (7). These precursors are
within its “head.”
integrated into the plasma membrane (8, 9)
The tobacco mosaic virus (center right), a
before undergoing proteolytic modification
plant pathogen, has a structure similar to that
(10). The cycle is completed by the release of
of M13, but contains ssRNA instead of DNA.
new virus particles (11).
The poliovirus, which causes poliomyelitis, is
The group of RNA viruses to which HIV
also an RNA virus. In the influenza virus, the
belongs are called retroviruses, because DNA
pathogen that causes viral flu, the nucleocap-
is produced from RNA in their replication cy-
sid is additionally surrounded by a coat de-
cle—the reverse of the usual direction of tran-
rived from the plasma membrane of the host
scription (DNA RNA).
cell (C). The coat carries viral proteins that are
involved in the infection process.
B. Capsid of the rhinovirus
Rhinoviruses cause the common cold. In these
viruses, the capsid is shaped like an icosahe-
dron—i. e., an object made up of 20 equilateral
triangles. Its surface is formed from three dif-
ferent proteins, which associate with one an-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Viruses
405
A. Viruses: examples
B. Capsid of the rhinovirus
Phage M13
ssDNA 7 kb
Helical
Coat
T4-Phage
ds DNA
Influenza virus
170 kbp
ssRNA (8 molecules) 3.6 kb
Eicosa-
complex
Nucleocapsid with coat
hedral
structure
head
Tail
1. Structure
Tobacco mosaic virus
ssRNA6.4 kb
Pentamer
Helical
Poliovirus
Phage DNA
ssRNA 7 kb
Bacterial cell
Eicosahedral capsid
30 nm
Hexamer
Capsid
2. Plant and animal
Eicosahedron of
180 monomers
1. Bacteriophages
pathogenic viruses
2. Diagram
C. Life cycle of the human immunodeficiency virus (HIV)
Glycoprotein
Other
GP120
enzymes
Infection
1
100 nm
2
Reverse
transcriptase
Core
Viral
RNA
Membrane
Viral
RNA
8
1
3
GP120
Precursors
of core proteins
RNA/
and enzymes
DNA
7
Translation
hybrid
2
mRNA
4
Viral
RNA
9
1
ds
Transcription
6
Mature
11
DNA
virus
5
particle
10
Integration
Reverse transcriptase
1
2.7.7.49
Cytoplasm
Ribonuclease H
Nucleus
Host DNA
2
3.1.26.4
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
406
Metabolic charts
Example
Metabolic charts
On p. 407, the initial step of the dark reactions
Explanations
in plant photosynthesis (in the Calvin cycle) is
shown at the top left.
The following 13 plates (pp. 407-419) provide
a concise schematic overview of the most
important metabolic pathways. Explanatory
text is deliberately omitted from them.
These “charts”:
• Contain details of metabolic pathways that
are only shown in outline in the main text
for reasons of space. This applies in partic-
ular to the synthesis and degradation of the
amino acids and nucleotides, and for some
aspects of carbohydrate and lipid metabo-
lism.
• Offer a quick overview of a specific path-
way, the metabolites that arise in it, and the
enzymes involved.
• Can be used for reference purposes and for
In this reaction, one molecule of ribulose-
revising material previously learned.
1,5-bisphosphate (metabolite 1) and one mol-
ecule of CO2 (metabolite 2) give rise to two
The most important intermediates are shown
molecules of 3-phosphoglycerate (metabolite
with numbers in the charts. The correspond-
3). The enzyme responsible has the EC
ing compounds can be identified using the
number 4.1.1.39. The annotated enzyme list
table on the same page.
shows that this refers to ribulose bisphosphate
In addition, at each step the four-figure EC
carboxylase (“rubisco” for short). Rubisco be-
number (see p. 88) for the enzyme responsi-
longs to enzyme class 4 (the lyases) and,
ble for a reaction is given in italics. The en-
within that group, to subclass 4.1 (the car-
zyme name and its systematic classification in
boxy-lyases). It contains copper as a cofactor
the system used by the Enzyme Catalogue are
([Cu]).
available in the following annotated enzyme
list (pp. 420-430), in which all of the enzymes
mentioned in this book are listed according to
their EC number. The book’s index is helpful
when looking for a specific enzyme in the
text.
In reactions that involve coenzymes, the
coenzyme names are also given (sometimes
in simplified form). Particularly important
starting, intermediate, or end products are
given with the full name, or as formulae.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic charts
407
A. Calvin cycle (plant chloroplasts)
6 CO2 + 18 ATP + 12 NADPH + 12 H
Hexose + 18 ADP + 18 P + 12 NADP
2x
5
5
5
5
5
5
Ribulose 1,5 - bis P
CO2
5.3.1.1
1
2
C3
4.1.1.39
6
6
3
3
4.1.2.13
ATP
ATP
2
ADP
7
2.7.2.3
3.1.3.11
C6
4
4
P
NADPH
NADPH
8
2
NADP
P
P
2.2.1.1
5
1.2.1.13
5
9
C4
15
6x
Glucose 6-
P
Glyceraldehyde 3-
phosphate
1
Ribulose 1,5-bisphosphate
10
2
Carbon dioxide
3
3-Phosphoglycerate
C7
3.1.3.37
P
4
1,3-Bisphosphoglycerate
5
Glyceraldehyde 3-phosphate
11
2.2.1.1
6
Dihydroxyacetone phosphate
7
Fructose 1,6-bisphosphate
8
Fructose 6-phosphate
12
C
12
C5
13
5
9
Erythrose 4-phosphate
10
Sedoheptulose 1,7-bisphosphate
5.1.3.4
5.3.1.6
11
Sedoheptulose 7-phosphate
12
Xylulose 5-phosphate
C5
C5
14
14
14
13
Ribose 5-phosphate
ATP
ATP
ATP
14
Ribulose 5-phosphate
ADP
ADP
ADP
15
2.7.1.19
2.7.1.19
Glucose 6-phosphate
1
1
1
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
408
Metabolic charts
A. Carbohydrate metabolism
5.1.3.1
5.3.1.6
Ribose 5-
8
9
10
10
Glycogen
phosphate
CO2
NADPH
1
P
2.2.1.1
NADP
UDP
7
11
12
2
H2O
2.2.1.2
P P
6
13
UTP
NADPH
5.4.2.2
2.2.1.1
3
P
P
NADP
Glucose
4
5
14
15
C6
3.1.3.9
5.3.1.9
3.1.3.11
Glucose 6-
2.7.1.1
phosphate
2.7.1.11
ATP
ADP
ATP
ADP
P
4.1.2.13
5.4.2.1
2.7.2.3
1.2.1.12
5.3.1.1
19
18
17
12
16
NADH
ATP
ADP
NADH
NAD
ADP
ATP
C3
H2O
NAD
Pyruvate
2.7.1.40
1.1.1.27
20
21
22
Lactate
25
CO2
GDP
ADP
ATP
Mitochondrion
GTP
Amino
Amino
23
23
21
Glycerol
24
acids
acids
6.4.1.1
C4
ADP
ATP
Fat
1
Glycogen
9
Ribose 5-phosphate
17
1,3-Bisphosphoglycerate
2
UDP-Glucose
10
Xylulose 5-phosphate
18
3-Phosphoglycerate
3
Glucose 1-phosphate
11
Sedoheptulose 7-phosphate
19
2-Phosphoglycerate
4
Glucose
12
Glyceraldehyde 3-phosphate
20
Phosphoenolpyruvate
5
Glucose 6-phosphate
13
Erythrose 4-phosphate
21
Pyruvate
22
Lactate
6
Gluconolactone 6-
P
14
Fructose 6-phosphate
23
Oxaloacetate
7
Gluconate 6-phosphate
15
Fructose 1,6-bisphosphate
24
Glycerol
8
Ribulose 5-phosphate
16
Glycerone-3-phosphate
25
Glycerol 3-phosphate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic charts
409
A. Biosynthesis of fats and membrane lipids
1
Glycolysis
2
ATP
ADP
NAD
NADH
Fatty acid
synthase
Acetyl
3
4
5
2
CoA
2.7.1.30
1.1.1.8
CoA
CoA
1.1.1.101
Ketone bodies
Acyl-CoA
6
7
8
Isoprenoids
NADP
NADPH
CoA
1
Pyruvate
2.7.7.41
9
10
2
Dihydroxyacetone 3-
P
Inositol
3
Acetyl CoA
CTP
P
P
CoA
CMP
4
Glycerol
Fat
11
12
5
Glycerol 3-phosphate
2.3.1.20
6
Acyl CoA
CMP
7
1-Acylglycerol 3-phosphate
2.7.1.32
2.7.7.15
8
1-Acylglycerone 3-phosphate
13
14
15
2.7.8.2
9
Phosphatidate
ATP
ADP
CTP
P
P
CMP
10
CDP-diacylglycerol
2.7.1.82
2.7.7.14
11
1,2-Diacylglycerol
16
17
18
2.7.8.1
12
Phosphatidylinositol
22
ATP
ADP
CTP
P
P
13
Choline
16
14
CDP-choline
19
Palmitoyl CoA
20
15
Phosphatidylcholine
16
Ethanolamine
21
Sphingosine
17
CDP-ethanolamine
6
UDP-Glc, UDP-Gal
Serine
22
UDP-GlcNAc
Phosphatidylethanolamine
18
CMP-NeuAc
19
Palmitoyl CoA
Ganglio-
23
Ceramide
sides
20
Phosphatidylserine
15
UDP-Gal
21
Sphingosine
11
UDP
22
Serine
23
Ceramide
Sphingo-
Cere-
24
Galactosylceramide
25
24
myelins
brosides
25
Sphingomyelin
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
410
Metabolic charts
A. Synthesis of ketone bodies and steroids
5a
CO
NADH
NAD
Pyruvate
2
5
6
1
Glycolysis
Glucose
1.1.1.30
NAD
2
Ketone bodies
CoA
NADH
4.1.3.5
2.3.1.16
Bile
4
3
2
salts
NADPH
Acetyl CoA
NADP
Glycine
Fatty acid
β-Oxi-
CoA
Taurine
synthesis
dation
11
10
2
ATP
2 H2O
7
Bile
2 [H]
2
ADP
CoA
acids
O2
Fatty acid
ATP
12
9
extension
ATP
AMP
ADP
8
PP
NADP
NADPH
PP
CO2
Cholesterol
2.5.1.1
2.5.1.10
13
14
15
16
17
2.5.1.21
O2
2 [H]
NADP
NADPH
2 [H] O2
H2O
1.14.99.9
21
20
19
18
1.1.1.51
NADPH
5.3.3.1
Isoprenoids
H2O
2 [H] O2
H2O
2 [H] O2
NADP
Proge-
Testo-
22
26
25
24
sterone
sterone
1.14.99.10
1.14.99.9
O2
O2
“Aromatase”
2 [H]
2 [H]
H2O
Aldosterone
H2O
Hormone
23
Estradiol
27
Cortisol
29
28
1
Pyruvate
10
Oleyl CoA
20
Dehydroepiandrosterone
2
Acetyl CoA
11
Mevalonate
21
Androstene-3,17-dione
3
Acetoacetyl CoA
12
Mevalonate 5-diphosphate
22
Testosterone
3-Hydroxy-3-methyl-
4
13
Isopentenyl diphosphate
23
Estradiol
glutaryl CoA
5
Acetoacetate
14
Geranyl diphosphate
24
Progesterone
5a
Acetone
15
Farnesyl diphosphate
25
17-OH-Progesterone
6
3-Hydroxybutyrate
16
Squalene
26
11-Deoxycortisol
7
Palmitate
17
Cholesterol
27
Cortisol
8
Palmitoyl CoA
18
Pregnenolone
28
11-Deoxycorticosterone
9
Stearoyl CoA
19
17-OH-Pregnenolone
29
Aldosterone
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic charts
411
A. Degradation of fats and phospholipids
Phospholipases
1
7
H2O
H2O
A1
3.1.1.32
A2
3.1.1.4
C
3.1.4.3
2
8
ATP
H2O
D
3.1.4.4
D
CoA
P
P
3
9
C
10
H2O
CoA
A1
A2
4
11
Adipose tissue
4
11
ATP
CoA
From uneven-
numbered fatty acids
10
ADP
1.3.99.3
6.4.1.3
5
9
12
16
17
NAD
H2O
ETFox
ETFred
Glucose
ATP
CO2
ADP
NADH
Respiratory chain
6
13
18
β-Oxidation
NAD
Pyruvate
15
CoA
Respiratory
NADH
chain
9'
14
19
2.3.1.16
1
Triacylglycerol
8
Free fatty acid
14
3-Oxoacyl CoA
2
Diacylglycerol
9
Acyl CoA
15
Acetyl CoA
3
Monoacylglycerol
9'
Shortened acyl CoA
16
Propionyl CoA
4
Glycerol
10
Carnitine
17
(S)-Methylmalonyl CoA
5
Glycerol 3-phosphate
11
Acyl carnitine
18
(R)-Methylmalonyl CoA
6
Glycerone 3-phosphate
12
2,3-Dehydroacyl CoA
19
Succinyl CoA
7
Phospholipid
13
3-Hydroxyacyl CoA
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
412
Metabolic charts
A. Biosynthesis of the essential amino acids
Isoleucine
Leucin
Valine
8
2-Oxo-
OG
glutarate
OG
OG
ATP
Glu
Threonine
Glu
Glu
P
P
4
7
6
2
Pyruvate
CO2
3
1
5
4.1.3.18
ATP
ADP
9
10
Cysteine
Methionine
2.7.2.4
NADPH
[CH3]
Aspartate
NADP
1.1.1.3
2.7.1.39
4.2.99.2
11
12
13
Threonine
H2O
P
NADPH
NADP
ADP
ATP
Glu
OG
Lysine
Histidine
15
P
PEP
14
16
17
18
Glu
8
OG
Trypto-
Phenyl-
Serine
20
19
phan
alanine
1
Pyruvate
8
Phosphoribosyl
15
Erythrose 4-phosphate
diphosphate
2
2-Oxo-3-deoxy-
2-Oxobutyrate
16
9
Aspartate
arabinoheptulosonate
3
2-Aceto-
7-phosphate
2-hydroxybutyrate
10
Aspartyl 4-phosphate
17
Chorismate
4
2-Oxo-4-methylvalerate
11
Aspartate 4-semialdehyde
18
Phenylpyruvate
5
2-Acetolactate
12
Homoserine
19
Anthranilate
6
2-Oxoisovalerate
13
Phosphohomoserine
20
N-(Phosphoribosyl)-
7
2-Oxoisocaproate
14
Phosphoenolpyruvate
anthranilate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic charts
413
A. Biosynthesis of the non-essential amino acids
1.14.16.1
Phenylalanine
Tyrosine
Glycolysis
Tetra-
THB
DHP
O2
H2O
hydro-
biopterin
2.6.1.52
1.1.1.95
3
2
1
2-Oxoglutarate
OG
Glu
NADH
NAD
5,10
THF
M-THF
Glu
OG
4
Serine
Methionine
8
Alanine
2.1.2.1
2.6.1.2
Glycine
Glu
OG
ATP
Gln
Glu
AMP
H2O
6
5
9
10
11
4.2.1.22
2.6.1.1
6.3.5.4
H2O
Aspar-
Aspar-
7
tate
agine
NH3
Tricarboxylic
Cysteine
Proline
acid cycle
NADP
1
3-Phosphoglycerate
2
3-Phosphohydroxypyruvate
NADPH
12
15
3
3-Phosphoserine
NH3
NADP
4
Glycine
NADH
Gluta-
mate
5
Cystathionine
NAD
NADPH
2.7.2.11
6
Homocysteine
13
14
7
2-Oxobutyrate
ATP
8
Pyruvate
ATP
ADP
NH3
Urea
9
Oxaloacetate
cycle
ADP
20
21
10
Aspartate
11
Asparagine
Glutamine
Arginine
Acetyl
12
2-Oxoglutarate
CoA
Ace-
13
Glutamate
tate
CoA
14
γ-Glutamyl phosphate
15
∆1-Pyrroline-5-carboxylate
16
19
ATP
16
N-Acetylglutamate
OG
P
17
N-Acetylglutamate 5-phosphate
ADP
Glu
18
N-Acetylglutamate semialdehyde
1.2.1.38
17
18
19
N-Acetylornithine
20
Ornithine
NADPH
NADP
21
Arginine
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
414
Metabolic charts
A. Amino acid degradation I
Iso-
1.4.4.2
Valine
b
Leucine
Glycine
leucine
CO2
2.1.2.10
a
c
OG
THF
5,10M-THF
Glu
4.1.2.5
Threo-
1 a, b, c
Serine
Glycine
2.1.2.1
nine
NAD
Branched-chain
CoA
α-ketoacid
NADH
18
dehydrogenase
NH3
complex
CO2
Histi-
2 a, b, c
17
Glu
OG
dine
[FAD]
CoQox
Alanine
[FADH2]
CoQ
2.6.1.2
red
NH3
3 a, b, c
ATP
16
15
19
a
c
H2O
b
CO2
H2O
ADP
H2O
4.2.1.18
4
10
13
14
20
H
O
Tricarboxylic
2
H2O
NAD
9
H2O
acid cycle
CoA
NADH
2.1.2.5
5
11
8
22
21
NAD
NH3
5F-THF
THF
NADH
NADH
NH3
H2O
ATP
CO2
ADP
NAD
Gluta-
Gluta-
6
12
7
6.4.1.3
mate
3.5.1.2
mine
CoA
2[H
1a
2-Oxoisovalerate
5
3-Hydroxyisobutyrate
15
Acetoacetate
1b
2-Oxo-3-methylvalerate
6
Methylmalonyl-semialdehyde
16
Acetyl CoA
1c
2-Oxoisocapronate
7
(S)-Methylmalonyl CoA
17
Pyruvate
2a
Isobutyryl CoA
8
(R)-Methylmalonyl CoA
18
Acetaldehyde
2b
2-Methylbutyryl CoA
9
Succinyl CoA
19
Urocanate
2c
Isovaleryl CoA
10
2-Methyl-3-hydroxybutyryl CoA
20
Imidazolone-
5-propionate
3a
Methylacrylyl CoA
11
2-Methylacetoacetyl CoA
3b
Tiglyl CoA
12
Propionyl CoA
21
N-Formimino-
glutamate
3c
3-Methylcrotonyl CoA
13
3-Methylglutaconyl CoA
4
3-Hydroxyisobutyryl CoA
14
3-Hydroxy-3-methylglutaryl CoA
22
2-Oxoglutarate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic charts
415
A. Amino acid degradation II
Non-enzymatic
Pyru-
reaction
2.6.1.3
1.13.11.20
Cys-
10
9
Methionine
Asparagine
vate
teine
ATP
H2O
SO2
Glu
OG
O2
β-Oxidation
P
PP
NH3
Acetyl
1
Aspartate
17
12
CoA
[CH3]
OG
Ade-
nosine
H2O
8 steps
Glu
2
16
11
Serine
8
H2O
Several
Trypto-
3
7
15
pathways
phan
H2O
Tricarboxylic
acid cycle
NH3
O2
6
Cysteine
4
21
14
Lysine
See pre-
CO2
H2O
vious page
NH3
Ascorbate
CoA
NADH
Dehydroascorbate
O2
NAD
[2 H]
5
CO2
2.6.1.5
1.14.16.1
Tyro-
Phenyl-
Glutamate
13
sine
alanine
NADH
Glu
OG
H2O
O2
NAD
THB
Non-
H2O
enzymatic
1.5.99.8
reaction
2.6.1.13
3.5.3.1
Argi-
Proline
18
19
20
nine
2[H]
H2O
Glu
OG
H2O
Urea
1
S-Adenosylmethionine
8
Oxaloacetate
15
4-Maleylacetoacetate
2
Homocysteine
9
Cysteine sulfinate
16
Fumarylacetoacetate
3
Cystathionine
10
3-Sulfinylpyruvate
17
Acetoacetate
4
2-Oxobutyrate
11
2-Oxoadipate
18
Glutamate 4-semialdehyde
5
Propionyl CoA
12
Crotonyl CoA
19
∆1-Pyrroline-5-carboxylate
6
Succinyl CoA
13
4-Hydroxyphenylpyruvate
20
Ornithine
7
Fumarate
14
Homogentisate
21
2-Oxoglutarate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
416
Metabolic charts
A. Ammonia metabolism
Nucleotide
Basic
From
Gln
Nucleobases
Toxic
protein
degradation
H2O
1
2
H2O
3.5.4.n
NH3
1.4.1.2
Glu
Glu
Ala
2.6.1.2
2
3
ATP
2 ATP
NADH
HCO3
ADP, Pi
2 ADP, Pi
NAD
Gln
4
Asp
5
2.1.3.3
6.3.4.5
8
H2O
Nitrogen donator in
ATP
synthesis of purine
bases, cytosine,
AMP, PP
GlcNAc, His, and Trp
7
9
6
Urea
cycle
3.5.3.1
4.3.2.1
10
H2O
Neutral,
non-toxic
Urea
1
Pyruvate
6
Fumarate
2
2-Oxoglutarate
7
Ornithine
3
Oxaloacetate
8
Citrulline
Kidney
4
Carbamoyl phosphate
9
Argininosuccinate
5
Malate
10
Arginine
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic charts
417
A. Biosynthesis of purine nucleotides
Ribose 5-
PRPP
phosphate
N10-Formyl-
THF
2.7.6.1
2.4.2.14
6.3.4.13
1
2
3
4
10
ATP
AMP
Gln
Glu
P P
Gly
ATP
ADP
F-THF
HCO3
ADP
ATP
ADP
Glu
ATP
Gln
THF
8
7
6
5
4.1.1.21
6.3.3.1
6.3.5.3
Asp
ATP
O
ADP
C
N
HN
Fumarate
2.1.2.3
3.5.4.10
N
N IMP
9
10
11
P
Rib
10F-THF
THF
H2O
Asp
ATP
NAD
Xanthine
1
Ribose 5-phosphate
mono-
ADP
NADH
phosphate
2
Phosphoribosyl
12
XMP
diphosphate (PRPP)
Gln
3
PR-amine
ATP
4
PR-glycineamide
ADP
5
PR-formylglycine-
Fumarate
amide
Glu
6
PR-formylglycine-
AMP
GMP
amidine
7
PR-5-aminoimidazole
NADPH
NADP
NADPH
NADP
8
PR-4-carboxy-5-
aminoimidazole
ADP
dADP
GDP
dGDP
1.17.4.1
1.17.4.1
9
PR-5-amino-4-
imidazolecarboxamide
Nucleoside
diphosphate
10
PR-5-formamido-
reductase
imidazole-4-carbox-
ATP
dATP
GTP
dGTP
amide
11
Inosine 5'-mono-
phosphate
12
Adenylosuccinic acid
PR = 5'-Phosphoribosyl-
RNA
DNA
RNA
DNA
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
418
Metabolic charts
A. Biosynthesis of the pyrimidine nucleotides and C1 metabolism
1
HCO3
P
2
Carbamoyl
Gln
ATP
ADP
Glu
Asp
H2O
FMN
FMNH2
phosphate
3
Carbamoyl
1
2
3
4
5
aspartate
6.3.5.5
2.1.3.2
3.5.2.3
1.3.99.11
4
Dihydroorotate
6
5
Orotate
DHF
5,10M-THF
P P
6
Phosphoribosyl
diphosphate
4..1.1.23
dTMP
dUMP
UMP
7
2.1.1.45
7
Orotate 5'-mono-
phosphate
P
Thymidylate
2[H
CO2
synthase
1.17.4.1
dTDP
dUDP
UDP
CDP
dCDP
1.17.4.1
ADP
ATP
Nucleoside
2[H
[NH
2
diphosphate
reductase
dTTP
UTP
CTP
dCTP
6.3.4.2
CTP
synthase
DNA
DNA
RNA
RNA
Tetrahydro-
Histidine
HCO
folate
THF
THF
ATP
N5-Formyl-
Methionine
ADP
THF
NADPH
NADP
NADPH
NADP
H
Homocysteine
H H
C
C
CH
3
THF
THF
THF
1.5.1.5
1.5.1.20
N5,N10-Methenyl-
N5,N10-Methylen-
N5-Methyl-
THF
THF
THF
H2O
dUMP
Glycine
HCO
dTMP
H2O
THF
Serine
N10-Formyl-THF
2.1.2.2, 2.1.2.3
1.5.1.3
Dihydro-
THF
DHF
folate
Purine ring (C-2, C-8)
NADP
NADPH
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Metabolic charts
419
A. Nucleotide degradation
Ade
AMP
GMP
dTMP
UMP
CMP
H2O
H2O
H2O
H2O H
H2O
2O
P
NH3
P
P
P
P
3.5.4.5
A
IMP
G
dT
U
C
H2O
P
P
P
H2O
NH3 H
2O
P
Rib
P
P Rib
P Rib
NH3
Inosine
I
Gua
Thy
Ura
P
H2O
NADPH
NADPH
Rib
P
NH3
NADP
NADP
1.1.3.22
1
2
7
11
H2O
H2O
H2O
H2O
H2O2
O2
O2
Primates,
H2O
2
birds, reptiles
Uric
8
12
acid
H2O
H2O
O2
Other mammals,
NH
NH3,
3,
molluscs
CO2
CO2
3
9
13
H2O
2-OG
2-OG
(Pyr)
Glu
Glu
Osseous fish
(Ala)
4
10
14
H2O
CO2
CO
2
2
5
Cartilaginous fish,
amphibians
6
Propionate
Acetate
3
1
Hypoxanthine (Hyp)
8
Carbamoyl-β-amino-
isobutyrate
2
Xanthine (Xan)
9
β-Aminoisobutyrate
4
3
Allantoin
10
2-Methyl-3-oxopropionate
4
Allantoic acid
11
Dihydrouracil
5
Glyoxylate
12
Carbamoyl-β-alanine
6
6
5
Urea
13
β-alanine
7
Dihydrothymine
14
3-oxopropionate
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
420
Annotated enzyme list
Annotated enzyme list
Only the enzymes mentioned in this atlas are listed here, from among the more than 2000
enzymes known. The enzyme names are based on the IUBMB’s of cial Enzyme nomenclature
1992. The additions shown in round brackets belong to the enzyme name, while prosthetic
groups and other cofactors are enclosed in square brackets. Common names of enzyme groups
are given in italics, and trivial names are shown in quotation marks.
Class 1: Oxidoreductases (catalyze reduction-oxidation reactions)
Subclass 1.n: What is the electron donor?
Sub-subclass
1.n.n: What is the electron acceptor?
1.1
A -CH-OH group is the donor
1.1.1
NAD(P)+ is the acceptor (dehydrogenases, reductases)
1.1.1.1
Alcohol dehydrogenase [Zn2+]
1.1.1.3
Homoserine dehydrogenase
1.1.1.8
Glycerol 3-phosphate dehydrogenase (NAD+)
1.1.1.21
Aldehyde reductase
1.1.1.27
Lactate dehydrogenase
1.1.1.30
3-Hydroxybutyrate dehydrogenase
1.1.1.31
3-Hydroxyisobutyrate dehydrogenase
1.1.1.34
Hydroxymethylglutaryl-CoA reductase (NADPH)
1.1.1.35
3-Hydroxyacyl-CoA dehydrogenase
1.1.1.37
Malate dehydrogenase
1.1.1.40
Malate dehydrogenase (oxaloacetate-decarboxylating, NADP+)—“malic enzyme”
1.1.1.41
Isocitrate dehydrogenase (NAD+)
1.1.1.42
Isocitrate dehydrogenase (NADP+)
1.1.1.44
Phosphogluconate dehydrogenase (decarboxylating)
1.1.1.49
Glucose 6-phosphate 1-dehydrogenase
1.1.1.51
3(or 17)β-Hydroxysteroid dehydrogenase
1.1.1.95
Phosphoglycerate dehydrogenase
1.1.1.100
3-Oxoacyl-[ACP] reductase
1.1.1.101
Acylglycerone phosphate reductase
1.1.1.105
Retinol dehydrogenase
1.1.1.145
3β-Hydroxy-∆5-steroid dehydrogenase
1.1.1.205
IMP dehydrogenase
1.1.3
Molecular oxygen is the acceptor (oxidases)
1.1.3.4
Glucose oxidase [FAD]
1.1.3.8
L-Gulonolactone oxidase
1.1.3.22
Xanthine oxidase [Fe, Mo, FAD]
1.1.99.5
Glycerol-3-phosphate dehydrogenase (FAD)
1.2
An aldehyde or keto group is the donor
1.2.1
NAD(P)+ is the acceptor (dehydrogenases)
1.2.1.3
Aldehyde dehydrogenase (NAD+)
1.2.1.11
Aspartate semialdehyde dehydrogenase
1.2.1.12
Glyceraldehyde 3-phosphate dehydrogenase
1.2.1.13
Glyceraldehyde 3-phosphate dehydrogenase (NADP+) (phosphorylating)
1.2.1.24
Succinate semialdehyde dehydrogenase
1.2.1.25
2-Oxoisovalerate dehydrogenase (acylating)
1.2.1.38
N-Acetyl-γ-glutamylphosphate reductase
1.2.1.41
Glutamylphosphate reductase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Annotated enzyme list
421
1.2.4
A disulfide is the acceptor
1.2.4.1
Pyruvate dehydrogenase (lipoamide) [TPP]
1.2.4.2
Oxoglutarate dehydrogenase (lipoamide) [TPP]
1.2.7
An Fe/S protein is the acceptor
1.2.7.2
2-Oxobutyrate synthase
1.3
A -CH-CH- group is the donor
1.3.1.10
Enoyl-[ACP] reductase (NADPH)
1.3.1.24
Biliverdin reductase
1.3.1.34
2,4-Dienoyl-CoA reductase
1.3.5.1
Succinate dehydrogenase (ubiquinone) [FAD, Fe2S2, Fe4S4], “complex II”
1.3.99.3
Acyl-CoA dehydrogenase [FAD]
1.3.99.11
Dihydroorotate dehydrogenase [FMN]
1.4
A -CH-NH2 group is the donor
1.4.1.2
Glutamate dehydrogenase
1.4.3.4
Amine oxidase [FAD], “monoamine oxidase (MAO)”
1.4.3.13
Protein lysine 6-oxidase [Cu]
1.4.4.2
Glycine dehydrogenase (decarboxylating) [PLP]
1.5
A -CH-NH group is the donor
1.5.1.2
Pyrroline-5-carboxylate reductase
1.5.1.3
Dihydrofolate reductase
1.5.1.5
Methylenetetrahydrofolate dehydrogenase (NADP+)
1.5.1.12
1-Pyrroline-5-carboxylate dehydrogenase
1.5.1.20
Methylenetetrahydrofolate reductase (NADPH) [FAD]
1.5.5.1
Electron-transferring flavoprotein (ETF) dehydrogenase [Fe4S4]
1.5.99.8
Proline dehydrogenase [FAD]
1.6
NAD(P)H is the donor
16.4.2
Glutathione reductase (NADPH) [FAD]
1.6.4.5
Thioredoxin reductase (NADPH) [FAD]
1.6.5.3
NADH dehydrogenase (ubiquinone) [FAD, Fe2S2, Fe4S4]—“complex I”
1.8
A sulfur group is the donor
1.8.1.4
Dihydrolipoamide dehydrogenase [FAD]
1.9
A heme group is the donor
1.9.3.1
Cytochrome c oxidase [heme, Cu, Zn] - “cytochrome oxidase,” “complex IV”
1.10
A diphenol is the donor
1.10.2.2
Ubiquinol cytochrome c reductase [heme, Fe2S2]—“complex III”
1.11
A peroxide is the acceptor (peroxidases)
1.11.1.6
Catalase [heme]
1.11.1.7
Peroxidase [heme]
1.11.1.9
Glutathione peroxidase [Se]
1.11.1.12
Lipid hydroperoxide glutathione peroxidase [Se]
1.13
Molecular oxygen is incorporated into the electron donor (oxygenases)
1.13.11
One donor, both O atoms are incorporated (dioxygenases)
1.13.11.5
Homogentisate 1,2-dioxygenase [Fe]
1.13.11.20
Cysteine dioxygenase [Fe]
1.13.11.27
4-Hydroxyphenylpyruvate dioxygenase [ascorbate]
1.13.11.n
Arachidonate lipoxygenases
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
422
Annotated enzyme list
1.14
Two donors, one O atom is incorporated into both (monooxygenases, hydroxylases)
1.14.11.2
Procollagen proline 4-dioxygenase [Fe, ascorbate]—“proline hydroxylase”
1.14.11.4
Procollagen lysine 5-dioxygenase [Fe, ascorbate]—“lysine hydroxylase”
1.14.13.13
Calcidiol 1-monooxygenase [heme]
1.14.15.4
Steroid 11β-monooxygenase [heme]
1.14.15.6
Cholesterol monooxygenase (side-chain-cleaving) [heme]
1.14.16.1
Phenylalanine 4-monooxygenase [Fe, tetrahydrobiopterin]
1.14.16.2
Tyrosine 3-monooxygenase [Fe, tetrahydrobiopterin]
1.14.17.1
Dopamine β-monooxygenase [Cu]
1.14.99.1
Prostaglandin H-synthase [heme]
1.14.99.3
Heme oxygenase (decyclizing) [heme]
1.14.99.5
Stearoyl-CoA desaturase [heme]
1.14.99.9
Steroid 17α-monooxygenase [heme]
1.14.99.10
Steroid 21-monooxygenase [heme]
1.15
A superoxide radical is the acceptor
1.15.1.1
Superoxide dismutase
1.17
A -CH2 group is the donor
1.17.4.1
Ribonucleoside diphosphate reductase [Fe]—“ribonucleotide reductase”
1.18
Reduced ferredoxin is the donor
1.18.1.2
Ferredoxin-NADP+ reductase [FAD]
1.18.6.1
Nitrogenase [Fe, Mo, Fe4S4]
Class 2: Transferases (catalyze the transfer of groups from one molecule to another)
Subclass
2.n: Which group is transferred?
2.1
A C1 group is transferred
2.1.1
A methyl group
2.1.1.2
Guanidinoacetate N-methyltransferase
2.1.1.6
Catechol O-methyltransferase
2.1.1.13
5-Methyltetrahydrofolate-homocysteine S-methyltransferase
2.1.1.28
Phenylethanolamine N-methyltransferase
2.1.1.45
Thymidylate synthase
2.1.1.67
Thiopurine methyltransferase
2.1.2
A formyl group
2.1.2.1
Glycine hydroxymethyltransferase [PLP]
2.1.2.2
Phosphoribosylglycinamide formyltransferase
2.1.2.3
Phosphoribosylaminoimidazolecarboxamide formyltransferase
2.1.2.5
Glutamate formiminotransferase [PLP]
2.1.2.10
Aminomethyltransferase
2.1.3
A carbamoyl group
2.1.3.2
Aspartate carbamoyltransferase [Zn2+]
2.1.3.3
Ornithine carbamoyltransferase
2.1.4
An amidino group
2.1.4.1
Glycine amidinotransferase
2.2
An aldehyde or ketone residue is transferred
2.2.1.1
Transketolase [TPP]
2.2.1.2
Transaldolase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Annotated enzyme list
423
2.3
An acyl group is transferred
2.3.1
With acyl-CoA as donor
2.3.1.1
Amino acid N-acetyltransferase
2.3.1.6
Choline O-acetyltransferase
2.3.1.12
Dihydrolipoamide acetyltransferase [lipoamide]
2.3.1.15
Glycerol 3-phosphate O-acyltransferase
2.3.1.16
Acetyl-CoA acyltransferase
2.3.1.20
Diacylglycerol O-acyltransferase
2.3.1.21
Carnitine O-palmitoyltransferase
2.3.1.22
Acylglycerol O-palmitoyltransferase
2.3.1.24
Sphingosine N-acyltransferase
2.3.1.37
5-Aminolevulinate synthase [PLP]
2.3.1.38
[ACP] S-acetyltransferase
2.3.1.39
[ACP] S-malonyltransferase
2.3.1.41
3-Oxoacyl-[ACP] synthase
2.3.1.42
Glycerone phosphate O-acyltransferase
2.3.1.43
Phosphatidylcholine-sterol acyltransferase—“lecithin-cholesterol acyltransferase
(LCAT)”
2.3.1.51
Acylgylcerol-3-phosphate O-acyltransferase
2.3.1.61
Dihydrolipoamide succinyltransferase
2.3.1.85
Fatty-acid synthase
2.3.2
An aminoacyl group is transferred
2.3.2.2
γ-glutamyltransferase
2.3.2.12
Peptidyltransferase (a ribozyme)
2.3.2.13
Protein-glutamine γ-glutamyltransferase [Ca]—“fibrin-stabilizing factor”
2.4
A glycosyl group is transferred
2.4.1
A hexose residue
2.4.1.1
Phosphorylase [PLP]—“glycogen (starch) phosphorylase”
2.4.1.11
Glycogen (starch) synthase
2.4.1.17
Glucuronosyltransferase
2.4.1.18
1,4-α-Glucan branching enzyme
2.4.1.25
4-α-Glucanotransferase
2.4.1.47
N-Acylsphingosine galactosyltransferase
2.4.1.119
Protein glycotransferase
2.4.2
A pentose residue
2.4.2.7
Adenine phosphoribosyltransferase
2.4.2.8
Hypoxanthine phosphoribosyltransferase
2.4.2.10
Orotate phosphoribosyltransferase
2.4.2.14
Amidophosphoribosyl transferase
2.5
An alkyl or aryl group is transferred
2.5.1.1
Dimethylallyltransferase
2.5.1.6
Methionine adenosyltransferase
2.5.1.10
Geranyltransferase
2.5.1.21
Farnesyl diphosphate farnesyltransferase
2.6
A nitrogen-containing group is transferred
2.6.1
An amino group (transaminases)
2.6.1.1
Aspartate transaminase [PLP]—“GOT”
2.6.1.2
Alanine transaminase [PLP]—“GPT”
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
424
Annotated enzyme list
2.6.1.3
Cysteine transaminase [PLP]
2.6.1.5
Tyrosine transaminase [PLP]
2.6.1.6
Leucine transaminase (PLP]
2.6.1.11
Acetylornithine transaminase [PLP]
2.6.1.13
Ornithine transaminase [PLP]
2.6.1.19
4-Aminobutyrate transaminase [PLP]
2.6.1.42
Branched-chain amino acid transaminase [PLP]
2.6.1.52
Phosphoserine transaminase [PLP]
2.7
A phosphorus-containing group is transferred (kinases)
2.7.1
With -CH-OH as acceptor
2.7.1.1
Hexokinase
2.7.1.3
Ketohexokinase
2.7.1.6
Galactokinase
2.7.1.11
6-Phosphofructokinase
2.7.1.19
Phosphoribulokinase
2.7.1.28
Triokinase (triosekinase)
2.7.1.30
Glycerol kinase
2.7.1.32
Choline kinase
2.7.1.36
Mevalonate kinase
2.7.1.37
Protein kinase
2.7.1.38
Phosphorylase kinase
2.7.1.39
Homoserine kinase
2.7.1.40
Pyruvate kinase
2.7.1.67
1-Phosphatidylinositol-4-kinase
2.7.1.68
1-Phosphatidylinositol 4-phosphate kinase
2.7.1.82
Ethanolamine kinase
2.7.1.99
[Pyruvate dehydrogenase] kinase
2.7.1.105
6-Phosphofructo-2-kinase
2.7.1.112
Protein tyrosine kinase
2.7.2
With -CO-OH as acceptor
2.7.2.3
Phosphoglycerate kinase
2.7.2.4
Aspartate kinase
2.7.2.8
Acetylglutamate kinase
2.7.2.11
Glutamate 5-kinase
2.7.3
With a nitrogen-containing group as acceptor
2.7.3.2
Creatine kinase
2.7.4
With a phosphate group as acceptor
2.7.4.2
Phosphomevalonate kinase
2.7.4.3
Adenylate kinase
2.7.4.4
Nucleoside phosphate kinase
2.7.4.6
Nucleoside diphosphate kinase
2.7.6
A diphosphate residue is transferred
2.7.6.1
Ribose phosphate pyrophosphokinase
2.7.7
A nucleotide is transferred
2.7.7.6
DNA-directed RNA polymerase—“RNA polymerase”
2.7.7.7
DNA-directed DNA polymerase—“DNA polymerase”
2.7.7.9
UTP-glucose-l-phosphate uridyltransferase
2.7.7.12
Hexose-1-phosphate uridyltransferase
2.7.7.14
Ethanolamine phosphate cytidyltransferase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Annotated enzyme list
425
2.7.7.15
Choline phosphate cytidyltransferase
2.7.7.41
Phosphatidate cytidyltransferase
2.7.7.49
RNA-directed DNA polymerase—“reverse transcriptase”
2.7.8
Another substituted phosphate is transferred
2.7.8.1
Ethanolaminephosphotransferase
2.7.8.2
Diacylglycerol cholinephosphotransferase
2.7.8.11
CDPdiacylglycerol-inositol 3-phosphatidyltransferase
2.7.8.16
1-Alkyl-2-acetylglycerol cholinephosphotransferase
2.7.8.17
N-Acetylglucosaminephosphotransferase
Class 3: Hydrolases (catalyze bond cleavage by hydrolysis)
Subclass
3.n: What kind of bond is hydrolyzed?
3.1
An ester bond is hydrolyzed (esterases)
3.1.1
In carboxylic acid esters
3.1.1.2
Arylesterase
3.1.1.3
Triacylglycerol lipase
3.1.1.4
Phospholipase A2
3.1.1.7
Acetylcholinesterase
3.1.1.13
Cholesterol esterase
3.1.1.17
Gluconolactonase
3.1.1.32
Phospholipase A1
3.1.1.34
Lipoprotein lipase, diacylglycerol lipase
3.1.2
In thioesters 3.1.2.4
3-Hydroxyisobutyryl-CoA hydrolase
3.1.2.14
Acyl-[ACP] hydrolase
3.1.3
In phosphoric acid monoesters (phosphatases)
3.1.3.1
Alkaline phosphatase [Zn2+]
3.1.3.2
Acid phosphatase
3.1.3.4
Phosphatidate phosphatase
3.1.3.9
Glucose 6-phosphatase
3.1.3.11
Fructose bisphosphatase
3.1.3.13
Bisphosphoglycerate phosphatase
3.1.3.16
Phosphoprotein phosphatase
3.1.3.37
Sedoheptulose bisphosphatase
3.1.3.43
[Pyruvate dehydrogenase] phosphatase
3.1.3.46
Fructose-2,6-bisphosphate 2-phosphatase
3.1.3.n
Polynucleotidases
3.1.4
In phosphoric acid diesters (phosphodiesterases)
3.1.4.1
Phosphodiesterase
3.1.4.3
Phospholipase C
3.1.4.4
Phospholipase D
3.1.4.17
3 ,5 -cNMP phosphodiesterase
3.1.4.35
3 ,5 -cGMP phosphodiesterase
3.1.4.45
N-Acetylglucosaminyl phosphodiesterase
3.1.21
In DNA
3.1.21.1
Deoxyribonuclease I
3.1.21.4
Site-specific deoxyribonuclease (type II)—“restriction endonuclease”
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
426
Annotated enzyme list
3.10.26-7
In RNA
3.1.26.4
Ribonuclease H
3.1.27.5
Pancreatic ribonuclease
3.2
A glycosidic bond is hydrolyzed (glycosidases)
3.2.1
In O-glycosides
3.2.1.1
α-Amylase
3.2.1.10
Oligo-1,6-glucosidase
3.2.1.17
Lysozyme
3.2.1.18
Neuraminidase
3.2.1.20
α-Glucosidase
3.2.1.23
β-Galactosidase
3.2.1.24
α -Mannosidase
3.2.1.26
β-Fructofuranosidase—“saccharase,” “invertase”
3.2.1.28
α,α-Trehalase
3.2.1.33
Amylo-1,6-glucosidase
3.2.1.48
Sucrose α-glucosidase
3.2.1.52
β-N-Acetylhexosaminidase
3.2.2.n
Nucleosidases
3.3
An ether bond is hydrolyzed
3.3.1.1
Adenosylhomocysteinase
3.4
A peptide bond is hydrolyzed (peptidases)
3.4.11
Aminopeptidases (N-terminal exopeptidases)
3.4.11.n
Various aminopeptidases [Zn2+]
3.4.13
Dipeptidases (act on dipeptides only)
3.4.13.n
Various dipeptidases [Zn2+]
3.4.15
Peptidyl dipeptidases (C-terminal exopeptidases, releasing dipeptides)
3.4.15.1
Peptidyl-dipeptidase A [Zn2+]—“angiotensin-converting enzyme (ACE)”
3.4.17
Carboxypeptidases (C-terminal exopeptidases)
3.4.17.1
Carboxypeptidase A [Zn2+]
3.4.17.2
Carboxypeptidase B [Zn2+]
3.4.17.8
Muramoylpentapeptide carboxypeptidase
3.4.21
Serine proteinases (endopeptidases)
3.4.21.1
Chymotrypsin
3.4.21.4
Trypsin
3.4.21.5
Thrombin
3.4.21.6
Coagulation factor Xa—“Stuart-Prower factor”
3.4.21.7
Plasmin
3.4.21.9
Enteropeptidase—“enterokinase”
3.4.21.21
Coagulation factor VIIa—“proconvertin”
3.4.21.22
Coagulation factor IXa—“Christmas factor”
3.4.21.27
Coagulation factor XIa—“plasma thromboplastin antecedent”
3.4.21.34
Plasma kallikrein
3.4.21.35
Tissue kallikrein
3.4.21.36
Elastase
3.4.21.38
Coagulation factor XIIa—“Hageman factor”
3.4.21.43
C3/C5 convertase (complement—classical pathway)
3.4.21.47
C3/C5 convertase (complement—alternative pathway)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Annotated enzyme list
427
3.4.21.68
Plasminogen activator (tissue)—“tissue plasminogen activator (t-PA)”
3.4.21.73
Plasminogen activator (urine)—“urokinase”
3.4.22
Cysteine proteinases (endopeptidases)
3.4.22.2
Papain
3.4.23
Aspartate proteinases (endopeptidases)
3.4.23.1
Pepsin A
3.4.23.2
Pepsin B
3.4.23.3
Gastricsin (pepsin C)
3.4.23.4
Chymosin
3.4.23.15
Renin
3.4.24
Metalloproteinases (endopeptidases)
3.4.24.7
Collagenase
3.4.99
Other peptidases
3.4.99.36
Signal peptidase
3.5
Another amide bond is hydrolyzed (amidases)
3.5.1.1
Asparaginase
3.5.1.2
Glutaminase
3.5.1.16
Acetylornithine deacetylase [Zn2+]
3.5.2.3
Dihydroorotase
3.5.2.7
Imidazolonepropionase
3.5.3.1
Arginase
3.5.4.6
AMP deaminase
3.5.4.9
Methylenetetrahydrofolate cyclohydrolase
3.5.4.10
IMP cyclohydrolase
3.6
An anhydride bond is hydrolyzed
3.6.1.6
Nucleoside diphosphatase
3.6.1.32
Myosin ATPase
3.6.1.34
H+-transporting ATP synthase—“ATP synthase,” “complex V”
3.6.1.35
H+-transporting ATPase
3.6.1.36
H+/K+-exchanging ATPase
3.6.1.37
Na+/K+-exchanging ATPase—“Na+/K+-ATPase”
3.6.1.38
Ca2+-transporting ATPase
3.7
A C-C bond is hydrolyzed
3.7.1.2
Fumarylacetoacetase
Class 4: Lyases (cleave or form bonds without oxidative or hydrolytic steps)
Subclass 4.n: What kind of bond is formed or cleaved?
4.1
A C-C bond is formed or cleaved
4.1.1
Carboxy-lyases (carboxylases, decarboxylases)
4.1.1.1
Pyruvate decarboxylase [TPP]
4.1.1.15
Glutamate decarboxylase [PLP]
4.1.1.21
Phosphoribosylaminoimidazole carboxylase
4.1.1.23
Orotidine-5 -phosphate decarboxylase
4.1.1.28
Aromatic L-amino acid decarboxylase [PLP]
4.1.1.32
Phosphoenolpyruvate carboxykinase (GTP)
4.1.1.39
Ribulose bisphosphate carboxylase [Cu]—“rubisco”
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
428
Annotated enzyme list
4.1.2
Acting on aldehydes or ketones
4.1.2.5
Threonine aldolase [PLP]
4.1.2.13
Fructose bisphosphate aldolase—“aldolase”
4.1.3.4
Hydroxymethylglutaryl-CoA lyase
4.1.3.5
Hydroxymethylglutaryl-CoA synthase
4.1.3.7
Citrate synthase
4.1.3.8
ATP-citrate lyase
4.1.3.18
Acetolactate synthase [TPP, flavin]
4.1.99
Other C-C lyases
4.1.99.3
Deoxyribodipyrimidine photolyase [FAD]—“photolyase”
4.2
A C-O bond is formed or cleaved
4.2.1
Hydrolyases (hydratases, dehydratases)
4.2.1.1
Carbonate dehydratase [Zn2+]—“carbonic anhydrase”
4.2.1.2
Fumarate hydratase—“fumarase”
4.2.1.3
Aconitate hydratase [Fe4S4]—“aconitase”
4.2.1.11
Phosphopyruvate hydratase—“enolase”
4.2.1.13
Serine dehydratase
4.2.1.17
Enoyl-CoA hydratase
4.2.1.18
Methylglutaconyl-CoA hydratase
4.2.1.22
Cystathionine β-synthase [PLP]
4.2.1.24
Porphobilinogen synthase
4.2.1.49
Urocanate hydratase
4.2.1.61
3-Hydroxypalmitoyl-[ACP] dehydratase
4.2.1.75
Uroporphyrinogen III synthase
4.2.99
Other C-O lyases
4.2.99.2
Threonine synthase [PLP]
4.3
A C-N bond is formed or cleaved
4.3.1
Ammonia lyases 4.3.1.3
Histidine ammonia lyase
4.3.1.8
Hydroxymethylbilane synthase
4.3.2
Amidine lyases 4.3.2.1
Argininosuccinate lyase
4.3.2.2
Adenylosuccinate lyase
4.4
A C-S bond is formed or cleaved
4.4.1.1
Cystathionine γ-lyase [PLP]
4.6
A P-O bond is formed or cleaved
4.6.1.1
Adenylate cyclase
4.6.1.2
Guanylate cyclase
Class 5: Isomerases (catalyze changes within one molecule)
Subclass 5.n: What kind of isomerization is taking place?
5.1
A racemization or epimerization (epimerases)
5.1.3.1
Ribulose phosphate 3-epimerase
5.1.3.2
UDPglucose 4-epimerase
5.1.3.4
L-Ribulose phosphate 4-epimerase
5.1.99.1
Methylmalonyl-CoA epimerase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Annotated enzyme list
429
5.2
A cis-trans isomerization
5.2.1.2
Maleylacetoacetate isomerase
5.2.1.3
Retinal isomerase
5.2.1.8
Peptidyl proline cis-trans-isomerase
5.3
An intramolecular electron transfer
5.3.1.1
Triose phosphate isomerase
5.3.1.6
Ribose 5-phosphate isomerase
5.3.1.9
Glucose 6-phosphate isomerase
5.3.3.1
Steroid ∆-isomerase
5.3.3.8
Enoyl-CoA isomerase
5.3.4.1
Protein disulfide isomerase
5.4
An intramolecular group transfer (mutases)
5.4.2.1
Phosphoglycerate mutase
5.4.2.2
Phosphoglucomutase
5.4.2.4
Bisphosphoglycerate mutase
5.4.99.2
Methylmalonyl-CoA mutase [cobamide]
5.99
Another kind of isomerization
5.99.1.2
DNA topoisomerase (type I)—“DNA helicase”
5.99.1.3
DNA topoisomerase (ATP-hydrolyzing, type II)—“DNA gyrase”
Class 6: Ligases (join two molecules with hydrolysis of an “energy-rich” bond)
Subclass
6.n: What kind of bond is formed?
6.1
A C-O bond is formed
6.1.1.n
(Amino acid)-tRNA ligases (aminoacyl-tRNA synthetases)
6.2
A C-S bond is formed
6.2.1.1
Acetate-CoA ligase
6.2.1.3
Long-chain fatty-acid-CoA ligase
6.2.1.4
Succinate-CoA ligase (GDP-forming)—“thiokinase”
6.3
A C-N bond is formed
6.3.1.2
Glutamate-NH3 ligase—“glutamine synthetase”
6.3.2.6
Phosphoribosylaminoimidazolesuccinocarboxamide synthase (sorry!)
6.3.3.1
Phosphoribosylformylglycinamidine cycloligase
6.3.3.2
5-Formyltetrahydrofolate cycloligase
6.3.4.2
CTP synthase
6.3.4.4
Adenylosuccinate synthase
6.3.4.5
Argininosuccinate synthase
6.3.4.13
Phosphoribosylamine glycine ligase
6.3.4.16
Carbamoylphosphate synthase (NH3)
6.3.5.2
GMP synthase (glutamine-hydrolyzing)
6.3.5.3
Phosphoribosylformylglycinamidine synthase
6.3.5.4
Asparagine synthase (glutamine-hydrolyzing)
6.3.5.5
Carbamoylphosphate synthase (glutamine-hydrolyzing)
6.4
A C-C bond is formed
6.4.1.1
Pyruvate carboxylase [biotin]
6.4.1.2
Acetyl-CoA carboxylase [biotin]
6.4.1.3
Propionyl-CoA carboxylase [biotin]
6.4.1.4
Methylcrotonyl-CoA carboxylase [biotin]
6.5
A P-O bond is formed
6.5.1.1
DNA ligase (ATP)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Abbreviations
431
Abbreviations
Abbreviations for amino acids, p. 60
For bases and nucleosides, p. 80
For monosaccharides, p. 38
AA
Amino acid
FAD
Flavin adenine dinucleotide
ACE
Angiotensin-converting en-
Fd
Ferredoxin
zyme (peptidyl-dipeptidase A)
FFA
Free fatty acid
ACP
Acyl carrier protein
fMet
N-formylmethionine
ACTH
Adrenocorticotropic hormone
FMN
Flavin mononucleotide
(corticotropin)
Fp
Flavoprotein (containing FMN
ADH
Antidiuretic hormone (adiure-
or FAD)
tin, vasopressin)
GABA
γ-Aminobutyric acid
ADP
Adenosine 5 -diphosphate
GDP
Guanosine 5 -diphosphate
AIDS
Acquired immunodeficiency
Glut
Glucose transporter
syndrome
GMP
Guanosine 5 -monophosphate
ALA
5-Aminolevulinic acid
GSH
Reduced glutathione
AMP
Adenosine 5 -monophosphate
GSSG
Oxidized glutathione
ANF
Atrial natriuretic factor
GTP
Guanosine 5 -triphosphate
ANP
Atrial natriuretic peptide
h
hour
(= ANF)
HAT medium Medium containing hypoxan-
ATP
Adenosine 5 -triphosphate
thine, aminopterin, and thymi-
AVP
Arginine vasopressin
dine
b
Base
Hb
Hemoglobin
bp
Base pair
HDL
High-density lipoprotein
BPG
2,3-Bisphosphoglycerate
HIV
Human immunodeficiency
cAMP
3 ,5 -Cyclic AMP
virus
CAP
Catabolite activator protein
HLA
Human leukocyte-associated
CDK
Cyclin-dependent protein
antigen
kinase (in cell cycle)
HMG-CoA
3-Hydroxy-3-methylglutaryl-
cDNA
Complementary DNA
CoA
CDP
Cytidine 5 -diphosphate
HMP
Hexose monophosphate path-
cGMP
3 ,5 -Cyclic GMP
way
CIA
Chemoluminescence immuno-
hnRNA
Heterogeneous nuclear ribo-
assay
nucleic acid
CMP
Cytidine 5 -monophosphate
HPLC
High-performance liquid chro-
CoA
Coenzyme A
matography
CoQ
Coenzyme Q (ubiquinone)
hsp
Heat-shock protein
CSF
colony-stimulating factor
IDL
Intermediate-density lipopro-
CTP
Cytidine 5 -triphosphate
tein
d
Deoxy-
IF
Intermediary filament
Da
Dalton (atomic mass unit)
IFN
Interferon
DAG
Diacylglycerol
Ig
Immunoglobulin
dd
Dideoxy-
IL
Interleukin
DH
Dehydrogenase
InsP3
Inositol 1,4,5-trisphosphate
DNA
Deoxyribonucleic acid
IPTG
Isopropylthiogalactoside
dsDNA
Double-stranded DNA
IRS
Insulin-receptor substrate
EA
Ethanolamine
kDa
Kilodalton (103 atomic mass
EIA
Enzyme-linked immunoassay
units)
ER
Endoplasmic reticulum
Km
Michaelis constant
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
432
Abbreviations
LDH
Lactate dehydrogenase
QH2
Reduced coenzyme Q (ubiqui-
LDL
Low-density lipoprotein
nol)
M
Molarity (mol L-1)
R
Gas constant
Mab
Monoclonal antibody
rER
Rough endoplasmic reticulum
MAP kinase
Mitogen-activated protein
RES
Reticuloendothelial system
kinase
RFLP
Restriction fragment length
MHC
Major histocompatability
polymorphism
complex
RIA
Radioimmunoassay
MPF
Maturation-promoting factor
RNA
Ribonucleic acid
mRNA
Messenger ribonucleic acid
ROS
Reactive oxygen species
N
Nucleotide with any base
RP
Reversed phase (of silica gel)
NAD+
Oxidized nicotinamide adenine
rRNA
Ribosomal ribonucleic acid
dinucleotide
S
Svedberg (unit of sedimenta-
NADH
Reduced nicotinamide adenine
tion coef cient)
dinucleotide
SAH
S-adenosyl L-homocysteine
NADP+
Oxidized nicotinamide adenine
SAM
S-adenosyl L-methionine
dinucleotide phosphate
SDS
Sodium dodecylsulfate
NADPH
Reduced nicotinamide adenine
sER
Smooth endoplasmic reticulum
dinucleotide phosphate
sn
Stereospecific numbering
NeuAc
N-acetylneuraminic acid
snRNA
Small nuclear ribonucleic acid
nm
Nanometer (10-9 m)
SR
Sarcoplasmic reticulum
ODH
2-Oxoglutarate dehydrogenase
ssDNA
Single-stranded DNA
PAGE
Polyacrylamide gel electropho-
TBG
Thyroxine-binding globulin
resis
THB
Tetrahydrobiopterin
Pan
Pantetheine
THF
Tetrahydrofolate
PAPS
Phosphoadenosine
TLC
Thin-layer chromatography
phosphosulfate
TPP
Thiamine diphosphate
PCR
Polymerase chain reaction
TRH
Thyrotropin-releasing hor-
PDH
Pyruvate dehydrogenase
mone (thyroliberin)
PEG
Polyethylene glycol
Tris
Tris(hydroxymethyl)aminome-
PEP
Phosphoenolpyruvate
thane
pH
pH value
tRNA
Transfer ribonucleic acid
Pi
Inorganic phosphate
TSH
Thyroid-stimulating hormone
PK
Protein kinase
(thyrotropin)
PLP
Pyridoxal phosphate
UDP
Uridine 5 -diphosphate
PP
Protein phosphatase
UMP
Uridine 5 -monophosphate
PPP
Pentose phosphate pathway
UTP
Uridine 5 -triphosphate
PQ
Plastoquinone
UV
Ultraviolet radiation
PRPP
5-Phosphoribosyl 1-diphos-
Vmax, V
Maximal velocity (of an
phate
enzyme)
PS
Photosystem
VLDL
Very-low-density lipoprotein
PTH
Parathyroid hormone
Q
Oxidized coenzyme Q (ubiqui-
none)
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Quantities and units
433
Quantities and units
1. SI base units
Quantity
SI unit
Symbol
Remarks
Length
Meter
m
1 yard (yd) = 0.9144 m
1 inch (in) = 0.0254 m
1 Å = 10-10 m = 0.1 nm
Mass
Kilogram
kg
1 pound (lb) = 0.4536 kg
Time
Second
s
Current strength
Ampere
A
Temperature
Kelvin
K
°C (degree Celsius) = K - 273.2
Fahrenheit: °C = 5/9 (°F - 32)
Light
Candela
Cd
Amount of substance
Mol
mol
2 Derived units
Quantity
Unit
Symbol
Derivation
Remarks
Frequency
Hertz
Hz
-1
S
Volume
Liter
L
10-3 m3
1 U.S. gallon (gal) = 3.785 L
Force
Newton
N
kg m s-2
Pressure
Pascal
Pa
N m-2
1 bar = 105 Pa
1 mmHg = 133.3 Pa
Energy, work, heat
Joule
J
N m
1 calorie (cal) = 4.1868 J
Power
Watt
W
J s-1
Electrical charge
Coulomb
C
A s
Voltage
Volt
V
W A-1
Concentration
Molarity
M
mol L-1
Molecular mass
Dalton
Da
1.6605
10-24
g
Molar mass
-
-
g
Molecular weight
-
Mr
-
Nondimensional
Reaction rate
v
mol s-1
Catalytic activity
Katal
kat
mol s-1
1 unit (U) = 1.67
10-8 kat
Specific activity
-
-
kat
(kg
Usually: U
(mg enzyme)-1
enzyme)-1
Sedimentation
Svedberg
S
10-13 s
coef cient
Radioactivity
Becquerel
Bq
Decays s-1
1 curie (Ci) = 3.7
1010 Bq
3 Multiples and fractions
4 Important constants
Factor Prefix Sym- Example
General gas
R = 8.314 J mol-1 K-1
bol
constant, R
23
Loschmidt
N = 6.0225
10
109
Giga G
GHz = 109
hertz
(Avogadro)
106
Mega M
MPa = 106
pascal
number, N
103
Kilo
k
kJ = 103 joule
-1
(number of
10-3
Milli
m
mM = 10-3 mol
L
particles per
10-6
Micro
µ
µV = 10-6 volt
mol)
10-9
Nano
n
nkat = 10-9 katal
-1
Faraday constant F F = 96480 C mol
10-12
Pico
p
pm = 10-12 meter
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
434
Further reading
Further reading
Textbooks
Alberts B, Bray D, Lewis J, Raff M, Roberts K, Watson JD. The molecular biology of the cell. 4th ed.
New York: Garland Science, 2002.
Berg JM, Tymoczko JL, Stryer L. Biochemistry. 5th ed. New York: Freeman, 2002.
Devlin TM, editor. Textbook of biochemistry: with clinical correlations. 5th ed. New York: Wiley-
Liss, 2002.
Granner DK, Mayes PA, Rodwell VW. Murray RK. Harper’s illustrated biochemistry. 26th ed. New
York : McGraw-Hill/Appleton and Lange, 2003.
Lodish H, Darnell J, Baltimore D. Molecular cell biology. 3rd ed. New York: Scientific American
Books, New York, 1995.
Mathews CK, van Holde KE, Ahern KG. Biochemistry. 3rd ed. San Francisco: Cummings, 2000.
Nelson DL, Cox MM. Lehninger principles of biochemistry. 3rd ed. New York: Worth, 2000.
Voet D, Voet JG. Biochemistry. 3rd ed. New York: Wiley, 2004.
Reference works
Branden C, Tooze J. Introduction to protein structure. New York: Garland, 1991.
Janeway CA, Travers P, editors. Immunobiology. 5th ed. New York: Garland, 2001.
Michal G, editor. Biochemical pathways: an atlas of biochemistry and molecular biology. New
York: Wiley, 1999.
Nature Publishing Group. Encyclopedia of life sciences. http://www.els.net [an Internet ency-
clopedia with up-to-date overview articles in every field of biochemistry and cell biology].
Webb EC, editor. Enzyme nomenclature 1992. San Diego: International Union of Biochemistry
and Molecular Biology/Academic Press, 1992.
Selected periodicals (journals and yearbooks)
Annual Review of Biochemistry. Annual Reviews, Inc., Palo Alto, CA, USA
[the most important collection of biochemical reviews].
Current Biology, Current Opinion in Cell Biology, Current Opinion in Structural Biology, and
related journals in this series. Current Biology, Ltd. London [short up-to-date reviews].
Trends in Biochemical Sciences. Elsevier Trends Journals, Cambridge, United Kingdom [the
“newspaper” for biochemists; of cial publication of the International Union of Biochemistry
and Molecular Biology (IUBMB)].
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Source credits
435
Source credits
Individual graphic elements in the following plates are based on the following sources, used
with the kind permission of the authors and publishers concerned.
Page
Figure
Source
65
Goodsell DS, Trends Biochem Sci 1993; 18: 65-8
201
A
Beckman Instruments, Munich, Bulletin no. DS-555A, p. 6, Fig. 6
203
B
Goodsell DS, Trends Biochem Sci 1991; 16: 203-6, Figs. 1a and 1b
207
C
Stryer L. Biochemistry. New York: Freeman, 1988, p. 945, Fig. 36-47
207
C
Alberts B, et al. The molecular biology of the cell. New York: Gar-
land, 1989, p. 663, Fig. 11-73B and p. 634, Fig. 11-36
279
A, B
Voet D, Voet JG. Biochemistry. New York: Wiley, 1990, p. 305,
Fig. 11-45 and p. 306, Fig. 11-47
295
A
Voet D, Voet JG. Biochemistry. New York: Wiley, 1990, p. 1097,
Fig. 34-13
297
A
Voet D, Voet JG. Biochemistry. New York: Wiley, 1990, p. 1112,
Fig. 34-33
297
A
Janeway CA, Travers P. Immunology. Heidelberg, Germany: Spek-
trum, 1994, p. 164, Fig. 4.3.c
333
B
Voet D, Voet JG. Biochemistry. New York: Wiley, 1990, p. 1126,
Fig. 34-55
335
B
Darnell J, et al. Molecular cell biology. 2nd ed. New York: Freeman,
1990, p. 923, Fig. 23-26
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
437
Index
Numbers in italics indicate figures.
A
see also individual sugars
ACP see acyl carrier protein
acetylation 62, 244
acquired immune system
A-DNA 86, 87
acetylcholine 352
294
AA see amino acids
receptors 222, 335, 354
acquired immunodeficiency
ABO blood groups 292, 293
acetylcholinesterase 354,
syndrome (AIDS) 404
absorption
355, 425
ACTase see aspartate carba-
A 102
N-acetylglucosamine
moyltransferase
light 102, 103, 128, 129
(GlcNAc)
ACTH see adrenocortico-
absorptive state 308
40, 44a
tropic hormone (cortico-
acceptors 6
phosphoglycosidase 234,
tropin)
electrons 88
235
actin 204, 205, 332
site (A) 250
phosphotransferase 234,
F-actin 64, 65, 332
ACE see angiotensin-con-
235, 425
G-actin 332
verting enzyme
acetylglutamate kinase 424
actin-associated proteins
acetaldehyde see ethanal
β-N-acetylhexosaminidase
204
acetate 13, 354
426
actinomycin D 254, 255
acetate-CoA ligase 320, 321,
N-acetylmuraminic acid
action potential 222, 348,
354, 355, 429
(MurNAc) 40
350, 351
acetic acid 49, 52, 53
N-acetylneuraminic acid
activating enzymes 114
activated see acetyl-CoA
(NeuAc) 38
activation, T-cell 296-7
acetoacetate 180
acetylornithine
activation energy (Ea) 22, 23,
acetoacetic acid 160
deacetylase 427
25, 90
acetoacetone 312
transaminase 424
active centers 74, 90
acetoacetyl-CoA 312
acetylsalicylic acid 317
enzymes 95
acetolactate synthase 428
acid amide bonds 12
essential amino acids 99
acetylcholine 270
acid anhydrides 10
lactate dehydrogenase
N-acetyl glutamate 182, 183
acid phosphatase 425
100
acetyl lipoamide 135
acid—amide bonds 13
trypsin 177
acetyl-CoA 12, 52, 53, 110,
acid—anhydride bonds 150
active transport 218, 220
163
acid—base
acyl carnitine 164, 165
acyltransferase 165, 312,
balance 274, 288-9, 289,
acyl carrier proteins (ACP)
313, 423
326-7
S-acetyltransferase 168,
carboxylase 160, 161, 162,
catalysis 90
169, 423
163, 168, 429
reactions 14, 15, 18, 19
acyl carrier proteins (ACP),
cholesterol 172, 173
acidic amino acids 60
S-malonyltransferase
ethanol metabolism 320
acidic monosaccharides 38,
168, 169, 423
fatty acid degradation
39
S-ferase 423
164, 165, 167
see also N-acetylneura-
acyl-[ACP] hydrolase 168,
pyruvate kinase inhibitor
minic acid; D-galactur-
169, 425
158
onic acid; D-glucuronic
acyl-CoA 52, 141, 164, 168,
tricarboxylic acid cycle
acid; iduronic acid
170
136, 137, 138, 139
acidoses, metabolic 160, 161,
dehydrogenase 164, 165,
van der Waals model 13
288, 326
421
N-acetyl-galactosaminyl
fat biosynthesis 170, 171
transferase 292, 293
acids 15, 18, 30-1, 31
fatty acid degradation
N-acetyl-γ-glutamylphos-
aconitate 9
162, 163
phate reductase 420
hydratase (aconitase) 8,
acylation 62, 380
acetylated amino sugars 38
9, 106, 136, 137, 428
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
438
Index
acylglycerol O-palmitoyl
aerobic oxidation 146, 147
allantoic acid 187
transferase 170, 171, 423
agarose 40, 41, 262
allantoin 186, 187
acylglycerol-3-phosphate
agents, alkylating 256, 402,
allolactose 119
O-acyltransferase 170,
403
allopurinol 186, 187
171, 423
AIDS 404
allosteric effects 2-3, 114,
acylglycerone phosphate
air 29
115, 116
reductase 420
ALA see 5-aminolevulinic
BPG 282, 284
N-acylsphingosine galacto-
acid
hemoglobin 116, 280
syltransferase 423
alanine 60, 61, 178, 184, 185
allosteric enzymes 92, 93,
adaptive immune system
β-alanine 12, 13, 62, 63,
96, 116
294
106
allosteric inhibitors 96
adaptor proteins 388
degradation 186
allosteric proteins 72
Grb-2 388
cycle 338, 339
allotypic variation 302
SOS 388
seryl 65, 66, 67
α-amanitin 242
additions 14, 15
transaminase 178, 179,
α1-globulins 276
adenine 12, 13, 80, 81
181, 423
α2-globulins 276
phosphoribosyltransfera-
albumins 194, 195, 277
α-keratin 71
se 187, 423
alcohol 10
alternative pathway,
adenosine 80, 81, 123
blood 321
complement 298
5’-diphosphate (ADP) see
dehydrogenase 64, 420
Amanita phalloides 242
ADP
fermentation 148, 149
amidases 89, 427
5’-monophosphate
liver damage 320, 321
amidine lyases 428
(AMP) see AMP
primary 11
amidophosphoribosyl
5’-triphosphate (ATP) see
secondary 11
transferase 402, 403, 423
ATP
alcohol dehydrogenase 65,
amine oxidase 62, 421
S-adenosyl-L-homocysteine
320, 321
amines 10
(SAH) 111
fermentation 148, 149
biogenic 62, 63, 352
S-adenosyl-L-methionine
alcohols 15
primary 11
(SAM) 110, 111
sugar 38, 39
secondary 11
adenosylcobalamin 108
aldehyde 11
tertiary 11
adenosylhomocysteinase
dehydrogenase 320, 321,
Aminita phalloides 204
426
420
amino acid N-acetyltrans-
adenylate
reductase 310, 311, 420
ferase 423
cyclase 120, 121, 384, 385,
aldehydes 10, 15
D-amino acid oxidase 58
386, 387, 428
aldimes 10
amino acid tRNA ligases 248
kinase 72, 73, 336, 337,
aldimine 10, 108, 178, 179
amino acids (AA) 35, 58-63,
424
aldohexoses 34, 38
146
adenylosuccinate
D-glucose see D-glucose
activation 236, 248-9, 249
lyase 428
aldolase see fructose bis-
aromatic 60, 184
synthase 429
phosphate adolase
biosynthesis 184-5, 185,
ADH see antidiuretic hor-
aldopentoses 38
412-13
mone
see also L-arabinose;
branched chain 184, 185
adhesive proteins 346
D-ribose; D-xylose
charge 58
adipocytes 46
aldoses 39
degradation 180-1, 181,
adipose tissues 160, 161, 162,
aldosterone 56, 57, 374, 376
414-15
308
aliphatic amino acids 60
essential see essential
adiuretin see antidiuretic
alkaline phosphatase 425
amino acids
hormone
alkanes 46, 47
excretion 324
ADP 190, 191, 212
alkanols 46
function 58, 59
adrenaline see epinephrine
alkenes 15
glucogenic 180
adrenocorticotropic hor-
1-alkyl-2-acetylglycerol
gluconeogenesis 154
mone (ACTH)
choline phosphotrans-
ketogenic 180
adriamycin see doxorubicin
ferase 170, 171, 425
metabolism 306, 338
aerobic glycolysis 150
alkylating agents 256, 402,
neurotransmitters 352
aerobic metabolism 122
403
optical activity 58, 59
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
439
proteinogenic 58, 60-1,
amylose 41, 42, 43
antiparallel pleated sheet 68
61
anabolic pathways 112, 113,
antiport 212, 218
non-proteinogenic 62-
138, 139, 338
antisense DNA 264
3
anaerobic conditions 112,
apatites 340, 341
resorption 272
146
APC see antigen-presenting
signaling 380, 381
anaerobic glycolysis 150,
cells
translation 60
336, 338
apoferritin 287
amino sugars 39, 44
anaerobic oxidation 146, 147
apolar molecules 28
aminoacyl
analogs
apoproteins 278
adenylate 248, 249
substrates 96, 97
apoptosis 210, 396-7, 397
tRNAs 248, 249, 252
transition states 97
aquaporin-1 220, 221
4-aminobutyrate transami-
analysis
arabinan 41
nase 356, 357, 424
analyzers 36
L-arabinose 38, 39, 42
aminodipeptidases 425
biochemical 102, 103
arachidonate lipoxygenases
aminoglycosides 254
proteins 78
390, 391, 421
β-aminoisobutyrate 186,
anaplerotic effect 180
arachidonic acid 48, 49, 390
187
anaplerotic pathways 139
arginase 183, 427
5-aminolevulinate 192, 193
anchors, lipid 214
arginine 60, 61, 182, 183
synthase 192, 193, 423
anemia 286
lactate dehydrogenase
aminomethyltransferase
sickle-cell 249, 264, 265
100
422
ANF see atrial natriuretic
vasopressin (AVP) see
aminopeptidases 176, 177,
factor
antidiuretic hormone
426
angiotensin 330, 331
argininosuccinate 182, 183
aminopropanol 63
angiotensin-converting en-
lyase 183, 428
aminopterin 304
zyme (ACE) 426
synthase 183, 429
aminotransferases 89
anhydrides
aromatic amino acids 60,
ammonia 11
mixed 11
184
bases 30, 31
phosphoric acid 11
L-amino acid decarboxy-
degradation 112
animals
lase 352, 353, 427
excretion 326, 327
cells 197
aromatic family 185
lyases 428
types of 182
aromatic ring systems 4
metabolism 178, 179, 416
anions 15, 27
arylesterase 316, 317, 425
urea cycle 182, 183
anomers 36
ascorbates 104, 368
ammonium ions (NH4+) 31,
ANP see atrial natriuretic
L-ascorbic acid see vitamin C
182, 183, 326
peptide
asialoglycoproteins 276
ammonotelic animals 182
antenna pigments 132
Asp-tRNA ligase 248, 249
AMP 158, 186, 190, 191
anthracyclines 402, 403
asparaginase 427
deaminase 336, 337, 427
anti-oncogenes 400
asparagine 60, 61
amphibolic function 136
antibiotics 225, 254-5, 255
synthase 429
amphibolic pathways 112,
antibodies 300-3, 303
aspartate 63, 178, 182, 183
113
monoclonal 304-5, 305
carbamoyltransferase
amphipathic molecules 28,
plasma cells 294
(ACTase) 116, 117, 188,
29
anticodons 82, 86, 236, 248,
422
amphiphilic molecules 28
249
family 184, 185
ampicillin 255
antidiuretic hormone (ADH)
kinase 424
amplification 398
220
kinetics 117
DNA 262
antigen-presenting cells
proteinases 177, 427
amylases
(APCs) 294
purine ring 188, 189
α-amylase 268, 269, 272,
antigens
pyrimidine ring 188, 189
273, 426
blood groups 292
semialdehyde dehydro-
brewing 148, 149
receptors 296, 297
genase 420
amylo-1,6-glucosidase 156,
antihemophilic factor A 291
transaminase 178, 179,
157, 426
antimetabolites 402, 403
183, 212, 213, 423
amylopectin 41, 42, 43
antioxidants 104, 284, 285,
aspartic acid 60, 61
amyloplasts 42
364
atomic oxygen 25
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
440
Index
ATP 35, 112, 122-3, 123
axons 348
bilirubin 194, 195, 287
ATP, charge density 123
diglucuronides 194, 195
dATP 240
monoglucuronides 194
energetic coupling 16,
B
biliverdin 194, 195
122, 124
reductase 194, 195, 421
exchange 212
B cell 161, 274, 275, 294
binding sites 250
formation 122, 123
B-DNA 84, 87
aminoacyl tRNA 252
glucose oxidation 147
bacteria
biogenic amines 12, 62, 63,
hydrolysis 122, 123
carbohydrates 35
352
nucleotide biosynthesis
cell components 202-3,
biomolecules 10-13
190, 191
203
biosynthesis
oxidative
cyanobacteria 132
heme 192-3, 193, 210
phosphorylation 123
lactic acid fermentation
NAD+ 208
phosphate transfer 123
148, 149
biotin see vitamin H
structure 122
nitrogen fixation 184, 185
biotransformations 226,
synthase see ATP syn-
polysaccharides 41
306, 316-17, 317
thase
symbiosis 184
BiP 233
synthesis 112, 122, 142-5,
bacteriophages 260, 404
1,3-bisphosphoglycerate
210
bacteriorhodopsin 126, 127
124, 125, 150, 151, 155,
NADH+H+ 142, 143
bacteroids 184, 185
282, 283
proton driven 126, 127
ball-and-stick models 6
2,3-bisphosphoglycerate see
ATP synthase 122, 126, 142,
basal transcription complex
BPG
143, 210
242, 244, 245
bisphosphoglycerate
catalytic units 142, 143
base pairs (bp) 84
mutase 283, 429
H+-transporting 140, 141,
bases (b) 30-1, 31, 80-1
phosphatase 283, 425
143, 427
ammonia 30, 31, 182
bisubstrate kinetics 95
light reactions 128, 129
chloride ions 31
blood
proton channels 142, 143
conjugated 18, 30
alcohol 321
respiratory control 144,
DNA ratios 84
cells 274, 275
145
hydroxyl ions 31
clotting 290-1, 291
ATP-citrate lyase 428
water 31
composition 274
ATP-dependent phosphory-
basic amino acids 60, see
ethanol 320, 321
lation 114
also individual amino
function 274, 275
ATPases
acids
groups 292-3, 293
Ca2+ 220
bcl-2 396
metabolites 274
F type 220
beer 148, 149
plasma 31
H+-transporting 427
Beer—Lambert law 102
serum 274
H+/K+-exchanging 427
behencic acid 49
blood—brain barrier 356
P type 220
bell-shaped curves 94, 95
boat conformation 54, 55
V type 220
benzene 4, 5
bodies, residual 234
atrial natriuretic factor
benzo(a)pyrenes 256, 257
Bohr effect 282
(ANF), receptor 388, 389
p-benzoquinone 33
bonds
atrial natriuretic peptide
β-granules 160, 161
acid—amide 13
(ANP) 388, 389
β turns 69
acid—anhydride 150
autocrine effects
β-oxidation 210
angles 7
eicosanoids 390
β-configurations 12
chemical see chemical
hormones 372, 373
β-globulins 276, 286
bonds
autotrophs 112
bicarbonate, resorption 326
disulfides 62, 72, 73, 106,
AVP see arginine vasopres-
bile 194, 268, 269
190
sin
pigments 194
double 4
axial (a) position 34
salts 314, 315
electron 6
bile acids 56, 57, 172, 312
hydrogen 72
conjugated 314
lengths 7
primary 56, 314-15
peptide 66-7
secondary 56, 314-15
phosphoric acid 122
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
441
polarity 6, 7
store 226
hemoglobin 280, 282
single 4
calcium-transporting ATP-
carbonate dehydratase 270,
ase 427
271, 283, 289, 428
bone 340, 341
calmodulin 386, 387
proton secretion 326, 327
calcium 342
calorimetry 20, 21
carbonic anhydrase see car-
remodeling 342, 343
Calvin cycle 128, 129, 130,
bonate dehydratase
Bowman’s capsule 323
131, 407
carbonium ions 15
BPG 282, 283
cAMP 120, 380, 384, 386,
carbonyl groups 11
brain 356, 357
387
carboxy-lyases 427
gluconeogenesis 154
lactose operon 119
N-carboxybiotin 108
branched-chain amino acid
phosphodiesterase 120,
γ-carboxyglutamate 62, 290
transaminase 424
121
carboxyl groups 11
brewing 149
camphor 52, 53
carboxylases see carboxy-
Brunner glands 268
CAP see catabolite activator
lyases
buffers 30, 31
protein
γ-carboxylation 62, 290
blood plasma 288, 289
cap, RNA modification 246
carboxylic acid 10, 11, 15, 48,
capacity 30, 288
capacity factor 16
49
dialysis 78, 79
capric acid 49
activated 106
function, organ metabo-
caproic acid 49
amides 10, 11
lism 308-9
capsids, rhinovirus 404
ester 10, 11
building blocks, lipids 46
N-carbamoyl aspartate 188,
carboxypeptidases 176, 177,
1,3,-butadiene 4, 5
189
425-6
butyric acid 49
carbamoyl phosphate 182,
caries 340
183, 188, 189
carnitine 164, 165
synthase 182, 183, 188,
acyltransferase 164, 165
C
429
O-palmitoyltransferase
carbohydrate metabolism
163, 165, 423
C peptide, insulin 76, 77,
160, 306, 310-11, 388
shuttle 212
160, 161
metabolic charts 408, 409
carotenoids 46, 47
C segment 302
regulation 158-9, 159
β-carotene 132, 133
C state 116
carbohydrates 34-45, 35
carrageenan 40, 41
C terminus 66
bacteria 35
carrier electrophoresis 276
c-oncogenes see proto-on-
disaccharides see disac-
caspases 396
cogenes
charides
catabolic pathways 112, 113,
C-peptides 76
hydrolysis 266
138, 139
C1
membranes 216
catabolite activator protein
metabolism 418
metabolism see carbohy-
(CAP) 64, 65, 118, 119
transferases 89
drate metabolism
catalase 24, 25, 166, 284,
caesium chloride 201
monosaccharides see
285, 421
calcidol 1-monooxygenase
monosaccharides
catalysis 24-5, 25
330, 331, 422
nutrients 360
acid—base 90
calciol see vitamin D
oligosaccharides see oli-
covalent 90
calcitonin 342, 343
gosaccharides
cycles 100, 143, 221
calcitriol 56, 57, 330, 342,
polysaccharides see poly-
enzymes 90-1
374, 376
saccharides
hydrogen peroxide 25
calcium
reserve 156
iodide 25
ATPase 220
resorption 266
lactate dehydrogenase
bone 342
see also glycolipids; gly-
100
functions 342, 343
coproteins
membranes 216
homeostasis 342, 343
carbamoyl phosphate 116,
catalysts 88, 92, see also en-
ions 328, 334, 335, 386,
182, 183, 188
zymes
387
carbon dioxide 35, 108
catalytic units, ATP synthase
metabolism 342-3
bicarbonate buffer 288
142, 143
reservoir 210
degradation 112
catechol O-methyltransfer-
sarcoplasmic pump 220
fixation 130
ase 316, 317, 422
signaling 342
transport 282-3, 283
catecholamines 352, 353
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
442
Index
cathepsins 176
galactosyl 51
55
cations 27
glucosylceramide 50, 51
cholic acid 56, 57, 314, 315
CD4 296
cGMP 386
choline 50, 51, 141, 354
CD8 296
esterase 358, 359
kinase 424
cdc2 see cyclin-dependent
phosphodiesterase 425
O-acetyltransferase 354,
protein kinase 1 (CDK1)
photoreceptors 358
355, 423
CDK see cyclin-dependent
chain termination method
phosphate cytidyltrans-
protein kinase
260
ferase 425
cDNA 260
chains
chondroitin 6-sulfate 347
CDP 190, 191
respiratory 106, 112, 126,
Christmas factor 291
choline 110, 111
140-1, 210
chromatin 208, 238, 239
diacylglycerol 170, 171
side 76, 77
chromatography 55, 78
diacylglycerol-inositol 3-
chair conformation 34, 54,
chromosomes 238
phosphatidyl transfer-
55
chylomicrons 163, 272, 278
ase 170, 171, 425
channels
chymosin 427
cell bodies 348
ion 126, 218, 222-4, 350,
chymotrypsin 94, 95, 268,
cell cycle 394-5, 395
384
269, 426
cells
K+ 222
CIA see chemoluminescence
animal 196
ligand-gated 222
immunoassay
antigen-presenting
passive 222
circadian rhythm 372
(APCs) 294
protein 218
cirrhosis 320
division 400, 401
voltage-gated Na+ 222
cis—trans isomerases 89
fractionation 198-9
chaperones 232, 233
cis—trans isomers 8, 9, 54,
inhibition 402
chaperonins 232, 233
55, 429
interaction 216
charge 59
cisplatin 402
movement 206
chemical bonds 4-7, 5
citrate 9, 137, 158
necrosis 396
chemical energy 112, 122
fat metabolism 163
organelles 196, 198
chemical potential 12, 16, 22
lyase 139
plant 196
chemical reactions 14-15
synthase 136, 137, 138,
proliferation 396, 397
chemical work 17
428
structure 196-7
chemokines 392
tricarboxylic acid cycle
transformation 401
chemoluminescent immu-
144, 145
transport 206
noassay (CIA)
citric acid 136
vacuoles 234
chemotaxis 298
cycle 141, 147
cellular immunity 294
chenodeoxycholic acid 56,
citronellal 52, 53
cellulose 34, 41, 42, 43
57, 314
citrulline 62, 63, 182, 183
hemicellulose 43
chiral center 8, 12, 35, 49
classic pathway, comple-
cement 340
chitin 40, 41
ment 298
centers
chloramphenicol 254, 255
clathrate, structure 28, 29
active 74, 90
chloride ions, bases 31
cleavage, homolytic 108
reaction 132
chlorophylls 128, 129, 132,
clones 258, 304
central nervous system see
133
closed-loop feedback con-
CNS
chloroplasts 196, 407, 407
trol 372
centrifugation 201
lumen 128, 129
clotting 290-1, 291
density gradient 200, 201
cholecalciferol see vitamin D
CMP 187
equipment 200
cholecystokinin 270
3’,5’-cNMP phosphodiester-
isopyknic 200
cholestane 54, 55
ase 425
principles 200-1
cholestanol 54, 55
CNS, signal transmission
steps 198
cholesterol 52, 53, 56, 57, 65,
348-9
zonal 200
312, 315
co-lipase 268, 269
centrosome 206
biosynthesis 172, 173
co-receptors 296
cephalin see phosphatidyl-
esterase 425
CO2 see carbon dioxide
ethanolamine
membranes 214, 215
CoA see acetyl CoA; coen-
cephalosporins 254, 255
monooxygenase 422
zyme A
cerebrosides 46, 47, 51
van der Waals models 54,
coactivator/mediator com-
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
443
plex 244
complement 298-9
R 117, 280, 281
coagulation factors 291, 426
activation 298, 299
ring 54, 55
cobalamine see vitamin B12
alternative pathway 298
space 66
cobalt 108, 109
C1—C9 298, 299
stabilizing interactions
coding strands 85
C3/C5 convertase 298,
72, 73
codogenic strands 85
299, 426
T 117, 280, 281
codons 236
C5 convertase 298, 299
zig-zag (Z) 86
anticodons 82, 86, 236,
classic pathway 298
conjugates 316
248, 249
factors 298
steroid hormones 110
start 248, 250
complementary DNA
conservation
stop 248, 250
(cDNA) see cDNA
differences 98
coenzyme A (CoA) 106, 107,
complex glycoproteins 45
energy 106, 126-7, 127
134
complex I 140
constitution 6
acetyl CoA see acetyl CoA
complex II see succinate de-
contraction
coenzyme Q (CoQ) 104, 136,
hydrogenase
muscles 332-3
140, 212
complex III 140
control 334-5
ATP synthesis 142, 143
complex IV 140
control elements 118
isoprenoid anchors 52, 53
complex V see ATP synthase,
transcription 244
oxidized 212
H+-transporting
copper ions (Cu) 132, 133,
redox systems 105
complexes
142, 143
reduced 147
complex lipids 170-1
coproporphyrinogen III 192,
cytochrome c reduc-
complex oligosaccharides
193
tase complex 141, 421
44, 230
CoQ see coenzyme Q
coenzymes 104-11, 105
complex
core oligosaccharide 230
availability 114
basal transcription 242,
core structures 44
coenzyme A see coen-
244, 245
Cori cycle 338, 339
zyme A
coactivator/mediator 244
corrin 108
coenzyme Q see coen-
daunomycin—DNA 254
corticotropin see adreno-
zyme Q
enzyme—substrate 92
corticotropic hormone
definition 104
enzymes 91, 95
cortisol 56, 57, 159, 338, 374,
energetic coupling 106,
fatty acid—albumin 163
376, 378
107
formation 72
gluconeogenesis 120, 121,
flavin see flavin
initiation, 70S 250
154
group-transferring 106-7,
lipoprotein 312
cosubstrates 104
107, 108-9, 109
membrane attack 298
cotton fibers 42
heme 106, 107
multienzyme 134
coupling
NAD see NAD
oxoglutarate dehydro-
energetic 16, 18, 106, 112
nucleotide triphosphates
genase 144, 145
ATP 16, 122, 124, 125
110, 122-3
pores 208
covalency 6, 90, 108
redox 104, 105, 106-7, 107
prothrombinase 291
creatine 324
soluble 104
repressor—allolactose 119
kinase 98, 336, 337, 424
vitamins 364
translocator 232
metabolism 336, 337
cofactors 46, 104
vitamin B2 366
phosphate 122, 336
collagenase 427
compounds 11
creatinine 336
collagens 64, 70, 71, 346
concentration
cristae 210
biosynthesis 344, 345
gradients 126, 127
CSF see colony-stimulating
bone 340
substrates 97, 116
factor
helix 65, 68, 69, 344
and velocity 93, 97
CTP 190, 191
procollagen 344
condensation, chromosomal
dCTP 190, 191, 240
structure 344, 345
238
synthase 190, 191, 429
tropocollagen 344
cones 358
cultures, primary 304
collision complexes 90
conformation 8, 9, 35
curves, bell-shaped 94, 95
colony-stimulating factors
A 86
cyanobacteria, photosystem
(CSF) 392
cis—trans 8, 9, 54, 55, 429
(PS) I 132
combustion 21
enzymes 92
cycles, catalytic 100, 143
competitive inhibitors 96, 97
native 72, 74
3’5’-cyclic AMP see cAMP
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
444
Index
3’5’-cyclic GMP see cGMP
D series monosaccharides
desulfurations 316
cyclic phosphorylation 130,
34
detoxification 306
131
DAG see diacylglycerol
development, cytokines 392
cyclin 394
dATP 240
dextrans 40, 41
B 394
daunomycin 254, 255
dGTP 240
cyclin-dependent protein
dCDP 190
DH see dehydrogenases
kinases (CDK) 394
dCTP 190, 191, 240
diabetes mellitus 76, 160-1
CDK1 394
ddNTPs 260, 261
ketone bodies 312
cyclophosphamide 402
deamination 178-80, 179,
diacylglycerol (DAG) 48, 49,
cystathionine
180, 181
170, 171, 386, 387
β-syntase 428
decarboxylases see carboxy-
O-acyltransferase 170,
γ-lyase 428
lyases
171, 423
cysteamine 12, 13, 62, 63
decarboxylation 62, 63, 108
choline phosphotransfer-
cysteine 60, 61, 63
oxidative 134
ase 425
acid—base balance 288
defense, blood 274
diacylglycerols (DAG), lipase
dioxygenase 421
deficiency
425
proteinases 177, 427
minerals 362
dialysis 78, 79
transaminase 424
vitamins 364
2,4-dienoyl-CoA reductase
cystinuria 322
genetic code 248
167, 421
Cyt P450 see cytochrome
degradation 112
differentiation, hormones
P450
amino acids 180
370
cytidine 80, 81
cyclin 394
diffusion
5’-diphosphate (CDP) see
nucleotides 186-7
facilitated 218
CDP
dehydration 108
free 218
5’-monophosphate
dehydrogenases (DH) 89,
digestion 266-73, 370
(CMP) see CMP
104, 145
digestive juices 268, 269
5’-triphosphate (CTP) see
dehydroascorbic acid 104
dihydrofolate reductase
CTP
denaturation 72
242, 304, 402, 403
cytochalasins 204
dendrites 348
dihydrogen phosphate 11
cytochrome c 140, 396
density gradient
dihydrolipoamide
oxidase 132, 133, 140, 141,
centrifugation 201
acetyltransferase 134,
421
dentine 340
135, 423
proton pumps 126, 127
2-deoxy-D-ribose 39
dehydrogenase 1, 34, 135,
cytochromes 106
deoxyaldoses 38, 39
421
b/f complex 128, 129
see also 2-deoxy-D-
succinyltransferase 423
c see cytochrome c
ribose; L-fucose
dihydroorotase 427
P450 172, 316, 318-19,
deoxycholic acid 56, 57, 314,
dihydroorotate 141, 188, 189
319
315
dehydrogenase 421
cytokines 342, 392-3, 393
deoxyhemoglobin 281
dihydrouridine 82, 83
cytoplasm 196, 202
deoxyribodipyrimidine
dihydroxyacetone
biochemical functions
photolyase 428
phosphate see glycerone-
203
deoxyribonuclease I 425
1-phosphate
nuclear interactions 208,
deoxyribonucleic acid see
3,4-dihyroxyphenylalanine
209
DNA
see dopa
pH 31
deoxyribonucleosides 80
dimers 116, 117
cytoplasmic pathway 228
triphosphates see dNTPs,
dimethylallyl
cytosine 80, 81
ddNTPs
diphosphate 173
cytoskeleton 196, 204-7, 216
2-deoxythymidine-5’-mono-
transferase 423
cytosol, fractionation 198
phosphate see dTMP
dimethylbenzimidazole 108
cytostatic drugs 402-3
dephosphorylation 114
2,4-dinitrophenol (DNP) see
cytotoxic T cells 294
depolarization 222
DNP
channels 350
dioxygenases 89
membranes 348
dipeptidases 176, 177, 425-6
D
dermatan sulfate 347
dipeptides 66
desmin 332
disaccharides 38, 39, 272,
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
445
273
dolichol 53, 53
dehydrogenase 164, 165,
disruption 198
diphosphate 230
421
dissociation curves 58
domains
electronegativity 6
dissolution 20, 21
dimers 116
electroneutral processes
disulfides 10-11
receptors 224
218
bonds 62, 72, 73, 106, 190,
SH2 392
electrons 14
300
steroid receptors 378
acceptors 88
dithiol 106
donors 6
configurations 2, 3
diuresis 328
electrons 88
donors 88
DNA 80, 84-5
dopa 62, 63, 352
excitation 128, 129
amplification 262
L-dopa 6, 7, 63
oxidized forms 32
antisense 264
dopamine 62, 63, 352
p4
B-DNA 84, 86
β-monooxygenase 352,
pairs 6
cDNA 260
353, 422
potential 18, 19
cloning 258-9, 259
double helix 84, 86
reduced forms 32
conformation 86-7, 87
double stranded (dsDNA)
reducing equivalent 33
electrophoresis 262, 263
see dsDNA
singlet state 130
fingerprinting 264, 265
doxorubicin 402
transfers 33
genetic coding 84
drugs, cytostatic 402-3
transport chain 128, 140
germ-line 302
dTMP 80, 81, 187, 191
electrophoresis 263, 277
gyrase see DNA topoiso-
dTTP 190, 191, 240
carrier 276
merase (ATP-hydro-
dUMP 190
DNA 262, 263
lyzing, type II)
dynamics, proteins 72, 73
lactate dehydrogenase
helicase see DNA topoi-
(LDH) 99
somerase (type I)
SDS gel 78, 79
E
libraries 260, 261
elementary fibrils 42, 43
ligase 256, 257, 429
elements 3
nucleus 208
e- see electrons
eliminations 14, 15
polymerases 240, 241
EA see ethanolamine
eliminative deamination
see also individual
Eadie—Hofstee plots 96,
180
polymerases
97
elongation 242
primers 240
EC number 88
Escherichia coli 251
proteins 65
ecdysone 56, 57
factors 252
sequencing 260-1, 261
ECM see extracellular matrix
elution, proteins 78, 79
structure 84, 85
EcoRI sites 258, 259
embryo, hemoglobin 280l
templates 240
effectors
emulsions 28, 270
zig-zag 86
allosteric 114, 115, 116
enamel 340
caspases 396
Dna K 232
enantiomers 8
DNA polymerases 240, 241
domain 224
enclosure, membranes 216
repair 256, 257
EIA see enzyme-linked im-
endergonic processes 16, 17,
DNA topoisomerases 240
munoassay
112
(ATP-hydrolyzing, type II)
eicosanoids 46, 47, 390-1,
endergonic transfer 18
391
240, 429
endocrine effects 372, 373
(type I) 240, 245, 429
autocrine effects 390
endocytosis 234, 235
DNA-binding proteins, on-
paracrine effects 390
receptor-mediated 278
cogenes 398
signaling 46, 390
endonucleases, restriction
DNA-dependent DNA poly-
elastase 268, 269, 426
258, 259, 425
merases 240, 242
electrical dipole 26
endopeptidases see metal-
DNA-directed DNA poly-
electrical work 17
loproteinases
merases 424
electrochemical gradients
endoplasmic reticulum (ER)
DNA-directed RNA poly-
126, 127, 128
196, 226, 227
merases 240, 242, 243,
electrolytes, recycling 328-
gluconeogenesis 154
245, 424
9, 329
protein folding 232, 233
DNP 144
electron-transferring flavo-
protein synthesis 231
dNTPs 260, 261, 262
protein (ETF) 164
rough (rER) 226, 232
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
446
Index
smooth (sER) 172, 173,
classes 88, 89
elongation 251
226, 227
see also individual
lactose operon 118
endosomes 196, 234
classes
protein biosynthesis 251
endosymbiotic theory 210
complement cascade
pyruvate dehydrogenase
endothermic reactions 20
298
134, 135
energetic coupling 16, 17, 18,
complexes 91, 95
replication 240, 241
112, 124-5, 125
conformations 92
termination 251
ATP 16, 122, 124
glucose determination
translation 251
coenzymes 106, 107
102, 103
essential amino acids (EAA)
energetics 16-17, 17
inhibitors see inhibitors
60, 174, 360
coupling see energetic
interconversion 114
active centers 99
coupling
isoenzymes 98, 99
biosynthesis 184, 185,
protein folding 74, 75
isosteric 92, 93
410, 412
energy
kinetics 92-5
non-essential 184, 413
activation 25
markers 198
essential fatty acids 46, 48
chemical 112, 122
nomenclature 420-30
ester bonds 12
conservation 106, 126-7,
pH 94, 95
esterases 89
127
proteolytic 177
esterification 36, 37
definition 16
reaction specificity 88
esters 15, 47
metabolism 336, 356
regulatory mechanisms
estradiol 56, 57, 374, 376
requirements 360, 361
114
estrogen 342
engineering, genetic 258
serum diagnosis 98, 178
receptor 72, 73
enhancers 118
substrate specificity 88
ethanal 320
enol phosphate 124
temperature 94, 95
ethanol 27, 320-1, 321
enolase see phosphopyru-
turnover 88, 89
ethanolamine (EA) 50, 51,
vate hydratase
watersplitting 130
62, 63
enoyl-[ACP] reductase 168,
see also individual en-
kinase 424
169, 421
zymes
phosphate cytidyltrans-
enoyl-CoA
epimers 36
ferase 424
hydratase 165, 166, 428
epimerases 89, 428
phosphotransferase 425
isomerase 166, 167, 429
epimerization 36, 37
ethers 10, 11
enterohepatic circulation
epinephrine 159, 163, 352,
ethidium bromide 262, 263
194, 314
380
euchromatin 208, 238
enteropeptidase (enteroki-
episodic release 372
eukaryotes 196, 197, 250,
nase) 176, 270, 426
equilibrium 16, 17, 18-19
251
enthalpy 20-1, 21
constant (K) 18, 19
event-regulated hormones
folding 74
nitrogen 174, 175
372
free 142, 143
potential 126, 350
excision repair 256
entropy 20-1, 21
equitorial (e) position 34
excitation, electrons 128,
envelope conformation 54,
ER see endoplasmic reticu-
129
55
lum
excretion
enzyme—substrate com-
ergosterol 56, 57
ammonia 326, 327
plexes 92
ERK see MAPK
hormones 370
enzyme-catalyzed reactions
erucic acid 49
kidneys 322
90, 93
erythrocytes 274
liver 306
enzyme-linked immunoas-
gluconeogenesis 154
protons 326
say (EIA) 304
metabolism 284-5, 285
exergonic processes 16, 17,
enzymes
erythromycin 254, 255
18, 112
activating 114
erythropoiesis 286
exocytosis 226, 228, 229
active centers 95
erythropoietin 330
neurotransmitters 348
activity 88, 89
erythrose-4-phosphate 153
exons 242, 243
allosteric 92, 93
Escherichia coli
exopeptidases 176, 177
analysis 102-3
aspartate carbamoyl-
exosomes 196
basics 88-9
transferase 116
exothermic reactions 20
catalysis 90-1, 216
cell components 202-3
expression plasmids 262
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
447
extensin 42, 43
Fe—Mo protein 184, 185
flavodoxin 72, 73
extracellular matrix (ECM)
Fe-protoporphyrin IX 192
flavoprotein (Fp)
346-7, 347
feedback inhibition 114, 115,
5-fluorouracil 402
158
fMet see N-formylmethione
purines 188
folate 366
F
fermentation 112, 146, 148-
folic acid 254, 255
9, 149
food 34, 266, 360
F type ATPase 220
ferredoxin 184, 185
foreign genes 264
F-actin 64, 65, 332
ferredoxin-NADP+ reduc-
forks, replication 240
Fabales 184
tase 422
formate 4, 5
facilitated diffusion 218
ferritins 286, 287
formic acid 49
factors, intrinsic 268, 269
ferredoxin (Fd) 128, 129
N-formylmethionine (fMet)
FAD see flavin adenine di-
fetal hemoglobin 280
250, 251
nucleotide
FFA see free fatty acids
5-formyltetrahydrofolate
Faraday constant 19
fibers
cycloligase 429
farnesol 52, 53
cotton 42
Fp see flavoprotein
farnesyl diphosphate 173
muscle 332, 337
fragments, Okazaki 240
farnesyltransferase 423
red muscle 336
frameshift mutation 257
Fas system 396
resorption 266
free diffusion 218
fat-soluble vitamins 46
white muscle 336
free electron pairs 6
fats 47, 48-9
fibrils 42, 344
free enthalpy (G) 16, 142,
biosynthesis 171, 409
elementary 42, 43
143
brown 144, 145
fibrin stabilizing factor 291
free fatty acids (FFA) 48
degradation 410-11
fibrin-stabilizing factor see
free radicals 32, 190, 256
digestion 270, 271
protein-glutamine γ-glu-
frucose-1,6-bisphosphate
metabolism 162-3, 163,
tamyltransferase
151
312
fibrinogen 291
β-fructofuranosidase 38,
nutrients 360
fibrinolysis 292-3, 293
426
structure 48, 49, 51
fibroin 70
fructose
synthesis, liver 162
fibronectins 346, 347
-1,6-bisphosphatase 154,
uncouplers 144, 145
filaments
155
fatty acid synthase 12, 168,
intermediate 70, 71, 204,
bisphosphate 425
169, 423
205, 206, 207
1,6-bisphosphate 150,
fat metabolism 162, 163
keratin 70, 71, 206
154, 159
fatty acid—albumin
microfilaments 204, 206,
2,6-bisphosphate 158, 159
complex 163
207
2,6-bisphosphate 2-
fatty acid-CoA ligase 163,
protofilaments 71, 204
phosphatase 425
272, 273
films
bisphosphate, adolase
fatty acids 46, 47, 48-9, 146,
amphipathic 28
310, 311, 428
312
surface 29
D-fructose 36, 37, 39
degradation 163, 164-5,
filtration
1-phosphate 38, 310
165, 166-7
cell 198
6-phosphate 150, 151,
essential 46, 48
ultrafiltration 322
153, 155, 159
liver 162
first-order reactions 22, 23
fructoses
long 166
Fischer projections 9, 35, 59
gluconeogenesis 328
muscle fibers 336
fixation, carbon dioxide 130
metabolism 310, 311
odd-numbered 166, 167
flavin 104, 140
uptake 272
resorption 272
redox systems 33
fucose 34, 44
synthesis 113, 163, 168-9,
flavin adenine dinucleotide
L-fucose 39
169
(FAD) 32, 81, 104
fucosyltransferase 292, 293
transport 164, 165
pyruvate dehydrogenase
fuel, lipids 46
unsaturated 48, 166, 167
134
fumarate 136, 137, 182, 183,
fatty liver 320
flavin mononucleotide
189
Fd see ferredoxin
(FMN) 32, 104, 105
dehydrogenase 136, 137
Fe protein 184-5
ATP synthesis 142, 143
hydratase (fumarase) 183,
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
448
Index
428
permeation chromato-
gluconolactone 37, 103
fumaric acid 8, 9
graphy 78
D-glucopyranose 34, 35
fumarylacetoacetase 427
genes
glucose 34, 35, 65, 146, 162
functional groups 10
expression 236-7
adipose tissue 160, 161
furanose 34
foreign 264
brain 356
genetic information 236,
chain 35
237
D-glucose 34, 36, 39, 44
G
libraries 260, 261
enzymic determination
multiple 302
102, 103
G proteins
probes 260
gluconeogenesis 154, 155
Ras 388
regulation 118
glycogen 156, 157
signal transduction 384,
structural 118
glycolysis 150, 151
385
therapy 264, 265, 402
insulin 121, 158, 159, 160,
G0 phase 394
genetic code 85, 248, 249
161
G1 phase 394
genetic engineering 258,
kidney 160, 161
GABA 63, 356, 357
264-5
lactose operon 118
galactokinase 310, 311, 424
genetics, molecular 236-65
liver 160, 161
galactose 44
genomes 236, 238-9
metabolism 120, 121
D-galactose 38, 39, 44
geometric isomers 8
muscle 160, 161
metabolism 310, 311
geraniol 52, 53
oxidase 102, 103, 420
uptake 272
geranyl diphosphate 173
oxidation 36, 146, 147
galactose 1-phosphate 310
geranyltransferase 423
reduction 36
galactosidase 119
germ-line DNA 302
transporter Glut-1 220,
β-galactosidase 426
Gibbs—Helmholtz equation
221
galactosylceramide 50, 51
20, 21
uptake 272
galactosyltransferase 292,
glandular hormones 372
glucose oxidation,
293
GlcNAc see N-acetyl-D-
hexokinase 147
gallbladder 195
glucosamine; N-acetyl-
glucose 1-phosphate 156,
gallstones 314
glucosamine
157, 159, 310
GalNAc see N-acetyl-D-gal-
GlcA see D-glucuronic acid
glucose 6-phosphatase 154,
actosamine
globular proteins 72-3
155, 157, 425
γ-aminobutyric acid see
globulins 276, 277
glucose 6-phosphate 37,
GABA
β-globulins 286
150, 151, 152, 157, 159
γ-carboxyglutamate 62, 290
glucagon 163, 308, 380
dehydrogenase 152, 153,
γ-globulins 276
gluconeogenesis 154, 158,
420
γ-glutamyltransferase 423
159
gluconeogenesis 154, 155
gangliosides 46, 47, 50, 51,
glucagon receptor 121
isomerase 151, 429
214, 215
1,4-α-glucan, branching
pentose phosphate
gas constant (R)
enzyme 157, 423
pathway 153
gases, transport 282-3
α-1,4-glucan, chain 157
α-glucosidase 224
gastric juice 268, 269
4-α-glucanotransferase 157,
glucoside formation 36
pH 31
423
glucosuria 160
gastricsin (pepsin C) 427
D-glucofuranose 35
glucosylceramide 50
gastrin 270
glycogen 120
glucuronate 37
gastrointestinal hormones
gluconeogenesis 35, 113,
glucuronic acid 36, 44
270
310, 311
β-glucuronidases 194
GDP 190, 191
amino acids 138, 154, 155,
glucuronosyltransferase
gels 79
180
194, 195, 316, 317, 423
electrophoresis
cortisol 120, 121
glutamate 178, 326, 357
DNA 263
glucagon 154, 158, 159
5-kinase 424
lactate dehydrogenase
insulin 160, 161
decarboxylase 356, 357,
(LDH) 99
recycling 328, 329
427
SDS 78, 79
gluconic acid 36
dehydrogenase 178, 179,
filtration 78, 79
gluconolactonase 153, 425
181, 183, 326, 327, 421
particles 78, 79
family 184, 185
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
449
formiminotransferase
α-glycerophosphate 141
GMP 186, 187, 190, 191
422
glycerophosphate shuttle
synthase 429
neurotransmitters 356
212, 213
Golgi apparatus 196, 226,
glutamate-NH3 ligase 429
glycine 60, 61, 192, 193, 314
227
glutamic acid 60, 61
amidinotransferase 336,
gonane 54, 55
glutaminase 178, 179, 181,
337, 422
GOT see aspartate transa-
327, 356, 357, 427
collagen 70
minase
glutamine 60, 61, 178, 326,
dehydrogenase 421
gout 186, 187
357
hydroxymethyltransfer-
gp130 392
brain 356
ase 422
GPT see alanine transami-
gluconeogenesis 328
glycinuria 322
nase
muscle metabolism 338
glycosaminoglycans 110
grana 128, 129
purine biosynthesis 188,
glycocalyx 214, 230
granulocytes 274, 275
189
glycocholic acid 314, 315
Grb-2 388
pyrimidine biosynthesis
glycogen 34, 35, 40, 41, 120,
GroEL 232, 233
188, 189
121
GroES 232, 233
synthase 64
balance 157
grooves, strands 86
synthetase 65, 124, 125,
liver 156
group transfer
356, 357, 429
metabolism 156-7, 157
coenzymes 106-7, 107,
glutamylphosphate reduc-
muscle 156, 336
108-9, 109
tase 420
polymer 41
potential 12, 19
glutathione 106, 284, 285
(starch) phosphorylase
reactions 18, 19
disulfide bonds 72
see phosphorylase
group X 108
oxidized
(starch) synthase 423
groups
peroxidase 284, 285, 421
synthase 120, 121, 156,
hydroxyalkyl 106
reductase 284, 285, 421
157, 158, 159
leaving 14
glyceral-3-phosphate dehy-
kinase 3 (GSK-3) 388
prosthetic 104, 134, 142
drogenase 212, 213
glycogenin 34, 35, 156
growth
glyceral 5-phosphate 153
glycolipids 35, 46, 47, 50-1
factors 342
glyceraldehyde 310
membranes 214
hormones 370
D-glyceraldehyde 34
structure 50, 51
GSH see glutathione, re-
glyceraldehyde 3-phos-
glycolysis 35, 113, 146, 150-
duced
phate 150, 151, 153, 155
9, 151, 336, 338
GSK-3 see glycogen syn-
glyceraldehyde-3-phos-
glycoproteins 35, 230
thase kinase-3
phate dehydrogenase
complex 45
GSSG see glutathione, oxi-
124, 125, 420
forms 45, 45
dized
glucose oxidation 147
mannose-rich 45
GTP 136, 190, 191
glycerophosphate shuttle
O-linked 45
dGTP 240
212, 213
glycosaminoglycans 44-5,
GTP-binding proteins 398
glycolysis 150, 151
346
guanidine
glycerol 27, 48, 49, 154, 170,
N-glycosidic bonds 80
2’-O-methylguanidine 82,
171
glycosidases 89, 426
83
gluconeogenesis 328
O-glycosidases 231
7-methylguanidine 82, 83
kinase 155, 170, 171, 424
N-glycosides 36, 37
guanidinoacetate 336
glycerol 3-phosphate 154,
N-glycosidic bonds 12, 13,
N-methyltransferase 336,
155, 170, 171
44, 123
337, 422
dehydrogenase 155, 170,
N-glycosidic links 44
guanine 80, 81, 186, 187
171, 212, 213, 420
O-glycosidic links 44
guanosine 80, 81, 186
O-acyltransferase 170,
glycosylation 62, 230
5’-diphosphate (GDP) see
171, 423
collagen 344
GDP
glycerone phosphate O-
proteins 230, 231
5’-monophosphate
acyltransferase 423
sequence 230
(GMP) see GMP
glycerone 1-phosphate 310
glycosylceramide 50, 51
glycerone 3-phosphate 151,
glycosyltransferases 89, 231
154, 170, 171
glyoxylic acid cycle 138
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
450
Index
5’-triphosphate (GTP) see
210
deacetylase 244, 245
GTP
catalase 25
histones 64, 238, 239
guanylate cyclase 358, 359,
coenzymes 106, 107, 140,
non-histone proteins 238
388, 389, 428
141
phosphorylation 238
L-gulonolactone oxidase 420
degradation 194-5, 195
HIV 404, 405
gyrase 240
groups 280
HMG-CoA 172, 173, 312
inhibitors 254
iron 286
dehydrogenase 420
monooxygenases 318
lyase 312, 313, 428
oxygenase 194, 195, 422
reductase 172, 173
H
oxygenated 287
synthase 312, 313, 428
hemiacetals 10, 11, 34
HMP see hexose mono-
H+ see hydronium ions
hemicellulose 42
phosphate pathway
(H3O+); protons
hemochromatoses 286
hnRNA 236, 242
H+-transporting ATP
hemoglobin (Hb) 64, 65,
immunoglobulins 302
synthase see ATP syn-
280-1, 281
splicing 246, 247
thase, H+-transporting
allosteric effect 116, 280
homeostasis 322, 326, 328
H+-transporting ATPase 427
buffers 288
blood 274
H+/K+-exchanging ATPase
transport 282, 283
calcium 342, 343
270, 271, 427
hemosiderin 286, 287
cytokines 392
Hageman factor 291
Henderson—Hasselbalch
hormones 370
hair 70
equation 18, 19
see also acid—base bal-
Hartnup’s disease 322
heparin 290, 347
ance
HAT medium 304
hepatic glycogen 156
homocysteine 110
Haworth projections 34, 35
hepatic metabolism 306
homogenization 198
Hb see hemoglobin
hepatocytes 163, 306, 307
homogentisate 1,2-dioxyge-
HCO-, resorption 326
heterochromatin 208, 238
nase 421
HDL 278
heterogenous nuclear ribo-
homoglycans 40
heart
nucleic acid (hnRNA) see
homoserine
absorptive state 308
hnRNA
dehydrogenase 420
muscle 332
heteroglycans 40
kinase 424
heat
heterotrophs 112, 128
honey 38
loss 20
Hevea brasiliensis 52
hops 148
of reaction 20, 21
hexokinase 147, 159, 424
hormone response ele-
heat-shock proteins (hsp)
hexose monophosphate
ments (HREs) 245, 378
232, 378
pathway (HMP) see pen-
hormone-sensitive lipase
heavy chains, IgG 300
tose phosphate pathway
162, 163
helicase 240
hexose-1-phosphate uridyl-
hormones 315, 322, 370-1,
helices
transferase 310, 311, 424
371
1-helix receptors 384
hexoses 39
action 120, 121, 378-9
7-helix receptors 225,
high-density lipoprotein
biosynthesis 370, 383
384
(HDL) see HDL
carbohydrate metabolism
α-helix 67, 68, 69, 71
high-performance liquid
158
α-keratin 70, 71
chromatography (HPLC)
closed-loop feedback
collagen 68, 70, 344
Hill coef cient (H) 116
control 372
DNA 84, 86
hippurate 324
degradation 383
double 84, 86
histamine 270, 380
differentiation 370
left-handed 86
histidine 59, 60, 61, 63, 184,
digestive processes 370
pitch 86
185
effects 370, 372, 373
superhelix 70, 71
ammonia lyase 428
episodic release 372
triple helix 70, 71, 344
dissociation curves 58
event-regulated 372
helper T cells 294
lactate dehydrogenase
excretion 370
heme
100
gastrointestinal 270
ATP synthesis 142, 143
histone
glandular 372
b 192
acetyl transferase 244,
glucose metabolism 121
biosynthesis 192-3, 193,
245
growth 370
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
451
hierarchy 372, 373
redox systems 33
drogenase 420
homeostasis 370
reducing equivalents 33
17β-hydroxysteroid dehy-
hydrophilic 120, 121, 380-
sulfide 11
drogenase 420
9
hydrolases 88, 89, 234, 425
5-hydroxytryptophan 63
inactivation 383
hydrolysis 266, 267
hydroxyurea 402
ion concentrations 370
G of 18
hyperbilirubinemias 194
juvenile 52, 53
hydrolytic cleavage 316
hyperbolic plots 93, 97
lipophilic 120, 374-5, 375,
hydrolytic deamination 179,
hyperglycemia 160, 161
379
180
hyperlipidemia 160, 161
lipophilic see lipophilic
hydrolyzable lipids 46
hyperpolarization 348
hormones
non-hydrolyzable 46
hyperuricemia 186, 187
metabolism 120-1, 370
hydronium ions (H3O+) 30,
hypervitaminosis 364
neurohormones 348, 349
31
hypoiodide 25
peptide 380, 381, 382-3
hydroperoxyl radical 33
hypovitaminosis 364
plasma levels 372-3, 373
hydrophilic hormones 120,
hypoxanthine 186, 187, 304
proteohormones 380, 381
121, 380-9, 385
phosphoribosyltransfer-
proteolysis 382
hydrophilic molecules 26, 28
ase 186, 187, 402, 403,
pulsatile release 372
hydrophilic signaling sub-
423
receptors 120
stances 392
regulation 120, 370, 371
hydrophobic effects 72
I
renal 330-1, 331
hydrophobic molecules 26,
second messengers 386-7
28
signal cascades 388-9
hydroquinone 33, 104
ice 26, 27
steroid see steroid hor-
hydroxy radicals 284
idiotypic variation 302
mones
3-hydroxy-3-methylglu-
IDL 278
tissue 370
taryl-CoA see HMG-CoA
iduronic acid 44
transport 370
3β-hydroxy-∆5-steroid de-
IF see intermediary fila-
vitamins 364
hydrogenase 420
ments
HPLC see high performance
β-hydroxyacyl-CoA 164
IFN see interferons
liquid chromatography
3-hydroxyacyl-CoA
Ig see immunoglobulins
HRE see hormone response
dehydrogenase 165, 420
IL see interleukins
elements
hydroxyalkyl groups 106
imidazolonepropionase 427
hsp see heat-shock proteins
hydroxyalkyl thiamine
imino acids 179
human
diphosphate 135
immune response 294-5,
body 31
3-hydroxybutyrate 141, 312
295
immunodeficiency virus
dehydrogenase 420
immune system
(HIV) see HIV
β-hydroxybutyric acid 160
acquired 294
humoral immunity 294
3-hydroxyisobutyrate dehy-
cytokines 392
hyaluronate 346, 347
drogenase 312, 313, 420
inate 294
hyaluronic acid 41, 44, 45
3-hydroxyisobutyryl-CoA
immunoassays 304, 305
hybridomas 304
hydrolase 425
immunoglobulins (Ig)
hydration 26, 27, 44
hydroxyl ions (OH-) 30, 31
classes 301
domains 300
shell 90
3-hydroxylacyl-CoA 141
hydride ions 32, 33, 104
hydroxylation 62, 70
heavy chains 302
hydrocarbons 46, 47
collagen 344
IgA 300
hydrochloric acid 266, 270,
hydroxymethylbilane syn-
IgD 300
271
thase 192, 193, 428
IgE 300
hydrogen
hydroxymethylglutaryl-CoA
IgG 44, 45, 64, 65
atoms 33
see HMG-CoA
domains 300, 301
bonds 6, 7, 26, 68, 72
3-hydroxypalmitoyl-[ACP]
IgM 296, 300
carbonate 182
dehydratase 168,169, 428
light chains 302, 303
chloride 30, 31
4-hydroxyphenylpyruvate
superfamily 296
ions 288, 289
dioxygenase 421
variability 302, 303
peroxide 24, 186, 284
hydroxyproline, collagen 70
IMP 186, 187
catalysis 25
3β-hydroxysteroid dehy-
biosynthesis 188, 189,
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
452
Index
190, 191
proinsulin 76, 160, 161
inorganic 202
cyclohydrolase 427
receptor see insulin re-
products 30
dehydrogenase 420
ceptor
IPTG see isopropylthioga-
innate immune system 294
signal transduction 388,
lactoside
inducers 118
389
iron
catabolite activator pro-
insulin receptor 121, 224,
deficiency 286
tein (CAP) 118
225, 388
distribution 287
induction, transcription 114,
substrate (IRS) 388
ion, hemoglobin 280
262
intensity factor 16
metabolism 286-7, 287
influenza virus 404
intercalators 225, 254
protein 184-5
inhibition, feedback 114, 115
daunomycin—DNA com-
uptake 286
inhibitors 96-7, 97
plex 254
iron—sulfur centers 140, 141,
allosteric 96
interconversion 134
142, 143
competitive 96, 97
enzymes 114, 115
iron—sulfur clusters 106,
gyrase 254
processes 144, 394
107, 286, 287
irreversible 96
pyruvate dehydrogenase
iron-protoporphyrin IX 192
kinetics 96, 97
134
IRS see insulin-receptor
reversible 96
regulation 118, 120
substrate
translation 254
interferons (IFN) 392
islets of Langerhans 160
initiation
interleukins (IL) 392
isoalloxazine see flavin
complexes, 70 S 250, 251
IL-6 receptor 392
(2R,3S)-isocitrate 9, 127
factors 250
intermediary metabolism
isocitrate 136, 141
transcription 242, 244,
112, 322
dehydrogenase 136, 137,
245
intermediate filaments (IF)
420
inorganic components
204, 205, 206, 207
glucose oxidation 147
food resorption 266
α-keratin 70, 71
tricarboxylic cycle 144,
urine 324
intermediate-density
145
inorganic ions 2, 202
lipoprotein see IDL
isoelectric point 58, 59
inorganic phosphate (Pi)
intermediates 142, 143
isoenzymes 99
125, 150
intermembrane space 210
pattern 98
inosine 5’-monophosphate
international units (U) 88
isolation, proteins 78
(IMP) see IMP
interphase 394
isoleucine 60, 61
inositol 1, 4, 5-trisphosphate
intestines, pH 31
isomerases 88, 89, 428
(InsP3) 170, 171, 386, 387
intramolecular transferases
cis—trans-isomerases 89
insulation
89
coenzymes 108
lipids 46, 47
intrinsic factor 268, 269
isomerization, light-in-
membranes 216
introns 242, 243
duced 358
insulin 64, 70, 76, 77, 160,
inulin 40, 41
isomers
161, 380
invertase see β-fructofura-
geometric 8
A chain 76, 77
nosidase
isomerism 8-9
B chain 76, 77
iodide
isopentenyl diphosphate 52,
biosynthesis 151, 160-1
catalysis 25
53, 172, 173
C peptide 76, 77, 160, 161
hypoiodide 25
isopentenyl-AMP 53
circuit 373
iodothyronines 374
isoprenoids 52-3, 53, 172
deficiency 160, 161
ion channels 65, 126, 218,
isoprenylation 52
gluconeogenesis 160, 161
222-4, 350, 384
isopropylthiogalactoside
glucose 121, 158, 159, 160,
see also individual recep-
(IPTG)
161
tors
isosteric enzymes 92, 93
human 76
ionic strength 78
isotypes 302
monomer 76
ionizing radiation 256
isopyknic centrifugation
overexpression 76
ionotropic receptors 348,
200
pentose phosphate path-
354
way 152
ions
postabsorptive state 308,
calcium 328, 386
309
hormone concentrations
preproinsulin 160, 161
370
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
453
J
L
libraries
DNA 260, 261
genes 260, 261
J segments 302
L segments 302
Lieberkühn glands 268
Janus kinases (JAK) 392
lac-repressor 118, 119
jaundice 194
lactacidosis 288
life cycle, HIV 404
joules 16, 17
β-lactam antibiotics 254
ligand-gated channels 222
lactate 98, 99, 154, 155, 288
nicotinic acetylcholine
juvenile hormone 52, 53
receptor 222
dehydrogenase see lac-
ligands 118, 119
tate dehydrogenase
K
gluconeogenesis 328
modulation 114, 115
muscle 338
oncogenes 398
K+ channels 223
redox reactions 19
ligases 88, 89, 106, 429
K systems 116
lactate dehydrogenase
light
katal 88
(LDH) 99, 420
absorption 102, 103, 128,
keratan sulfate 247
activity 102, 103
129
keratin
catalytic cycle 100, 101
monochromatic 102
filaments 206
Cori cycle 338, 339
ultraviolet 256
α-keratin 70, 71
fermentation 148, 149
light chains
kernicterus 194
gel electrophoresis 99
biosynthesis 302
ketamine 179
gluconeogenesis 155
IgG 300
ketoacidosis 288
glucose oxidation 147
light-induced isomerization
ketoacidotic coma 312
isoenzymes 98
358
β-ketoacyl-CoA 164
structure 98
lignoceric acid 49
ketogenic amino acids 180
lactic acid 8, 9, 149
Lineweaver—Burk plots 92,
α-ketoglutarate see 2-oxo-
Lactobacillus 148, 149
93, 96
glutarate
lactoferrin 286, 287
linoleic acid 48, 49
ketohexokinase 310, 311,
lactose 38, 39
linolenic acid 48, 49
424
allolactose 119
lipase 268, 269
ketohexoses 38
lactose operon 118
lipid 46-57
D-fructose see D-fructose
operon 118, 119
alcohols 46
ketone bodies 160, 161, 162,
laminin 346
anchors 214
163, 172
lanosterol 173
classification 47
acid—base balance 288
lattices
complex 170-1
biosynthesis 312, 313
hexagonal 27
hydrolysis 266
brain 356
ice 26
hydroperoxide gluta-
synthesis 410
lauric acid 49
thione peroxidase 421
ketonemia 312
LDH see lactate dehydro-
lipoproteins 278
ketones 10, 11, 162
genase
membranes 214, 215, 216
ketonuria 160, 312
LDL 278
metabolism 160, 306,
ketopentoses 38
receptor 278
312-13, 313
D-ribulose see D-ribulose
leaving groups 14
resorption 272, 273
ketoses 39
lecithin see phosphatidyl-
role 46, 47
kidney, glucose 160, 161
choline
synthesis 226
kidneys 154, 288, 322-3, 323
lecithin-cholesterol acyl-
lipid-soluble vitamins 46,
kinetics
transferase (LCAT) see
364-5, 365
allosteric enzymes 116
phosphatidylcholine-ste-
lipoamide 106, 107, 134
aspartate 117
rol acyltransferase
arm 134, 135
bisubstrate 94, 95
lefthanded α-helix (aL) 68
lipogenesis 162
enzymes 92-3
Lesch—Nyhan syndrome
lipoic acid 62, 106
inhibition 96, 97
186
lipolysis 162
see also reaction kinetics
leucine 60, 61
lipophilic hormones 120,
374-5, 375, 378, 379
Km see Michaelis constant
transaminase 424
knall-gas 20
leukocytes 274
receptors 378, 398
reaction see oxyhydrogen
leukotrienes 390
lipoprotein 278-9, 279
reaction
liberins 372
lipase 162, 163, 278, 279,
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
454
Index
425
bodies
membrane receptors 224-5,
lithocholic acid 56, 57, 314
macroelements 2, 363
398
liver
macromolecules 202
membranes
absorptive state 308
macrophages 274, 275
attack complex 298
alcohol 320
major histocompatibility
carbohydrates 216
cirrhosis 320
complex see MHC
cell interaction 216
damage, alcohol 321
malate 137, 141
composition 216
liver, fat synthesis 162
dehydrogenase 136, 137,
cytoskeletal anchors 216
fatty 320
138, 139, 420
depolarization 348
fatty acids 162
gluconeogenesis 154,
electrochemical gra-
functions 306-7, 307
155
dients 126
glucose 160, 161
glucose oxidation 147
energy conservation 126-
glycogen 156
urea cycle 183
7
metabolism see liver me-
shuttle 212, 213
functions 216-17
tabolism
maleic acid 8, 9
hyperpolarization 348
postabsorptive state 308
maleylacetoacetate isomer-
lipids 214, 216, 409
postresorption 309
ase 429
micelles 28
resorption 309
malic enzyme 139
mitochondrial 210
urea 182
malonyl-CoA 162, 163
nuclear 208
liver metabolism 306, 307
malt 148
permeability 218, 219
ammonia 178, 179
maltose 38, 39
plasma 196, 214-15
glucose 120, 121
brewing 148, 149
potential 126, 127, 350
long-chain fatty-acid-CoA
D-mannitol 39
proteins 214, 216
ligase 429
mannose 44
proton gradients 126, 127
loops
D-mannose 36, 37, 38, 39,
receptors see membrane
chromosomes 238
44
receptors
mobile 99
mannose 6-phosphate 234
semipermeable 78
peptides 98
mannose-rich glycoproteins
transport 216-19
low-density lipoprotein see
45
uncouplers 144, 145
LDL
mannose-rich oligosacchar-
vesicles 28, 198
lumen, chloroplasts 128, 129
ides 44, 230
menaquinone 52, 53
lyases 88, 89, 427-8
α-mannosidase 426
menthol 52, 53
lymphocytes 161, 274, 275,
MAPK (MAP kinase) 388
6-mercaptopurine 402
294
markers
messenger RNA see mRNA
monoclonal antibodies
enzymes 198
messengers, second see sec-
304
molecules 198, 199
ond messengers
lymphokines 392
tumors 400
metabolic acidoses 160, 161,
Lynen cycle 312
mass action, law of 18, 19
288, 326
lysine 60, 61, 184
matrix
metabolic charts 406-10
hydroxylase see procolla-
extracellular 346-7
metabolic defects, gene
gen lysine 5-dioxyge-
space 210
therapy 265
nase
maturation, RNA 208, 236,
metabolic pathways 88
lysines, acetylation 238
242, 243
hormones 370
lysophosphatidate 170, 171
maturation-promoting fac-
metabolism 356-7
lysophospholipid 50, 51
tor (MPF)
acetylcholine 354, 355
lysosomes 196, 234-5, 235
maximum velocity (Vmax)
antimetabolites 402, 403
pH 31
92, 93
buffer function 308-9
lysozyme 42
mechanical damage 144
calcium 342-3
mechanical work 17
carbohydrate 388
mechanisms
creatine 336
M
ping-pong 90, 91, 94
energy, brain 356
regulatory 114-21
glycogen 157
M phase 394
sequential 90, 91, 94, 95
hormones 120-1, 370,
M13 phage 260, 404
mediators 224, 370, 388
376-7
mAb see monoclonal anti-
MEK 388
intermediary 112,113, 322
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
455
liver 306
93, 96
resorption 272, 273
muscle 336-7
Michaelis-Menten
motor end plate 335
regulation 115, 120-1
equation 92, 93
MPF see maturation-pro-
steroids 376-7
kinetics 92, 93
moting factor; mitosis-
metabolites 96, 158
model 96
promoting factor
activated 110-11, 111
microelements 362
mRNA 82, 236, 246
liver 306
microfibrils 42, 43
3’ modification 246, 247
metabotropic receptors 348,
microfilaments 204, 206,
5’ modification 246, 247
354
207
immunoglobulins 302
metalloproteinases 176, 177,
microtubules 204, 206, 207
POMC 382
427
milk products, fermentation
MstII 264, 265
metals
148, 149
mucus 268
complexes 33
minerals 360, 362-3, 363
multienzyme complexes
heavy 316
minibands, chromosomes
134
nonmetals 2
238
muramoylpentapeptide car-
methane 26, 27, 28, 29
mitochondria 196
boxypeptidase 254, 426
methemoglobin reductase
fatty acid degradation 163
murein 34, 35
284, 285
fractionation 198
polymers 41
methionine 60, 61, 110, 184,
gluconeogenesis 154
MurNAc see N-acetylmura-
250, 251
membrane 210
minic acid
acid—base balance 288
metabolic functions 210-
muscle
adenosyltransferase 423
11
absorptive state 308
methotrexate 402
protein import 232, 233
amino acid metabolism
methylation 110, 316
structure 210-11, 211
338, 339
methylcobalamin 108
transport 212-13, 213
calcium ions 335
methylcrotonyl-CoA car-
mitogen-activated protein
contraction see muscle
boxylase 429
kinase see MAPK
contraction
methylenetetrahydrofolate
mitosis-promoting factor
fibers see muscle fibers
dehydrogenase 421
(MPF) 394
glucose 160, 161
reducatase 421
mobile loops 99
glycogen 156, 336
α-methylglucoside 37
mobile phase 54
heart 332
methylglutaconyl-CoA hy-
modulation, ligands 114
metabolism see muscle
dratase 428
molecular genetics 236-65
metabolism
7-methylguanidine 82, 83
molecular models 76-7
motor end plate 335
2’-O-methylguanidine 82,
p-molecular orbitals 4
postabsorptive state 308
83
molecular oxygen 186
protein 338, 339
methylmalonyl-CoA 166,
molecular structure 6
sarcoplasmatic reticulum
167
molecules 7, 198, 202, 218
(SR) 335
epimerase 428
monoacylglycerols 48, 49,
skeletal 332
mutase 167, 429
170, 171
smooth 332, 338
methylnitrosamines 256,
2-monoacylglycerols 272
striated 333
257
monoamine oxidase (MAO)
muscle contraction 332-3,
O-methylnorepinephrine
see amine oxidase
333, 334-5
317
monochromatic light 102
muscle fibers 332, 336, 337
methyltetrahydrofolate cy-
monoclonal antibodies
muscle metabolism 336-9,
clohydrolase 427
(mAb) 304-5, 305
339
5-methyltetrahydrofolate-
monocytes 274, 275
mutarotation 36, 37
homocysteine S-methyl-
monokines 392
mutases 429
transferase 422
monomers, insulin 76
mutation 256-7
methyltransferases 108
monooxygenases 89, 104,
frameshift 256, 257
mevalonate 172, 173
106, 166, 318, 319
mutagens 256, 257
kinase 424
cytochrome P450-de-
point 248, 256, 257
mevalonyl diphosphate 173
pendent 318, 319
repair 256, 257
MHC proteins 224, 296
monosaccharides 34-5, 35
somatic 302
micelles 28, 29, 270
acidic 38, 39
myelomas 304
Michaelis constant (Km) 92,
reactions 36-9, 37, 39
myo-inositol 50, 51
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
456
Index
myofibrils 332
urea 182
419
myoglobin 72, 73, 282, 336
niacin 366
dinucleotides 80
myosin 64, 65, 332
nicotinamide 366
nucleotide triphosphates
ATPase 427
adenine dinucleotide
110, 122
tropomyosin 64, 65, 332,
oxidized see NAD+
see also individual nu-
334
reduced see NADH
cleotides
myristic acid 49
adenine dinucleotide
nucleus 196, 208-9, 209
phosphate
and cytoplasm 208, 209
fractionation 198
N
oxidized see NADP+
localization sequence 208
reduced see NADPH
membrane 208
N terminus 66
nicotinate 366
pores 208, 209
N-bound oligosaccharides
nicotinic acetylcholine
scaffolding 238
230
receptor 223
tumor suppressors 398
Na+/K+-ATPase 126, 127, 272,
nitric oxide synthase 388,
nutrients 112, 306, 360, 361
273, 326, 327, 350, 351,
389
427
nitrogen 11
O
NAD+ 99, 101, 104, 366
balance 174
/NADH ratio 144
fixation 184, 185
biosynthesis 208
monoxide (NO) 388, 389
O2 see oxygen
gluconeogenesis 155
nitrogenase 184, 185, 422
octet rule 2
redox reactions 19
norepinephrine 145, 163,
ODH see 2-oxoglutarate de-
NADH 101
317, 352
hydrogenase
dehydrogenase 141, 421
nuclear localization se-
OH- see hydroxyl ions (OH-)
glucose oxidation 147
quence 208
oil
NADH+H+
nuclear pores 208, 209
drop effect 28, 29
ATP synthesis 142, 143
nuclear tumor suppressors
emulsions 28
gluconeogenesis 155
398
Okazaki fragments 240
NADP+ 33, 104, 105, 366
nucleic acids
oleic acid 49
photosynthesis 128, 129,
bases 80, 81
oligo-1,6-glucosidase 426
131
genetic information 236
oligonucleotides 80, 81
NADPH+H+ 33, 152, 190, 191,
hydrolysis 266
oligosaccharides 44, 45, 230,
284
see also individual nucleic
272, 273
photosynthesis 128, 129
acids
OMP 188, 189
native conformation 72
nucleobases
oncogenes 398-9, 399, 400
nephron 322
components 188, 189
one-helix receptors 224
Nernst equation 126, 127
see also purine ring;
operators 118
nerve cells 348, 349
pyrimidine ring
opsin 358
nervonic acid 49
nucleolus 208
opsonization 298
net charge, amino acids 59
nucleophilic substitutions
optical activity 8
net outcome 136
14, 15
optical isomers 8, 9
NeuAc see N-acetylneura-
nucleosidases 426
optical rotation 36
minic acid
nucleoside 80, 81
orbitals 2, 4, 5
neural synapses 349
diphosphatase 427
organ metabolism 308-9
neural tissue 308
diphosphate kinase 190,
organelles 196, 198, 199
neuraminidases 276, 426
191, 424
organic compounds 2
neurohormones 348, 349
phosphate 106, 107
organic substances 360-1
neuropeptides, proteolysis
kinase 190, 191, 424
organisms
382
nucleoside diphosphate re-
heterotrophic 128
neurotransmitters 62, 63,
ductase 190, 191
phototrophic 128
348, 349, 352-3, 353
nucleosomes 65, 238
orientation, substrates 90,
GABA 62, 63, 356
nucleotides 80-1, 81
100
glutamate 356
biosynthesis 190-1, 191,
origin of replication 240
receptors 354-5, 355
417-18
ornithine 62, 63, 182, 183
neutral amino acids 60
degradation 186-7, 187,
carbamoyltransferase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
457
183, 422
photosynthesis 128, 129
genase
orotate phosphoribosyl-
redox systems 33
PDK-1 388
transferase 423
oxygen transport 282-3
pectins 42, 43
orotidine 5’-
hemoglobin 280, 282
PEG see polyethylene glycol
monophosphate see OMP
saturation curve 280, 282
penicillins 40, 254, 255
orotidine-5’-phosphate de-
2-oxyglutarate 137, 326
pentose phosphate pathway
carboxylase 427
dehydrogenase 134, 135
(PPP) 113, 152-3, 153
osteoblasts 342
complex 136, 137
and amino acid
osteoclasts 342
oxyhydrogen reaction 20, 21
biosynthesis 185
overexpression, insulin 76
pentoses 39
oxaloacetate 137, 155
PEP 124, 125, 150, 154, 155
oxaloacetic acid 136, 154
P
carboxykinase 139, 154,
oxidases 89, 104
155
oxidation 10, 11, 316
P type ATPase 220
carboxylase 158, 159
α-oxidation 166
P/O quotient 146
PEP-CK 242, 427
β-oxidation 113, 141, 210
p53 protein 394, 396
gene 243
oxidation, β-oxidation, fat
PAGE see polyacrylamide gel
transcription 245
metabolism 162, 163
electrophoresis
pepsin 94, 176, 268, 269,
fatty acid degradation
palmitate 162, 163, 169
270
164, 165, 166, 167
palmitic acid 49, 168
A 427
glucose 36, 37
Pan see pantetheine
B 427
ω-oxidation 166
pancreas 268, 269, 308
pepsinogen 268
terminal 136
pancreatic ribonuclease 426
peptidases 89, 176, 330, 331,
oxidative deamination 179-
pantetheine 106, 169
426
80
pantoic acid 12, 13, 106
peptide hormones 380, 381,
oxidative decarboxylation
pantothenate 12
382-3
134
pantothenic acid 106, 366
biosynthesis 382
oxidative phosphorylation
papain 300, 427
degradation 382
112, 140, 210
PAPS see phosphoadenosine
hormone-like 392-3
ATP 122, 123
phosphosulfate
inactivation 382
oxidized forms, electrons 32
paracrine effects 372, 373,
peptides
oxidizing agents 32
390
bonds 66-7, 67, 252
oxidoreductases 88, 89
parallel pleated sheet 68
C-peptides 76
oxoacid 179
parathyroid hormones
chains 66
oxoacid dehydrogenases
(PTH) 342
conformation 66, 67
134-5
Parkinson’s disease 62
hormones see peptide
3-oxoacid-CoA transferase
passive channels 222
hormones
180
passive transport 218
loops 98
3-oxoacyl-[ACP] reductase
pathways
neurotransmitters 352
168, 169, 420
amphibolic 112, 113
nomenclature 66, 67
3-oxoacyl-[ACP] synthase
anabolic 112, 113, 138,
sequence 66
168, 169, 423
139, 338
signal 228, 230
2-oxobutyrate synthase 421
anaplerotic 139
peptidoglycan 35
2-oxoglutarate 136, 178
catabolic 112, 113, 138,
peptidyl dipeptidases 330,
dehydrogenase (ODH)
139
331, 425
144, 145, 147, 421
cytoplasmic 228
peptidyl proline cis—trans-
glucose oxidation 147
pentose phosphate (PPP)
isomerase 233, 429
2-oxoisovalerate dehydro-
113, 152-3, 153
peptidyl site (P) 250, 252
genase 420
secretory 228
peptidyl-dipeptidase A see
oxygen 11
PCR 262-3, 263
angiotensin-converting
atomic 25
deoxyribonucleoside tri-
enzyme
phosphates 262
peptidyltransferase 423
primers 262
perinuclear space 208
RT-PCR 264, 265
periodic table 2, 3
PDH see pyruvate dehydro-
peroxisomes 196
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
458
Index
periplasm 35
phosulfate (PAPS) 110, 111
cine ligase 429
permeability, membranes
phosphodiesterase 120, 121,
phosphoribosylaminoimi-
218, 219
386, 387, 425
dazole
permease 119
phosphoenolpyruvate see
carboxamide formyl-
peroxidases 89, 102, 103,
PEP
transferase 422
106, 305, 421
phosphoenolpyruvate,
carboxylase 427
peroxide anions 284
carboxykinase see PEP-CK
succinocarboxamide
pH 18
6-phosphofructo-2-kinase
synthase 429
cells 30, 31
424
phosphoribosylformylglyci-
enzymes 94, 95
6-phosphofructokinase 158,
namidine synthase 429
fermentation 148
159
phosphoribosylglycinamide
pH, gradient 127
glucose oxidation 147
formyltransferase 422
membrane difference 126
glycolysis 150, 151
phosphoribulokinase 131
neutral 30
phosphoglucomutase 429
phosphoric acid, esters 10,
phages see bacteriophages
phosphogluconate
11, 80
phagocytosis 234, 235
6-phosphogluconate 152,
phosphoric acid—anhydride
phalloidin 204
153
11, 80
phenobarbital 194
dehydrogenase 153, 420
bonds 12, 13, 122, 123
phenylalanine 60, 61
6-phosphogluconolactone
phosphoric acid—ester,
4-monooxygenase 422
152, 153
bonds 13, 122, 123
phenylethanolamine N-
phosphoglycerate
phosphorus 11
methyltransferase 352,
2-phosphoglycerate 125,
phosphorylase 120, 121, 423
353, 422
150, 151, 155
carbohydrate metabolism
pheophytin 130, 131
3-phosphoglycerate 150,
159
phosphates 13
151, 155
glycogen metabolism
ATP transfer 123
dehydrogenase 420
156, 157
buffers 288
kinase 131, 147, 424
kinase 424
ions 328
glycolysis 150, 151
phosphorylation 62, 106
residues 12
mutase 151, 429
ATP-dependent 114
transport 212
phospholipases
cyclic 130, 131
phosphatidates 47, 50, 51,
A1 425
dephosphorylation 114
170, 171
A2 268, 390, 391, 425
histones 238
cytidyltransferase 170,
C 386, 387, 425
oxidative 112, 122, 140,
171, 425
D 425
210
lysophosphatidate 170,
phospholipids 46, 47, 50-1,
protein kinase A 120, 121
171
51
substrate-level 122, 124,
phosphatase 170, 171, 425
biosynthesis 171
150
phosphatides 46, 47, 51
blood clotting 290
phosphoserine transami-
phosphatidic acids 46
degradation 411
nase 424
phosphatidylcholine 50, 51,
lipid metabolism 312
phosphatidylinositols
170, 171, 212, 213
lysophospholipids 50
1-phosphatidylinositol-
sterol acyltransferase
membranes 214
4-kinase 424
278, 279, 423
structure 50, 51
1-phosphatidylinositol-
phosphatidylethanolamine
phosphomevalonate kinase
4-phosphate kinase
50, 170, 171, 212, 213
424
424
phosphatidylinositol 50,
4’-phosphopantetheine 12
phosphatidylinositol
170, 171, 212, 213
phosphoprotein
(PtdIns) 170
phosphatidylinositol 4, 5-
phosphatase 394, 395,
phosphatidylinositol-3-
bisphosphate (PIP2) 170,
425
kinase (PI3K) 388
171
phosphates 120, 121
phosphatidylinositol-4,5-
phosphatidylinositol-4-
phosphopyruvate hydratase
bisphosphate (PIP2, Ptd
phosphate kinase 170, 171
124, 428
InsP2) 170
phosphatidylserine 50, 170,
5-phosphoribosyl 1-diphos-
phosphotransferases 89
171, 212, 213
phate (PRPP) 188, 189
photolyase see deoxyribodi-
phosphoadenosine phos-
phosphoribosylamine gly-
pyrimidine photolyase
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
459
photoreactivation 256
131
electron transfer 19
photoreceptors 358, 359
Q cycle 126, 127
equilibrium 126, 350
photosynthesis 106, 112,
reduced (QH2) 126, 127
group transfer 19
128-9, 129, 130-1
see also coenzyme Q, re-
membrane 350
photosystems (PS) 128
duced
normal 130, 131
I 131, 132, 133
pleated sheets
proton transfer 19
II 130, 131, 133
α-pleated sheet 67
resting 126, 350, 351
phototrophs 128
β-pleated sheet 68, 69
PPP see protein phosphate
phylloquinone see vitamin K
plots
pathway
phytol 52, 53, 128, 129
hyperbolic 93, 97
PQ see plastoquinone
phytyl 52, 53
Lineweaver—Burk 92, 93
pRb 394
Pi see inorganic phosphate
PLP see pyridoxal phosphate
prealbumin 64, 65
π electrons 4
point mutations 248, 257
precipitation 79
PI3K see phosphatidylinosi-
polar molecules 28
precursors 114, 138
tol-3-kinase
polarimetry 36, 37
pregnenolone 376
pigments, antenna 132
polarity 218
prenylation 62, 214
ping-pong mechanisms 90,
polarization 6
preproinsulin 160, 161
91, 94
depolarization 222, 348,
preprotein 230
pitch 68
350
primary structure 76
helix 86
hyperpolarization 348,
primase 240
PK see protein kinase
350
primers
PK-B see protein kinase B
membranes 348
DNA 240
pKa value 18
repolarization 350
PCR 262
plants 42-3
polarizers 36
sequencing 260
plaques 260
poliovirus 404
proaccelerin 291
plasma 274, 275
polyacrylamide gel electro-
probes, gene 260
buffers 288, 289
phoresis (PAGE)
processes, electroneutral
cells 294
polyadenylate
218
hormones 372-3, 373
polymerase 246
procollagen 344
hydrogen ions 288, 289
tail 243, 246, 247
lysine 5-dioxygenase
kallikrein 426
polyethylene glycol (PEG)
344, 345, 422
membranes see plasma
polymerase chain reaction
proline 4-dioxygenase
membranes
see PCR
344, 345, 422
pH 31
polynucleotidases 425
proconvertin 291
proteins 276-7, 277, 288
polynucleotides 80, 81
proconvertin see coagula-
thromboplastin
polysaccharides 40-5, 41,
tion factor VIIa
antecedent see coagu-
42-3
product 22
lation factor XIa
polysomes 250, 251
inhibition 144
plasma membranes 196
POMC 382
proenzymes 176
composition 217
mRNA 382
progesterone 56, 57, 374,
functions 217
prepro-POMC 382
376
lipids 215
pro-POMC 382
proinsulin 76, 160, 161
permeability 219
pores 208, 209
prokaryotes 196, 197
proteins 215
porins 210
proline 60, 61
structure 214-15, 215
porphobilinogen 192, 193
collagen 70
transport 219, 221
synthase 192, 193, 428
dehydrogenase 421
plasma thromboplastin
porphyrias 192
hydroxylase see procolla-
antecedent 291
porphyrin systems 194
gen proline 4-dioxyge-
plasmids 258, 259, 262, 263
post-translational modifica-
nase
plasmin 426
tion 62, 63, 344
promoters 114, 118
plasminogen activator 427
postabsorptive state 308
proopiomelanocortin see
plastocyanin 128, 129
potassium channels 222, 233
POMC
plastoquinone (PQ) 52, 53
potential
Propionibacterium 148, 149
photosystem I 133
action 222, 348, 350, 351
propionic acid 49
photosystem II 128, 129,
difference ( P) 16
fermentation 148, 149
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
460
Index
propionyl-CoA 166, 167
catalysis 64
prothrombin 291
carboxylase 166, 167,
channels 218
prothrombinase complex
429
clotting factors 290
291
proprotein 230
defense 64
proto-oncogenes 398, 399,
prostacyclins 390
denaturation 74, 75
400
prostaglandins 390
dynamics 72, 73
protofilaments 70, 204
prostaglandin H-syn-
elution 78
proton channels, ATP syn-
thase 390, 391, 422
Escherichia coli 251
thase 142, 143
prosthetic groups 104, 134,
folding see protein folding
proton gradients 126, 140,
142
globular 72-3
210, 212
proteasomes 176, 177
glycosylation 230, 231
membranes 126, 127
protein
gradients 127
protons
disulfide isomerase 233,
heat-shock (hsp) 232
excretion 326
429
hyaluronate 346
gradient see proton
glycosyltransferase 231,
hydrolysis 266
gradient
423
isolation 78
motive force ( P) 126
lysine 6-oxidase 344, 345,
lysosomal 234, 235
pumps 126, 127
421
marking 174
secretion 327
phosphatases 114, 144,
maturation 230-3
transfer 14, 18, 19
145, 392
mediator 224
protoporphyrin IX 192, 193
tyrosine kinase 424
membranes 214, 215, 216
protoporphyrinogen IX 192,
see also proteins
metabolism see protein
193
protein folding 72, 74-5, 75,
metabolism
proximity, substrates 90,
174
mitochondrial import
100
endoplasmic reticulum
232, 233
PRPP see 5-phosphoribosyl-
233
motive force 127
1-diphosphate
energetics 74
movement 64
pseudouridine 82, 83
rough endoplasmic retic-
non-histone 238
PTH see parathyroid hor-
ulum 232
nutrients 360
mone
protein kinases (PK) 114,
overexpression 262, 263
pulsatile release, hormones
394, 395, 424
plasma 277
372
ERK 388
polyproteins 382
purines 80, 81
inactivating 144
post-translational
biosynthesis 188-9, 189,
MEK 388
modification 63, 174
190, 417
oncogenes 398
preprotein 230
degradation 186, 187
PDK-1 388
proprotein 230
derivatives 352
PK-A 120, 121, 386, 387
regulation 64
ring 188, 189
PK-B 388
regulatory 114, 119
pyranoses 34
Raf 388
resorption 266
pyridine nucleotides 32, see
second messengers 348
ribosomal 250
also NAD+; NADP+
protein metabolism 174-5,
single—strand-binding
pyridoxal 368
175, 306
240
phosphate (PLP) 62, 178,
muscle 338, 339
sorting 174, 228-9, 229
358, 368
protein-glutamine γ-gluta-
stabilization 74
transamination 108,
myltransferase 423
storage 64
109
proteinases 176, 290
structure 64, 68-71
see also vitamin B6
proteinogenic amino acids
synthesis 230-1, 231
complex
60-1, 61
transport 64, 220-1
pyridoxamine 179, 368
non-proteinogenic 62-3
proteoglycans 34, 35, 340,
phosphate 108, 178, 179
proteins 64-5, 65
346, 347
pyridoxol 368
adaptor 388
proteohormones 380, 381
pyrimidines 80, 81
adhesive 346
proteolysis 174, 176-7
biosynthesis 188-9, 189,
allosteric 72
enzymes 177
190, 418
analysis 78
hormones 382
degradation 186, 187
biosynthesis 174
neuropeptides 382
ring 188, 189
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
461
pyrrolines
R conformation 117
antigen 296, 297
1-pyrroline-5-carboxy-
hemoglobin 280, 281
co-receptors 296
late dehydrogenase
R state 116
cytokines 392
421
R/S system 8
domain 224
pyrroline-5-carboxylate
radiation 256
effects of 225
reductase 421
radicals
glucagon 121
pyruvate 35, 99, 154, 155,
free 32, 256
hormones 120
338
hydroperoxyl 33
insulin 224, 225, 388
2-oxoglutarate 141
hydroxyl 33, 284
ion channels 126, 218,
carboxylase 159, 429
semiquinone 32, 33, 130,
222-4, 350, 384
decarboxylase 427
131
ionotropic 348, 354
dehydrogenase see pyru-
superoxide 284
lipophilic hormones 378,
vate dehydrogenase
tyrosine 190, 191
398
glycolysis 150, 151
radioimmunoassays (RIA)
membrane 216, 224-5
kinase 151, 159, 424
Raf 388
metabotropic 348, 354
redox reactions 19, 101
Ramachandran plot 66
neurotransmitters 354-5,
transport 212
Ras 388
355
pyruvate carboxylase 138,
rate constants 22, 23
nicotinic acetylcholine
139, 155
reaction
222
pyruvate decarboxylase,
centers 132, 133
oncogenes 398
fermentation 148, 149
P680 130, 131
photoreceptors 358, 359
pyruvate dehydrogenase
heat of 20, 21
signaling see receptor
(PDH) 210, 421
reactions
signaling
-phosphatase 425
catalyzed 25
steroid 378
Escherichia coli 134, 135
endergonic 112
substrates 388
glucose oxidation 147
enzyme specificity 88,
synapses 348
kinase 424
89
T-cell 224
tricarboxylic cycle 144,
enzyme-catalyzed 90, 93
see also individual recep-
145
exergonic 112
tors
pyruvate kinase 147
first-order 22, 23
reciprocal velocity 93
glucagon 158, 159
kinetics 22-3
recombination
glycolysis 150, 151
order 22, 23
repair 256
rates 22, 23, 88
somatic 302, 303
second-order 22, 23
recycling 328-9, 329
Q
specificity 88, 89
redox
spontaneous 18
coenzymes 104, 105, 106-
Q see coenzyme Q, oxidized
substrate specificity 88, 89
7, 107, 140
(ubiquinone)
uncatalyzed 90
see also flavins; heme
Q cycle 126, 127
reactive oxygen species
groups; iron—sulfur
QH2 see coenzyme Q,
(ROS) 33, 284, 285
centers
reduced (ubiquinol);
reagents
potential (E) 18, 19, 32
plastoquinone
enzymes 102, 103
processes 32-3
quaternary structure 76
modifying 97
reactions see redox reac-
quinol system 32
rearrangements 14
tions
quinones 104, 131
see also isomerizations
series 32, 130, 131, 142
hydroquinone 33, 104
receptor signaling 216
systems see redox sys-
redox systems 33
signal recognition par-
tems
semiquinone radical 32,
ticle (SRP) 230
redox reactions 14, 15, 32,
33, 104
signal transfer 224
33
system 32
receptor-mediated endocy-
lactate 19, 100, 101
see also plastoquinone
tosis 278
NAD+ 18, 19, 101
receptors
pyruvate 19
1-helix 224, 384
redox systems 32
R
7-helix 224, 384
biological 32, 33
acetylcholine 354
coenzyme Q (CoQ) 105
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
462
Index
flavin 33
components 132, 140, 141
ribosomal RNA see rRNA
hydrogen peroxide 33
redox systems 142, 143
ribosomes 82, 236
respiratory chain 142, 143
respiratory control, ATP
eukaryotes 250, 251
reduced forms, electrons 32
synthase 144, 145
fractionation 198
reducing agents 32
resting potential 126, 350,
proteins 250
reductases 89
351
ribozymes 24, 88
reduction 316
restriction endonucleases
ribuloses
equivalents 32, 33, 212
258, 259, 264, 425
D-ribulose 38, 39, 128, 152
glucose 36, 37
restriction fragment length
ribulose phosphate 3-
Refsum disease 166
polymorphisms see RFLP
epimerase 428
regions, signal 228
reticuloendothelial system
L-ribulose phosphate 4-
regulation 114-21, 144-5
(RES)
epimerase 428
allosteric 116
retinal 62, 358, 364
ribulose 5-phosphate
carbohydrate metabolism
isomerase 358, 359, 429
152, 153
158-9
retinoate 52, 53
ribulose bisphosphate,
gene 118
retinoic acid 364
carboxylase 130, 131,
glycogen metabolism 156
retinol see vitamin A
427
interconversion 120
retinol, dehydrogenase 358,
Richardson diagrams 72, 73
transcription 118-20, 244
359, 420
rifamycin 254, 255
tricarboxylic cycle 144
retroviruses 404
righthanded helix
regulatory proteins 114, 118,
reverse transcriptase see
α-helix (αR) 68
119
RNA-directed DNA poly-
double 86
relative velocity 25
merase
RNA 80, 81, 82-3, 83, 87
renal hormones 330-1, 331
RFLP 265
conformation 86
renin 330, 331, 427
sickle-cell anemia 262,
hnRNA 236, 246, 247, 302
renin—angiotensin system
264
immunoglobulins 302
330, 331
rhesus (Rh) blood groups
maturation 208, 236, 242,
repair, mutations 256, 257
292
243, 246-7
replication 208
rhinovirus 404, 405
transcription 242, 243
DNA polymerases 240
Rhizobium 184
translation 83
Escherichia coli 240, 241
rhodopsin 224, 358
see also mRNA; rRNA;
forks 240
RIA see radioimmunoassay
snRNA; tRNA
genetic information 236,
riboflavin 366
RNA polymerase see DNA-
240-1
ribonucleic acid,
directed RNA polymerase
origin of 240
transcription 243
RNA-binding proteins 82
repolarization 350
ribonuclease
RNA-directed DNA poly-
repression, transcription
A 74, 75
merase 404, 405, 425
114
H 404, 405, 426
rods 358
repressors 118
pancreatic 426
root nodules 184, 185
rER see rough endoplasmic
ribonucleic acid (RNA) see
ROS see reactive oxygen
reticulum
RNA
species
RES see reticuloendothelial
ribonucleoside diphosphate
rotation 67
system
reductase 190, 191, 403,
rough endoplasmic reticu-
residual bodies 234
422
lum (rER) 226, 227, 231,
resonance 4, 5, 32, 66, 67
ribonucleotide
232
resorption 266, 267, 272-3,
reductase see ribonucleo-
roughage 42, 273
273
side diphosphate re-
rRNA 82
HCO- 326
ductase
5S-rRNA 86, 87
liver 309
reduction 191
RT-PCR 264, 265
minerals 362
ribose 12, 13, 123
rubber 52, 53
urine 322
phosphate pyrophospho-
rubisco see ribulase bis-
respiration 144, 145, 146-7
kinase 424
phosphate carboxylase
respiratory chain 106, 112,
D-ribose 34, 38, 39
126, 140-1, 210
ribose 5-phosphate 152, 153,
S
ATP synthesis 142
429
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
463
S see Svedberg units
sequential mechanisms 90,
oncogenes 398
S phase 394
91, 94, 95
proteins (STPs) 392
saccharase see β-fructofura-
sER see smooth endoplasmic
signal transmission
nosidase
reticulum
CNS 348-9
Saccharomyces cerevisiae
serine 50, 51, 60, 61, 63
synaptic 348
148, 149
dehydratase 181, 428
signaling
SAH see S-adenosyl-L-ho-
family 184, 185
amino acids 380, 381
mocysteine
proteases 176, 177, 298
calcium 342
salicylate 317
proteinases 177, 425
cytokines 392-3
saliva 268, 269
serotonin 62, 63
eicosanoids 46, 390-1
salt, precipitation 78, 79
serum 274
hydrophilic substances
SAM see S-adenosyl-L-me-
enzyme diagnosis 98, 178
392
thionine
seryl alanine 65, 66, 67
membranes 216
sarcomeres 332
seven-helix receptors 224
second messengers 224
sarcoplasmic Ca2+ pump
SH2 domain 392
steroids 46
220, 221
short tandem repeats (STRs)
silencers 118
sarcoplasmic reticulum (SR)
264
silk 70, 71
334, 335
shuttles 212
single—strand-binding pro-
scaffolding
sialic acid see N-acetylneur-
teins 240
cytoskeletal 206
aminic acid
single-stranded DNA
nuclear 238
sickle-cell anemia 249, 264,
(ssDNA)
Schiff’s base 108
265
site-specific
SDS gel electrophoresis 78,
sight 358-9
deoxyribonuclease see
79
sigmoidal substrate satura-
restriction endonucleases
second messengers 224, 348
tion curves 116
β-sitosterol 56, 57
glycolipids 50
signal
size, molecular 218
hormones 120, 121, 384,
cascades see signal cas-
skeletal muscles 332
386-7
cades
small intestines 31, 268,
see also individual mes-
peptidase 427
269
sengers
peptides 228, 230, 231
small nuclear ribonucleo-
second-order reactions 22,
receptor transfer 224
protein particles
23
regions 228
(snRNPs) 246
secondary structure 68-9, 76
signaling see signaling
small nuclear RNA (snRNA)
secretin 270
SRP see signal recognition
82
secretions
particle
smooth endoplasmic retic-
kidney 322
stop-transfer 230
ulum (sER) 172, 173, 226,
pancreas 268-9
transduction see signal
227
small intestines 268-9
transduction
smooth muscle 332, 338
secretory pathway 228
translocation 228, 229
sn see stereospecific num-
sedimentation coef cient
transmission see signal
bering
200
transmission
SNAREs 228, 229
sedoheptulose bisphospha-
signal cascades
snRNA, splicing 83
tase 425
hormones 388-9
snRNPs (snurps) 246
sedoheptulose 153
photoreceptors 358, 359
soap bubbles 28, 29
selenocysteine 6, 63
sodium
self-protection, blood 274
signal recognition particle
chloride dissolution 21
semiquinone radical 32, 130,
(SRP) 230, 231
dodecylsulfate see SDS
131
receptor 230
ions 328
sense strands 84
signal transducers and acti-
voltage-gated channel
separation, spontaneous 29
vators of transcription see
222, 223
sequences
STAT
solenoids 238
glycosylation 230
signal transduction 120
solubility 28, 29
nuclear localization 208
cytokines 392, 393
soluble coenzymes 104
peptides 66
G proteins 384, 385
solvents 26-7, 54
sequencing 261
insulin 388, 389
somatic mutation 302
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
464
Index
somatic recombination 302
hormones see steroid
lation 150
somatotropin 64, 65
hormones
substrates
receptor 64
receptors 378
activity 95
son of sevenless see SOS
signaling 46
analogs 96, 97
sorbitol 36, 37
steroid 11β-monooxyge-
bisubstrate kinetics 94, 95
D-sorbitol 39
nase 422
coenzymes 105
sorting, proteins 174, 228-9,
steroid 17α-monooxyge-
concentration 97, 116
229
nase 422
cosubstrates 104
SOS, adaptor proteins 388
steroid 21-monooxyge-
enzymes 94
space
nase 422
insulin 388
space, intermembrane 210
steroid
-isomerase 429
orientation 90, 100
matrix 210
structure 54
phosphorylation 122,
spectrin 206
synthesis 410
124, 125
spectrophotometry 102, 103
sterol esterase 268, 269
proximity 90, 100
sphingolipids 46, 47, 51, 170
sterol esters 47
receptors 388
sphingomyelin 214, 215
sterols 46, 54, 55, 56, 57
specificity 88, 89, 94
sphingophospholipid 51
see also cholesterol;
suicide 96, 97
sphingosine 51
ergosterol; β-sitos-
succinate 136, 137
N-acetyltransferase 423
terol; stigmasterol
dehydrogenase 136, 137,
spliceosomes 246, 247
stearylglycerols,
140, 421
splicing
tristearylglycerol 49
glucose oxidation 147
hnRNA 246, 247
stigmasterol 56, 57
semialdehyde dehydro-
snRNA 83
stop codons 248, 250
genase 356, 357, 420
spontaneous separation 29
stop-transfer signal 230
succinate-CoA ligase 136,
squalene 52, 53, 172, 173
storage, liver 306
137, 147
SR see sarcoplasmic reticu-
STP see signal transduction
(GDP-forming) 429
lum
protein
succinic acid 8, 9
ssDNA see single-stranded
strands
succinyl phosphate 125
DNA
double 85
succinyl-CoA 136, 137, 166,
stabilization
grooves 86
167
catalysis 90
matrix 85
heme biosynthesis 192,
transition states 100, 101
sense 85
193
starches 40, 42, 43
Streptococcus 148, 149
ligase 124, 125
brewing 148, 149
Streptomyces lividans 222,
sucrose 38, 39
start codons 248, 250
223
α-glucosidase 426
STAT 392
streptomycin 254
density gradients 201
statins 372
striated muscle 333
sugar 38
stationary phase 54
stroma 128, 129
alcohols 38, 39
stearic acid 49
STRs see short tandem
see also mannitol;
stearoyl-CoA desaturase 422
repeats
sorbitol
stercobilin 195
structural genes 118
amino 44
stercobilinogen 194, 195
structure 6
suicide substrates 96, 97
stereospecific numbering
dimers 116
sulfathiazole 255
(sn)
isomers 8
sulfatides 50, 51
stereospecificity 8
polysaccharides 40
sulfides, disulfide 11
steroid alcohols see sterols
primary 76
sulfonamides 254, 255
steroid hormones 56, 57,
proteins 68, 70-1
sulfur 11
172, 374, 377
secondary 76
sulfur-containing amino
biosynthesis 376
tertiary 76
acids 60
conjugates 110
quaternary 76
superoxide
inactivation 376
Stuart—Prower factor see
dismutase 284, 285, 422
metabolism 376-7, 377
coagulation factor Xa
radicals 284
see also individual
Stuart-Prower factor 291
surface films 29
hormones
substitution, nucleophilic 14
Svedberg unit (S)
steroids 46, 52, 53, 54-7, 55
substrate-level phosphory-
sweat 31
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
465
symbiosis
tetrahydrocortisol 317
thyrotropin 380
endosymbiotic theory
tetrahydrofolate (THF) 108,
thyrotropin-releasing hor-
210
110
mone (TRH) see thyroli-
nitrogen fixation 184, 185
N5-formyl-THF 108
berin
symport 212, 218, 220
N5-methyl-THF 108
thyroxine 62, 374
synapses 348
N5-methylene-THF 108
thyroxine-binding globulin
receptors 348
N10-formyl-THF 108
(TBG)
signal transmission 348,
tetraiodothyronine see thy-
TIM 232
349
roxine
tissue kallikrein 426
vesicles 354
tetramers 99
tissue plasminogen activa-
synaptobrevin 228, 229
tetrapyrroles 128, 129
tor (tPa) see plasminogen
synaptotagmin 228, 229
THB see tetrahydrobiopterin
activator (tissue)
Synechococcus elongatus
Thermodynamics, Second
tissues
130, 132
Law of 20
disruption 198
syntaxin 228, 229
thermogenin (UCP-1) 144,
hormones 370
synthases see lyases
145
titin 332
synthetases see ligases
THF see tetrahydrofolate
titration curves 30
thiamine see vitamin B1
TLC see thin-layer chroma-
thiamine
tography
T
diphosphate (TPP) 106,
TMP, dTMP 80, 81, 187
107, 366
TNF-a see tumor necrosis
T cell 274, 275, 294
pyruvate dehydrogen-
factor-a
activation 296-7, 297
ase 134, 135
tobacco mosaic virus 404
receptor 224, 225
see also vitamin B1
tocopherol see vitamin E
T form 117
thiazole ring 106
TOM 232
hemoglobin 280, 281
thin-layer chromatography
topoisomerases 240, 245,
T state 116
(TLC) 54, 55
429
T4 phage 404
thioesters 10, 11, 106, 125
toxicity, urea 182
TAFs see TATA box, binding
bonds 12, 13
TPP see thiamine diphos-
protein, associated fac-
thioethers 10
phate
tors (TAFs)
thiokinase see succinate-
trace elements 2, 360, 362-3
tail, polyadenylate 246
CoA ligase (GDP-form-
TRADD 396
TATA box 242, 243
ing); succinyl-CoA ligase
transducin 224
binding protein (TBP)
thiols 10, 11
transferrin 286
244, 245
thiopurine methyltransfer-
transacetylase 119
associated factors
ase 402, 403, 422
transaldolase 152, 153, 422
(TAFs) 244, 245
thioredoxin 190, 191
transaminases 94, 176, 177
taurine 314
reductase 190, 191, 421
transamination 134, 178-9,
taurocholic acid 314, 315
threonine 60, 61, 63
179, 180
TBG see thyroxine-binding
aldolase 428
pyridoxal phosphate 108,
globulin
synthase 428
109
TBP see TATA box, binding
thrombin 290, 426
transcription 208, 242-5,
protein
thrombocytes 274, 290
243
teeth 340, 341
thromboplastin 291
control 114, 115, 118-20,
temperature, enzymes 94, 95
thromboxanes 390
244-5
templates 84, 240
thylakoids 128, 129, 133
factors 114, 118, 119, 392,
terminal oxidation 136
thymidine 80, 81, 82, 304
394
termination, Escherichia coli
thymidylate synthase 190,
genetic information 236
251
191, 402, 403, 422
initiation 244, 245
terpenes 52, 53
thymine 81, 84, 186, 187
PEP-CK 244, 245
tertiary structure 76
dimers 256
regulation 244
testosterone 56, 57, 338,
thyroid-stimulating hor-
RNA 242, 243
374, 376
mone (TSH) see thyrotro-
transducin 72, 73, 358
tetracyclines 254
pin
transfer RNA see tRNA
tetrahydrobiopterin (THB)
thyroliberin 380
transferases 88, 89
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
466
Index
C1-transferases 89
triacylglycerol 48, 49, 163,
352, 353, 422
intermolecular 89
170, 171
transferrin 287
lipase 270, 271, 272, 273,
U
transformation 258, 398,
425
401
tricarboxylic acid cycle 113,
bio 226, 317
136-9, 137, 139, 210
ubiquinol see coenzyme Q,
tumors 400
and amino acid
reduced
transition states 15, 90, 100
biosynthesis 185
ubiquinone see coenzyme Q,
analogs 97
regulation 144, 145
oxidized
stabilization 100, 101
triiodothyronine 374
ubiquitin 62, 176, 177, 394
transketolase 152, 153, 422
see also thyroxine
UCP-1 see thermogenin
translation 174, 208
triokinase 310, 311, 424
UDP glucose 110, 111, 156
amino acids 60
triose phosphate isomerase
4-epimerase 310, 311, 428
UDP glucose 1-phosphate
elongation 252-3
150, 151, 429
Escherichia coli 250, 251,
Tris(hydroxymethyl)amino-
uridyltransferase 157
252, 253
methane
UDP glucuronic acid 194
genetic information 236,
tristearylglycerol, van der
UDP-glucuronosyltransfer-
248, 250-3
Waals model 49
ase 194
ultrafiltration 322
inhibitors 254
tRNAPhe 82, 83, 86, 87
initiation 250-1
aminoacyl 248
ultraviolet radiation (UV)
RNA 83
genetic information 236
256
termination 252-3
tropines 372
UMP 187, 188, 189, 190
translocation 252
tropomyosin 64, 65, 332,
uncouplers 144, 145
signals 228, 229
334
uniport 212, 218
translocator complexes
troponin 332, 334
units, international (U) 88
TIM 232
trypsin 94, 95, 177, 268, 269,
unsaturated fatty acids 48
TOM 232
426
uracil 80, 81, 84, 186, 187
translocon 230
trypsinogen 176, 177, 270
urea 324
transmembrane helix 214
tryptophan 60, 61
cycle 113, 182-3, 183
transport
TSH see thyroid-stimulating
degradation 112
active 218
hormone
liver 182
antibiotics 254
tubules 210, 221
ureotelic animals 182
antiport 212, 218
microtubules 204, 206,
uric acid 186, 187, 324
blood 274
207
uricase 186
CO2 282, 283
transverse 334
uricotelic animals 182
hemoglobin 282, 283
tubulins 204, 205
uridine 80, 81
hormones 370
tumor necrosis factor-α
5’-diphosphate see UDP
5’-monophosphate see
lipoproteins 279
(TNF-α) 396
mechanisms 220
tumor-suppressor genes
UMP
membrane regulation
394, 398
5’-triphosphate see UTP
216
tumors 400-1, 401
dihydrouridine 82, 83
mitochondrial 212-13
gene therapy 264, 265
pseudouridine 82, 83
urine 31, 322, 323, 324-5,
O2 282-3, 283
monoclonal antibodies
passive 218
302
325
plasma membranes 219
turnover
urobilin 194, 195
processes 218-19
enzymes 88, 89
urobilinogen 194, 195
proteins 220-1
number 92
urocanate hydratase 428
secondary 220
β-turns 68
urokinase see plasminogen
symport 212, 218, 220
type 1 diabetes 160
activator (urine)
uniport 212, 218
type 2 diabetes 160
uroporphyrinogen III 192,
transporters 35, 210, 212,
tyrosine 60, 61, 352
193
218, 220
kinases 224, 392
synthase 192, 193, 428
transthyretin 64, 65, 276
radical 190, 191
UTP 190
transverse tubules 334
transaminase 424
UTP-glucose-1-phosphate
α, α-trehalase 426
tyrosine-3-monooxygenase
uridyltransferase 424
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.
Index
467
UV see ultraviolet radiation
vitamins 365
emulsions 28
A (retinol) 52, 53, 364, 365
exclusion of 90
antioxidants 364
excretion 322
V
B1 (thiamine) 366, 367
hydrophobic interactions
B2 complex 366, 367
28-9
V segments 302
B6 complex 368, 369
nutrients 362
V systems 116
B12 (cobalamine) 108,
O2- 284
V type ATPase 220
368, 369
recycling 328-9, 329
V/J recombination 302, 303
C (L-ascorbic acid) 104,
solvents 26-7
vacuoles 43, 196, 234
105, 368, 369
structure 26
valerianic acid 49
coenzymes 364
water-soluble vitamins 366-
valine 60, 61
D (calciol, cholecalciferol)
9, 367, 369
van der Waals models 6, 7
52, 53, 364, 365
waterbinding polysacchar-
acetyl-CoA 13
deficiency diseases 364
ides 40
cholesterol 54, 55
E (tocopherol) 32, 52, 53,
watersplitting enzyme 130
tristearylglycerol 49
104, 364, 365
wavelength, constant 102,
variability, immunoglobu-
H (biotin) 62, 108, 109,
103
lins 302
368, 369
waxes 47
variation
hormones 364
work 17
allotypic 302
K (phylloquinone) 52, 53,
energy 16
idiotypic 302
104, 364, 365
wort 148, 149
vasopressin see antidiuretic
lipid-soluble 46, 364-5,
hormone (ADH)
365
vectors 258, 259
X
nutrients 360
velocity
water-soluble 366-9, 367,
and concentration 93, 97
369
xanthine 187
maximum 92, 93
VLDL 163, 278, 312
oxidase 186, 187, 420
reciprocal 93
xyloglucan 41
Vmax see maximum velocity
relative 25
voltage-gated Na+ channels
D-xylose 38, 39, 42
very-low-density
222
xylulose 5-phosphate 153
lipoprotein see VDLV
vesicles 28, 29
fractionation 198
W
Y
synaptic 354
vimentin 332
water 11, 27, 65, 296
yeasts 148, 149
viruses 404-5, 405
acids 31
DNA 264, 265
bacterial cells 202
Z
rhinovirus 404, 405
degradation 112
Koolman, Color Atlas of Biochemistry, 2nd edition © 2005 Thieme
All rights reserved. Usage subject to terms and conditions of license.